Lus10031648 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10031648 pacid=23155799 polypeptide=Lus10031648 locus=Lus10031648.g ID=Lus10031648.BGIv1.0 annot-version=v1.0
ATGTCCGCCATCCGGTACCGAGCTCCTCCACCTGGCCGGCCACGGCCCAACCACCAGTCTCCCGGAGAAGATGAGGAGCCTTTCAATATAATCCCAGTCC
ACAACCTCTTAGCAGACCATCCATCCCTCCGCTATCCGGAGGTCCGCGCCGCTGCCGCCGCCTTACGAGCAGTAGGGAACCTCCGCCGTCCGCCCTACGT
CCAATGGCATCCTTCAATGGACCTCCTGGACTGGCTGGCCCTCTTCTTTGGCTTCCAGCGCGACAGCGTCCGCAACCAGAGGGAGCATCTAGTGCTGCAC
TTGGCTAACGCGCAGATGCGGCTCACTCCGCCGCCGGACAACATCGACACACTCGACCACGGGGTTCTTCGGCGGTTCAGACGGAAATTGCTGAAGAACT
ACACTAGCTGGTGCTCATACCTCAACAAAAAGTCCAACATCTGGATCTCGGATCGGTCGAACCCGGATGTGCGGCGCGAGCTACTGTATGTGTCTCTTTA
TCTTCTGATTTGGGGCGAATCGGCCAATTTGCGGTTTATGCCTGAGTGTATTTGCTTCATTTTTCACAACATGGCGATGGAATTGAACAAGATACTTGAG
GATTATATGGATGAGAACACTGGACAGCCTATTATGCCGTCGTTCTCCGGCGAGAACGCGTTTTTGAACTCTGTTGTGAAACCGATTTACGATACTGTGA
AGGCCGAGGTGGAGAATAGTAAGAATGGGACTGCGCCTCACACGTCGTGGAGGAACTACGATGATATCAATGAGTACTTTTGGAGCAGGAGGTGCTTCGA
GAAGCTGAAGTGGCCGATTGATTTGGGGAGCAATTTCTTTGTGATTGATAGTAGGCAGAAGCATGTGGGGAAGACGGGGTTTGTTGAGCAGAGGTCATTC
TTGAACTTGTTTCGAAGCTTTGACAGGTTGTTTATAATGTTGATCCTTTTCCTACAGGCTGCCATAATCGTGGCTTGGGCAGAAAAGGGGTATCCATGGC
AGGCACTGGAAAGTAGGGAGGTTCAGGTTCGAGTTTTGTCCGTTTTCTTCACTTGGGCTGGATTGAGGTTCATACAGTCGTTGCTTGATATCATAATGCA
GCGTAGGTTGGTTTCTCGAGAGACTATGTGGCTCGGGGTTAGGATGGTGGCGAAAGCTGTTGTTGCTACTGGTTGGATTCTTGTATTTTCGGTGTTCTAT
GCAAGGATTTGGAGCCAACGAGATCGCGATAGAGGATGGTCTGGTGCAGCGAATAGGAGGGTAGTGACTTTCCTTGAGGTGGCTGGGGTGTTTGTGCTGC
CTGAGATATTGGCAACTGTTCTGTTTATAATTCCTTGGGTTAGGAACTTCCTTGAGAATACCAATTGGAGGATCTTCTACTTGTTGTCGTGGTGGTTCCA
AAGCAGGAGCTTTGTTGGCCGTGGACTAAGGGAAGGGCTAGTTGACAATATCAAGTACAGTCTGTTCTGGATTCTGGTGTTGGCTACAAAGTTCAGTTTC
AGCTATTTTCTACAGATCAAACCTATGGTTAGGCCATCCAAACTAATGCTGGATCTGAAGGATGTCACCTATGAATGGCATGAGTTTTTTGACAACAGCA
ACAGATTTGCAGTGGGATTACTGTGGCTGCCTGTGGTGTTGATCTACCTTATGGATCTGCAGATTTGGTACTCTATCTATTCCTCTTTCGTAGGGGCGGC
