External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G73320 356 / 4e-124
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
AT1G08125 135 / 1e-37
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
AT5G44170 75 / 6e-16
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT2G26810 63 / 2e-11
Putative methyltransferase family protein (.1.2.3)
AT5G49560 62 / 4e-11
Putative methyltransferase family protein (.1)
AT5G27400 54 / 5e-08
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT4G35987 53 / 5e-08
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
AT5G27410 54 / 7e-08
D-aminoacid aminotransferase-like PLP-dependent enzymes superfamily protein (.1.2)
AT3G50850 51 / 3e-07
Putative methyltransferase family protein (.1)
AT2G26200 49 / 2e-06
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021425
123 / 4e-33
AT1G08125 385 / 2e-135
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Lus10018640
65 / 4e-12
AT5G44170 348 / 1e-122
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10015166
65 / 7e-12
AT5G27400 377 / 9e-131
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10016141
63 / 3e-11
AT1G08125 183 / 7e-56
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Lus10031516
61 / 1e-10
AT5G27400 364 / 1e-125
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10039876
56 / 2e-09
AT5G44170 244 / 5e-83
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Lus10028415
55 / 1e-08
AT1G63855 248 / 2e-84
Putative methyltransferase family protein (.1.2.3)
Lus10041869
52 / 9e-08
AT1G63855 255 / 8e-87
Putative methyltransferase family protein (.1.2.3)
Lus10024993
50 / 1e-06
AT2G26200 726 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.017G153900
395 / 4e-140
AT1G73320 347 / 2e-120
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
Potri.009G006800
120 / 3e-32
AT1G08125 370 / 4e-129
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2.3)
Potri.017G017000
84 / 6e-19
AT5G44170 316 / 3e-110
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.001G148200
79 / 2e-17
AT5G44170 46 / 7e-06
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.009G068400
64 / 7e-12
AT2G26810 345 / 2e-121
Putative methyltransferase family protein (.1.2.3)
Potri.007G025600
62 / 5e-11
AT5G49560 289 / 2e-98
Putative methyltransferase family protein (.1)
Potri.005G109000
53 / 3e-08
AT1G63855 261 / 5e-89
Putative methyltransferase family protein (.1.2.3)
Potri.005G039000
51 / 4e-07
AT5G27400 361 / 6e-123
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.006G225600
49 / 2e-06
AT2G26200 739 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1)
Potri.017G031700
41 / 0.0008
AT2G43320 527 / 0.0
S-adenosyl-L-methionine-dependent methyltransferases superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF10294
Methyltransf_16
Lysine methyltransferase
Representative CDS sequence
>Lus10031894 pacid=23180618 polypeptide=Lus10031894 locus=Lus10031894.g ID=Lus10031894.BGIv1.0 annot-version=v1.0
ATGGCGAATCCGGCTACCGAAAGCTTAGAACAAGAAGAAGAGGAGGTGATGAATATGGGGAAGCTCGGGCTGTATGAAGCGAATGTGAGGTTGCTGAGTA
GCGAAGATTCGGAATCTGCCGCTGAGGAAATCATGCTGCTATGGGGAATCCAACAGCCCACCTTTTCGAAACCGAATGCTTATGTCTCCCAGTCCTCTCT
CCAGCTCCGCCTCGACGCTTGTGGTCACTCCCTCAACATTCTTCAGTCTCCTTCCTCCTTGGGTTCACCAGGAGTGACAGGATCAGTGATGTGGGACAGT
GGCATTGTACTAGGCAAGTTCTTGGAACACGCAGCTGATTCGAAGATGCTTGTTCTTCAAGGCAAGAAGGTTATCGAGCTTGGTTCGGGCTGTGGATTGG
TAGGATGCATAGCCTCTCTTCTGGGTGCAGAAGTGATACTCACTGATCTCCCCGACCGAATGCGGCTCCTGAGAAAGAACGTTGAGACCAATGTGGGCCA
CGAGCATCTACGTGGCTCTGCAGTCGTGGAGGAGCTCACTTGGGGAGACGACCCTGACCGGTCCATGATCGAACCCTTGCCCGATTACGTTGTGGGGTCC
GACGTGGTGTACAGCGAAGGAGCCGTGGGGGATCTGCTGGACACGCTCCTCAAACTCTGCGGACCAAAGACAACAGTGTTCATAGCTGGGGAGCTACGAA
ATGATGCAGTGCTTGAGTACTTCTTGGAGGAAGCCATGAAGGAGTTTATGGTAGGGAGAGTTGAGCAGTCCGAATGGCATCCGGATTACAACAGCCGCCG
GGTTGTGCTGTATATCATGGTCAGGAAGTAG
AA sequence
>Lus10031894 pacid=23180618 polypeptide=Lus10031894 locus=Lus10031894.g ID=Lus10031894.BGIv1.0 annot-version=v1.0
MANPATESLEQEEEEVMNMGKLGLYEANVRLLSSEDSESAAEEIMLLWGIQQPTFSKPNAYVSQSSLQLRLDACGHSLNILQSPSSLGSPGVTGSVMWDS
GIVLGKFLEHAADSKMLVLQGKKVIELGSGCGLVGCIASLLGAEVILTDLPDRMRLLRKNVETNVGHEHLRGSAVVEELTWGDDPDRSMIEPLPDYVVGS
DVVYSEGAVGDLLDTLLKLCGPKTTVFIAGELRNDAVLEYFLEEAMKEFMVGRVEQSEWHPDYNSRRVVLYIMVRK
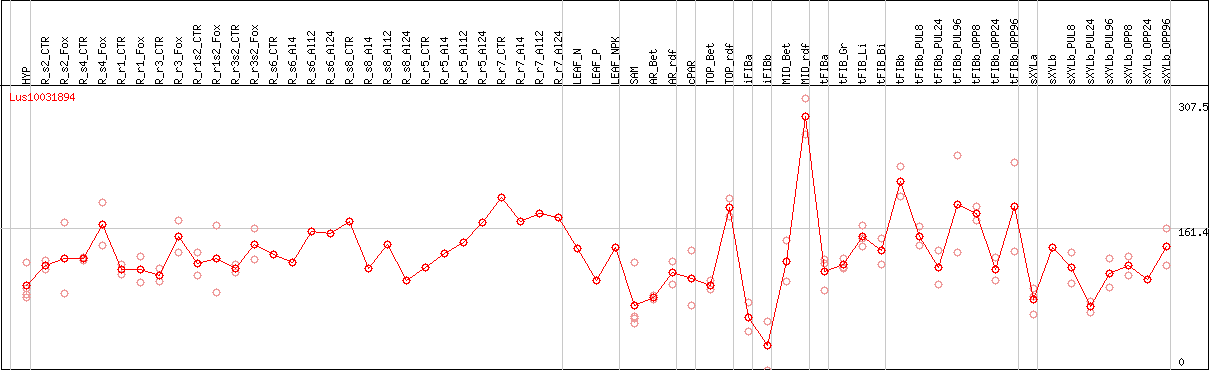
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031894 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.