Lus10031332 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10031332 pacid=23151170 polypeptide=Lus10031332 locus=Lus10031332.g ID=Lus10031332.BGIv1.0 annot-version=v1.0
ATGGTGTTCAAGAACAAGCTCTTCTTCTCGTCGAAGAAGTCGTCATCGGATGCATCAAGTCCGACCGATGGTGGTGGATCTTCCAACAGTCCTCGATCCA
TCGGGTCGAACTCCCCGATCCGGAGCAGCGAGAAGAAGAAGTCTAAGTCGTCTTCGTCTTTCTCGTCCTCGTCGAAGGACAACAGCCCCAGCTTTTCCTC
CGCGGGTGCCGCGGCGGGGGCGGCCGGGTGTAAGCAGGTTCAGGTGAAGGATGGGCTGAAGAAGAAGGAGAGCGTCGTGAAGGGGAAAGAGAGTGTTCAA
TTGATCGGTAAGGCTGCTTCTCCTGCTCATCATCCTAAGTCTCAGCCGCAAACGCCTGTTAAGCAAGGCGGAGGCGGAGGAGGCTTCTCTGCTTCGGTAC
TTGGTTCTACTTCTGCTTCACCTTCCGCTTCAGGGTCTAAGTTGAGGAAGGCTTTGGGGAGTGGTAAGGAGGATGCGTCGTCTGTGTCTCCGATATTGGC
TTCTTCGTTGGGGTTGAATAAGATAAAGACACGGTCAGGGCCATTGCCTCAGGAGAGTTTCTTTGGTTATAGAGGGGATAAGGGGTCTGGGGTGCTTGGT
TCCAGTAATTTGTCGAGGCCTGGTGGGGGGGATGGCGGATTCCGTTCGAATTCGGTGAAGAAGGAGGATGGTACTGGAAAGGGTAGGATGACAGGGTTTC
AGGACCGCAGTGGTAAAGCCGTTGATACTAGCAGCAATTGGGATCGGGTGTCTTCTGGTAGTGGAGCGCAGACTAGAGAGGTTACTCCAAATCCGCAGCC
AGGGATGAACTTGGACAATGGGGAACCATCTAACGATGCAGGTCGGAATGAGTCATCCTGGGGTCGTTCTGGTGGTGTGAGAAGTTCGGATGTTTTTACA
CCAGAGGAGTCTGAGTCGCCACGTCTCCAAGCTATCCTACGTCTAACTAGTGCACCAAGAAAGAGATTTCCCTCTGATATAAAAAGTTTTTCTCATGAAC
TCAATTCCAAAGGCGTGCGCCCATTCCCCTTTTGGAAGCCTCGTGGACTGAATAATGTGGAGCAAGTATTGGTTGTTATACGAGGAAAGTTTGACAAGGC
AAAGGAAGAAGTGAATTCGGATCTGGCTATCTTTGCAGCTGATCTTGTTGGAATCCTTGAAAAGAATGCAGAAAGTCATCCTGAATGGCAGGAAACTATT
GAGGACTTGTTGGTCTTGGCCAGGAGTTGTGCTATGCAATCACCTGGAGAGTTTTGGCTTCAATGTGAAAGTATAGTCCAAGATTTGGATGACAGACGCC
AAGAACTCCCTCCAGGAACGCTGAAGCAACTTCATACTAAAATGCTTTTCATTCTGACTAGGTGTACAAGACTTTTGCAGTTCCACAAAGAAACTGGACT
TCCCGATGATGAGAATATTTTTCAACTGCGTCAATCTAGGGTCTTACAAGTACACTCCGCAGAGAAACGCCTTCCTCCAGGTGCTGGAAGGGGCATGAAA
AGCTCCACAGCTTCAAAAGTTGCAAAAGGGACTTCAGTGACGAAATCGTACAGTCAAGAACAACACGGGCGAGGTTGGAACCGAGAGCCATTGGTACTGC
CTGCAAACTTTCTTTCTCCCACTGAGGCTTCCATTAATGTTGAGTCTCCTGGAGGTCGAGATAGGATGGCTTCCTGGAAAAAGCTTCCATCACCTGCTGC