GGTCGGTCTGTTTGATCATTTGGGTGAGATTCGGAACCTTCCTCAATTGAGGTTGAGGTTCCAGTTCTTTGCGAGTGCACTCCAGTTTAACTTGATGCCA
GAAGAGCAGTTGCTGAATGCTAGGGGTACTCTGAGGAGCAAATTCAAAGATTCGATTCATCGCTTAAAGTTGAGATATGGGCTTGGTCGGCCTTATAAGA
AGCTGGAATCCAATCAGGTTGAAGCAAACAAGTTTGCTATAGTTTGGAATGAGATTATAATGATTTTCAGGGAAGAGGATATCATCTCGGATCGAGAGGT
TGAGTTGCTGGAGTTGCCTCAGAATTCTTGGAATGTACAAGTTATTCGCTGGCCTTGCTTCCTTCTTTGCAATGAGTTGTTACTTGCTCTGAGCCAGGCA
AAAGAACTGGTGGATGCTCCTGACAAGTGGCTGTGGTACAAGATATGCAAGAATGAGTACAGGCGCTGTGCTGTTATAGAAGCTTATGACAGTATCAAGC
ACTTGCTACTTGAAATAATCAAGGTCAACACTGAAGAACATTCGATTGTGACTATTTTGTTCCAAGAAATTGACCACTCGCTGCAGATTGAAAAATTCAC
CAAGACTTTCAAGATGACTGCACTGCCTAATTTCCACGTGAAGCTGATAAAACTTCTTGACCTGTTGATGAAACCAACCAAGGATGTGAATCAGGTTGTG
AATACTCTCCAGGCTCTCTATGAGATTGCTATACGAGACTTTTTTAGAGAACAGAGAACTGTTGAACAGCTGAAGGAGGATGGATTAGTTCCTCATGATC
CTGCTGCCATGGCTGGGCATCTCTTTGAGAATGCAGTCGAACTTCCAGATTCTGCTCAGGAAACTTTGTACAGAAATGTAAGGCGCTTGCACACTATCCT
TACCTCCAGAGATTCAATGCACACTGTCCCAAAAAATCTGGAGGCCAGGCGTAGAATTGCCTTTTTCAGCAACTCCCTATTCATGAACATGCCGCATGCT
CCTCAAGTTGAGAAAATGATGGCCTTTAGTGTACTAACCCCATACTACAGTGAGGAAGTAATATACAGCAGAGAACAACTCCGAACTGAGAACGAGGATG
GGATTGCCACTCTGTACTACTTGCAAACTATCTATGATGATGAGTGGAAGAATTTTATTGAGAGGATGCGACGGGAAGGAATGTTGAATACTGATGAGAT
ATGGACGACCAAGTTGAGAGACCTCAGGCTCTGGGCATCTTACAGAGGACAAACACTTTCCCGTACTGTGAGAGGAATGATGTACTATTATCGGGCGCTG
AAGATGCTTGCTTTTCTTGATTCGGCCTCGGAGTTGGACATCAGGGAAGGTGCAAGGGAACTTGGTTCAATGCGGAGAGATTCGGGCCTGAATGGACAGG
GATCTGAAAGGTTTTCTTCTTCCCGACGCTTGAGCAGAAATAGCAGTTCGGTGAGTTTGTTGTTTAAGGGTCACGAGTATGGGACGGCTATTATGAAGTT
TACATATGTGGTTGCGTGTCAGATATATGGTACTCAGAAGGCTAAGAAGGATCCTCATGCGGAGGAAATTCTGTACTTGATGAAAAACAACGACGCTCTC
CGTGTAGCTTATGTTGATGAGAAAACTACAGGAAGAGATGTCACAGAATATTTCTCTGTGCTTGTCAAGTATGATGCTGAGTTGGAGAGAGAGGTGGAGA
TCTATAGGGTGAAGCTTCCTGGGCCTCTGAAGCTTGGAGAAGGAAAGCCGGAGAATCAGAGTCATGCTCTTATCTTCACCCGTGGCGATGCATTGCAGAC
CATTGACATGAACCAGGATAATTATTTCGAAGAGGCTCTCAAGATGCGGAATCTCTTGGAAGAATACAAGCAATATTATGGTATCAGGAAGCCTACCATC
