External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G20860 159 / 2e-48
ATNEK5
NIMA-related kinase 5 (.1)
AT3G44200 134 / 1e-37
IBO1, ATNEK6
"NIMA \(never in mitosis, gene A\)-related 6", NIMA-RELATED KINASE6, NIMA (never in mitosis, gene A)-related 6 (.1)
AT1G54510 130 / 1e-36
ATNEK1
NIMA-related serine/threonine kinase 1 (.1.2.3)
AT5G28290 123 / 5e-34
ATNEK3
NIMA-related kinase 3 (.1)
AT3G04810 122 / 1e-33
ATNEK2
NIMA-related kinase 2 (.1.2)
AT3G63280 120 / 3e-33
ATNEK4
NIMA-related kinase 4 (.1.2)
AT3G12200 107 / 3e-28
ATNEK7
NIMA-related kinase 7 (.1.2)
AT3G53930 57 / 2e-10
Protein kinase superfamily protein (.1.2)
AT1G73690 56 / 3e-10
CDKD1;1, AT;CDKD;1, CAK3AT
cyclin-dependent kinase D1;1 (.1)
AT1G30640 54 / 1e-09
Protein kinase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031834
210 / 1e-67
AT3G20860 563 / 0.0
NIMA-related kinase 5 (.1)
Lus10040499
145 / 8e-42
AT3G20860 476 / 2e-164
NIMA-related kinase 5 (.1)
Lus10011301
144 / 4e-41
AT3G20860 470 / 2e-157
NIMA-related kinase 5 (.1)
Lus10001857
126 / 5e-35
AT3G44200 854 / 0.0
"NIMA \(never in mitosis, gene A\)-related 6", NIMA-RELATED KINASE6, NIMA (never in mitosis, gene A)-related 6 (.1)
Lus10013338
126 / 7e-35
AT3G44200 845 / 0.0
"NIMA \(never in mitosis, gene A\)-related 6", NIMA-RELATED KINASE6, NIMA (never in mitosis, gene A)-related 6 (.1)
Lus10042632
125 / 8e-35
AT3G04810 671 / 0.0
NIMA-related kinase 2 (.1.2)
Lus10009186
125 / 1e-34
AT1G54510 695 / 0.0
NIMA-related serine/threonine kinase 1 (.1.2.3)
Lus10015928
125 / 1e-34
AT1G54510 694 / 0.0
NIMA-related serine/threonine kinase 1 (.1.2.3)
Lus10001783
124 / 2e-34
AT1G54510 705 / 0.0
NIMA-related serine/threonine kinase 1 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0016
PKinase
PF00069
Pkinase
Protein kinase domain
Representative CDS sequence
>Lus10031264 pacid=23151030 polypeptide=Lus10031264 locus=Lus10031264.g ID=Lus10031264.BGIv1.0 annot-version=v1.0
ATGGAAGGGAGGAGTACTAGAGGAGGAGAAGACGATTACGAAGTGAATGAGCAGATAGGGAGGGGGAGATTCGGAGCTGCGTTTCTCGTCATTCGTAAGT
CGGAGAACAAGAGGTATGTGATGAAGAAGATCAAGTTGGCTAGGCAGACTGAAAGGTTCAAGCAGACTGCTATCCAGGAGATGAACTTGATAGCCAAGTT
GAATCACCCGTACATAGTGGAGTACAAGGACTCCTGGGTTGGAAAGGAATGCGTGTGTATCGTTACAAATTACTGCGAGGGTGGCGACATGACTCAGGTG
ATAAGGAAAGCTAGAGGCTCTAAAGGTCTTGATTTTTGA
AA sequence
>Lus10031264 pacid=23151030 polypeptide=Lus10031264 locus=Lus10031264.g ID=Lus10031264.BGIv1.0 annot-version=v1.0
MEGRSTRGGEDDYEVNEQIGRGRFGAAFLVIRKSENKRYVMKKIKLARQTERFKQTAIQEMNLIAKLNHPYIVEYKDSWVGKECVCIVTNYCEGGDMTQV
IRKARGSKGLDF
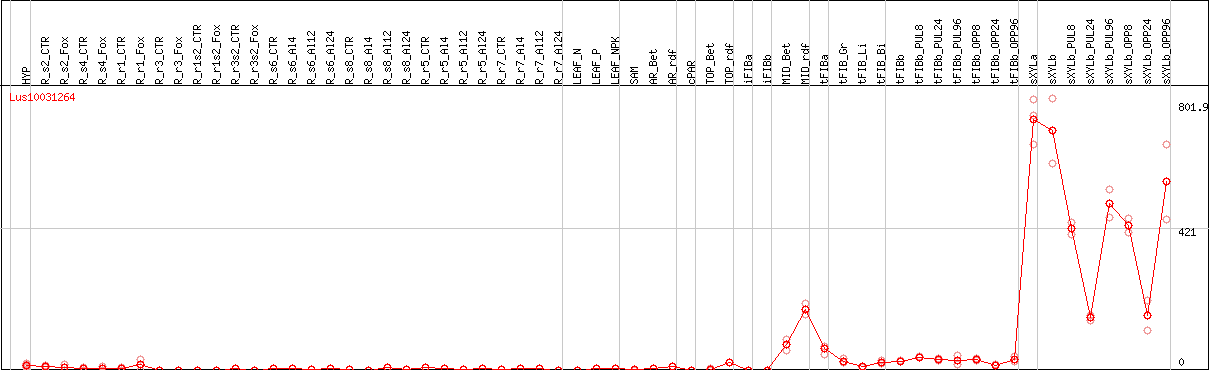
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031264 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.