AAAAGGTCCAAAGGAAATTGCTTCACCTAAGGATCAAAATGAGGTAAGGGCTGATCCTTTGAAAGTGCCTAGCAACACTAAAGGGATTTCTGATATCGAT
CTGCCTGCTGCTAAGCCATCTGAGGTCCCCACAGTGAAACTTGAACATTCTGCGAAGCACCAACATAAAGTTTCTTGGGGATACTGGGGTGACCAAAATC
CTGCAGATGAAAATTCAATTATTTGTCGAATTTGCGAGGAAGAGGTTCCAACTTCATATGTTGAAGACCATTCAATAATATGTGCGATTGCAGATCGTTA
TGATCAGAAAGGTCAAGGTGTGAATGAGCGCCTTTCAAAAATTGCAGAAACTCTTGAGAAAATGATGGAGTCTTTTGCACAGAAGGATATCCAACCTGCT
CTAGGGAGCCCTGATGTAGCAAAGGTGTCAAATTCTAGTGTTACTGAAGAGTCCGACATTCCATCTCCCAAGCTAAGTGACTGGTCTAGAAGAGGATCTG
AAGACATGCTCGATTGTTTTCCTGAAGCTGATAACTCGCCATTCACGGATGACATGAAGGGCTTGCCATCCATGTCATGCAGAACACGTTTTGGTCCAAA
ATCTGATCAGGGGATGGCCACTTCATCTGCTGGTAGCATGACTCCTAGATCTCCATTATTGACACCAAAGAACAGCCAGATAGACTTGCTTCTTGCTGGA
AAAGGTGGTTATCCAGAGCATGATGACCTTCCTCAGGATGATGGTTATCTGTTTTATTTTGCTAAGAGTACTGCTTGTTTGATACAGATGAATGAACTAG
CTGATATTGCAAAATGTGTGTCTACTATGCCTTTGGAGGATGAGCGTTCCGTGCCCTATTTATTATCCTGCCTTGAGGACCTGAGGGTTGTTATTGACAG
AAGAAAGTTTGATGCTCTTACTGTGGAAACTTTTGGGGCAAGGATTGAGAAACTTATACGAGAGAAGTATTTGCAGCTCTGTGAGCTAGTGGAAGATGAG
AAGGTTGATATAGGAAACACTGTGATCGATGAAGATGCTCCCTTGGAAGATGATGTAGTTAGGAGCCTGAGAACAAGCCCAATACACTCTTCGAAGGACC
GCACATCAATTGATGACTTTGAGATAATGAAGCCAATCAGTCGCGGGGCATTTGGGCGGGTATTCCTGGCTAAGAAAAGAACGACTGGAGATCTTTTTGC
CATAAAGGTCCTAAAGAAGGCTGATATGATACGCAAGAATGCTGTTGAGAGTATACTAGCAGAACGCGATATTCTAATTTCCGTGCGCAATCCATTTGTG
GTTCGTTTCTTCTATTCGTTTACTTGTCGTGAGAACTTGTACCTTGTGATGGAGTATTTAAACGGAGGCGACTTGTATTCATTATTACGTAATATGGGCT
GCTTAGATGAAGACGTTGCTCGGGTCTATATAGCCGAAGTCGTACTTGCATTGGAATATCTACACTCATTGCGCGTGGTTCATCGTGATCTGAAGCCTGA
TAATCTACTCATTGCACACGATGGTCATATCAAGTTAACAGATTTCGGGCTTTCGAAGGTCGGGCTCATCAACAGCACAGATGACTTGTCTGGTCCAGCG
GCTGGTGGAACTTATATGCTTGTGGACGACGAACCTGAACTGTCGACTTCAGATCAACAAGAAAGGAGGAAGAAACGTTCTGCTGTGGGCACACCCGACT
ATTTGGCACCAGAGATTCTTTTGGGAAGAGGACACGGCTTTACTGCTGATTGGTGGTCTGTGGGCGTCATTTTGTTCGAACTCATTGTTGGTATTCCTCC
TTTCAATGCAGAGCATCCTCAGATAATATTCGACAACATTCTCAATCGGAAGATACCGTGGCCCGAAGTACCTGACGAAATGAGCCCCGAAGCTCGAGAT