TTGGGAGTCAGGGAACATATCTTTACTGGGTCTGTATCATCACTGGCGTGGTTCATGTCTGCTCAGGAGACAAGTTTTGTTACTTTGGGACAGCGCGTCT
TGGCGAATCCTCTGAAAGTCAGAATGCACTATGGCCACCCTGATGTGTTCGATAGATTCTGGTTCATGACTCGCGGTGGCCTCAGTAAAGCTTCCAGGGT
CATTAACATCAGTGAAGACATATTTGCTGGCTTCAATTGCACGTTGCGAGGTGGAAACATCACCCATCATGAATACATCCAGGTAGGGAAAGGAAGAGAT
GTCGGTCTTAATCAAATATCCATGTTCGAAGCCAAGGTTGCCAGTGGGAATGGTGAGCAAATTCTGAGCAGAGATGTCTACAGATTGGGACACAGGTTGG
ATTTCTTCAGGATGCTCTCATTCTTTTACACGACCGTGGGATTCTTTTTCAACACCATGATGGTGATCCTGACTGTGTATGCCTTTCTATGGGGACGACT
TTATCTGGCTCTCAGTGGTGTTGAGGGTTCTGCTTTGGCAGACGCCAGTAACAATAGAGCTCTCGGTGCCATTTTGAATCAGCAGTTTATAATCCAACTC
GGTCTATTCACTGCGCTCCCGATGATCGTGGAGAATTCTCTGGAGCAAGGATTTCTCCAAGCTATTTGGGATTTCCTGACAATGCAGCTCCAGCTGTCAT
CTGTGTTCTACACATTCTCCATGGGAACTCGCTCTCATTACTTCGGCCGCACGATTCTCCATGGCGGGGCAAAGTATCGAGCGACTGGACGAGGTTTCGT
CGTGCAGCACAAGAGCTTTGCCGAGAATTACCGACTCTATGCTCGTAGCCATTTTGTCAAGGCGATAGAACTCGGTTTGATCCTTATCGTTTACGCTACC
CACAGTCCTGTCGCCAAGGCTACGTTTGTGTACATAGCACTCACCATCTCCAGTTGGTTTCTGGTAATGTCGTGGATAATGGCTCCGTTTGTATTTAATC
CATCTGGATTCGACTGGTTGAAGACCGTGTACGACTTCGACGATTTCATGAACTGGATCTGGTACAGGGGAAGTGTGTTTGCAAAAGCTGAGCAGAGTTG
GGAAAAATGGTGGGAAGAGGAGCAGGATCATCTCAGAACCACCAGGCTCCTAGGCAAGTGCGTCGAGATCGTTTTGGACCTCAGGTTCTTCTTTTTCCAG
TTCGGGATCGTCTACCAAATGGGTATTGCAGCTAAAAGCACCAGCATTTTTGTCTATCTCCTCTCCTGGATATACATCTTCGTCGTCGTGGCTATATTCG
TGCTGATAGTGTACGCTCGCGAGAAATATGCTGCGAAGGAACACATCTACTACCGTCTTGTCCAGTTTCTTGTCATCGTGCTGGCCATACTCGTGATGAT
TGCACTGCTGGAGTTCACATCTCTCAGTTTTGCCGACATCTTCACCAGTATGCTGGCGTTTATCCCCACTGGCTGGGGAATGCTGTTGATCGCCCAGGTA
TTCCGGCCATGGCTGCAATCCACCATCCTCTGGGAGCTCGTCGTCTCGGTGGCTCGTTTATACGACATTATGTTTGGAGTGATCGTAATGACCCCTGTCG
CGTTCTTGTCGTGGATGCCTGGGTTCCAAGCCATGCAGACTAGGATACTGTTCAACGAAGCGTTCAGTAGAGGTCTCCGAATTATGCAGATCGTCACAGG
GAAGAAGTCTATGCTTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10031648 pacid=23155799 polypeptide=Lus10031648 locus=Lus10031648.g ID=Lus10031648.BGIv1.0 annot-version=v1.0
MSAIRYRAPPPGRPRPNHQSPGEDEEPFNIIPVHNLLADHPSLRYPEVRAAAAALRAVGNLRRPPYVQWHPSMDLLDWLALFFGFQRDSVRNQREHLVLH
LANAQMRLTPPPDNIDTLDHGVLRRFRRKLLKNYTSWCSYLNKKSNIWISDRSNPDVRRELLYVSLYLLIWGESANLRFMPECICFIFHNMAMELNKILE
DYMDENTGQPIMPSFSGENAFLNSVVKPIYDTVKAEVENSKNGTAPHTSWRNYDDINEYFWSRRCFEKLKWPIDLGSNFFVIDSRQKHVGKTGFVEQRSF
LNLFRSFDRLFIMLILFLQAAIIVAWAEKGYPWQALESREVQVRVLSVFFTWAGLRFIQSLLDIIMQRRLVSRETMWLGVRMVAKAVVATGWILVFSVFY
ARIWSQRDRDRGWSGAANRRVVTFLEVAGVFVLPEILATVLFIIPWVRNFLENTNWRIFYLLSWWFQSRSFVGRGLREGLVDNIKYSLFWILVLATKFSF
SYFLQIKPMVRPSKLMLDLKDVTYEWHEFFDNSNRFAVGLLWLPVVLIYLMDLQIWYSIYSSFVGAAVGLFDHLGEIRNLPQLRLRFQFFASALQFNLMP
EEQLLNARGTLRSKFKDSIHRLKLRYGLGRPYKKLESNQVEANKFAIVWNEIIMIFREEDIISDREVELLELPQNSWNVQVIRWPCFLLCNELLLALSQA
KELVDAPDKWLWYKICKNEYRRCAVIEAYDSIKHLLLEIIKVNTEEHSIVTILFQEIDHSLQIEKFTKTFKMTALPNFHVKLIKLLDLLMKPTKDVNQVV
NTLQALYEIAIRDFFREQRTVEQLKEDGLVPHDPAAMAGHLFENAVELPDSAQETLYRNVRRLHTILTSRDSMHTVPKNLEARRRIAFFSNSLFMNMPHA
PQVEKMMAFSVLTPYYSEEVIYSREQLRTENEDGIATLYYLQTIYDDEWKNFIERMRREGMLNTDEIWTTKLRDLRLWASYRGQTLSRTVRGMMYYYRAL
KMLAFLDSASELDIREGARELGSMRRDSGLNGQGSERFSSSRRLSRNSSSVSLLFKGHEYGTAIMKFTYVVACQIYGTQKAKKDPHAEEILYLMKNNDAL
RVAYVDEKTTGRDVTEYFSVLVKYDAELEREVEIYRVKLPGPLKLGEGKPENQSHALIFTRGDALQTIDMNQDNYFEEALKMRNLLEEYKQYYGIRKPTI
LGVREHIFTGSVSSLAWFMSAQETSFVTLGQRVLANPLKVRMHYGHPDVFDRFWFMTRGGLSKASRVINISEDIFAGFNCTLRGGNITHHEYIQVGKGRD
VGLNQISMFEAKVASGNGEQILSRDVYRLGHRLDFFRMLSFFYTTVGFFFNTMMVILTVYAFLWGRLYLALSGVEGSALADASNNRALGAILNQQFIIQL
GLFTALPMIVENSLEQGFLQAIWDFLTMQLQLSSVFYTFSMGTRSHYFGRTILHGGAKYRATGRGFVVQHKSFAENYRLYARSHFVKAIELGLILIVYAT
HSPVAKATFVYIALTISSWFLVMSWIMAPFVFNPSGFDWLKTVYDFDDFMNWIWYRGSVFAKAEQSWEKWWEEEQDHLRTTRLLGKCVEIVLDLRFFFFQ
FGIVYQMGIAAKSTSIFVYLLSWIYIFVVVAIFVLIVYAREKYAAKEHIYYRLVQFLVIVLAILVMIALLEFTSLSFADIFTSMLAFIPTGWGMLLIAQV
FRPWLQSTILWELVVSVARLYDIMFGVIVMTPVAFLSWMPGFQAMQTRILFNEAFSRGLRIMQIVTGKKSML
|
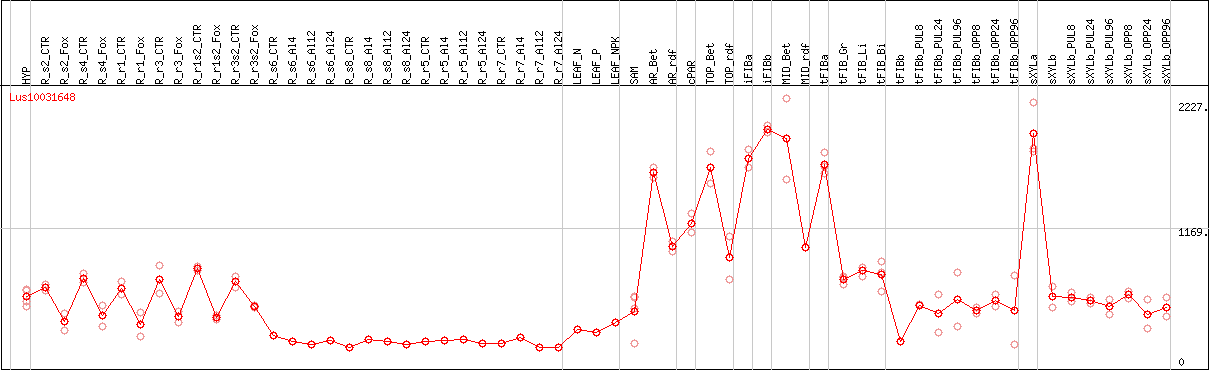
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031648 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