ATTATCGATAGATTTTTAACAGAAGATCCTCATATGAGACTCGGAGCCAGAGGTGCATCAGAGGTAAAGCAACACGTGTTCTTCAAAAACATCAACTGGG
ACACACTGGCAAGGCAAAAAGCTGCATTCGTGCCCGCTTCAGACAGCGCTCTCGACACGAGCTACTTCACAAGCCGGTACACACGGAGCAGGGAGATGAA
TGTGGAGGTCTCGCAGAGTTCGATTCTGAATCTTTCCCAGCTCGCGTCGATCAACTACGATCTTCTATCAAAGGGTTGGAAGGATGATCCTCCTACAAGC
TCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10031332 pacid=23151170 polypeptide=Lus10031332 locus=Lus10031332.g ID=Lus10031332.BGIv1.0 annot-version=v1.0
MVFKNKLFFSSKKSSSDASSPTDGGGSSNSPRSIGSNSPIRSSEKKKSKSSSSFSSSSKDNSPSFSSAGAAAGAAGCKQVQVKDGLKKKESVVKGKESVQ
LIGKAASPAHHPKSQPQTPVKQGGGGGGFSASVLGSTSASPSASGSKLRKALGSGKEDASSVSPILASSLGLNKIKTRSGPLPQESFFGYRGDKGSGVLG
SSNLSRPGGGDGGFRSNSVKKEDGTGKGRMTGFQDRSGKAVDTSSNWDRVSSGSGAQTREVTPNPQPGMNLDNGEPSNDAGRNESSWGRSGGVRSSDVFT
PEESESPRLQAILRLTSAPRKRFPSDIKSFSHELNSKGVRPFPFWKPRGLNNVEQVLVVIRGKFDKAKEEVNSDLAIFAADLVGILEKNAESHPEWQETI
EDLLVLARSCAMQSPGEFWLQCESIVQDLDDRRQELPPGTLKQLHTKMLFILTRCTRLLQFHKETGLPDDENIFQLRQSRVLQVHSAEKRLPPGAGRGMK
SSTASKVAKGTSVTKSYSQEQHGRGWNREPLVLPANFLSPTEASINVESPGGRDRMASWKKLPSPAAKGPKEIASPKDQNEVRADPLKVPSNTKGISDID
LPAAKPSEVPTVKLEHSAKHQHKVSWGYWGDQNPADENSIICRICEEEVPTSYVEDHSIICAIADRYDQKGQGVNERLSKIAETLEKMMESFAQKDIQPA
LGSPDVAKVSNSSVTEESDIPSPKLSDWSRRGSEDMLDCFPEADNSPFTDDMKGLPSMSCRTRFGPKSDQGMATSSAGSMTPRSPLLTPKNSQIDLLLAG
KGGYPEHDDLPQDDGYLFYFAKSTACLIQMNELADIAKCVSTMPLEDERSVPYLLSCLEDLRVVIDRRKFDALTVETFGARIEKLIREKYLQLCELVEDE
KVDIGNTVIDEDAPLEDDVVRSLRTSPIHSSKDRTSIDDFEIMKPISRGAFGRVFLAKKRTTGDLFAIKVLKKADMIRKNAVESILAERDILISVRNPFV
VRFFYSFTCRENLYLVMEYLNGGDLYSLLRNMGCLDEDVARVYIAEVVLALEYLHSLRVVHRDLKPDNLLIAHDGHIKLTDFGLSKVGLINSTDDLSGPA
AGGTYMLVDDEPELSTSDQQERRKKRSAVGTPDYLAPEILLGRGHGFTADWWSVGVILFELIVGIPPFNAEHPQIIFDNILNRKIPWPEVPDEMSPEARD
IIDRFLTEDPHMRLGARGASEVKQHVFFKNINWDTLARQKAAFVPASDSALDTSYFTSRYTRSREMNVEVSQSSILNLSQLASINYDLLSKGWKDDPPTS
S
|
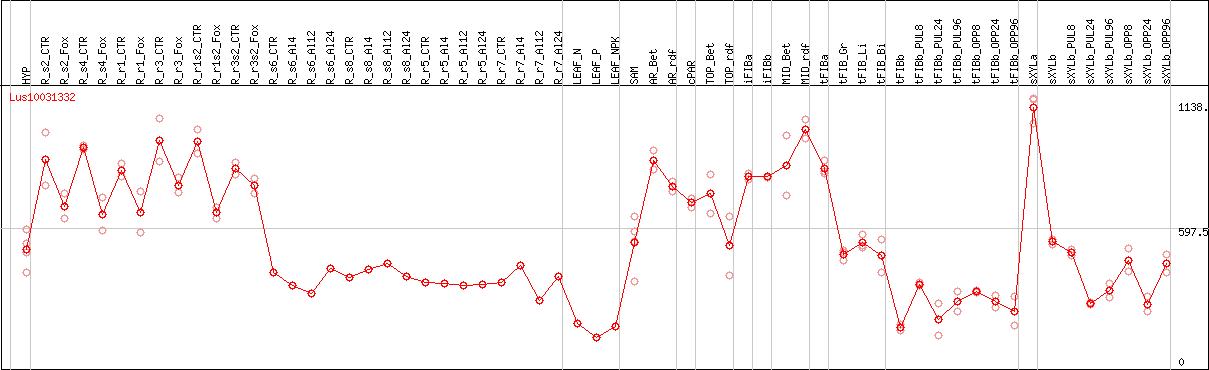
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031332 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
