Lus10031945 [FLAX]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Poplar homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Lus10031945 pacid=23161100 polypeptide=Lus10031945 locus=Lus10031945.g ID=Lus10031945.BGIv1.0 annot-version=v1.0
ATGTCTCTCCCTCCGTTGGAGTGTGTGCATATCACAGAAGACCATCTCCGTGAATGGAAAAATGGCAACGCTAACTATAGGGTTACTGAACCTGTACCCC
TGCTTCGTTTCCTCTATGAGCTCTGCTGGACGATGGTTCGGGGGGAATTGCCATTCCAAAAGTGCAAACTAGCTCTGGATTCGGTGGTGTTTCCTGATAC
AGTGTCAAAGGGCGAGTTGAGTTCTACCTTCGCTGATATAATTACCCAAATGGCATTAGATCTTACGATGCCTGGAGAGTACCGTGCACGTCTAATAAAG
CTGGCAAAGTGGTTAGTGGAATCTGCGCAGGTCCCATTGAGGCTGCTACAGGAGCGATGTGAGGAGGAGTTTCTGTTTGAGGGAGAGATGATAAAGATAA
AGGTTCAAGATTTGAAGGGAAAGGAGGTCAGAGTGAATACTCGTCTTCTTTACCAGCAAACAAAGTTCAATTTGCTTCGAGAAGAGAGTGAGGGTTATGC
CAAACTGATTACTCTTCTTTACCGAGGATCGGAAGATACAACTGCAAATGCATCAGCAGCTACAATAGGCATCATCAAGTCGTTAATTGGGCATTTTGAT
TTGGACCCAAACCGTGTTTTTGATGTTGTTTTAGAGTGTTTTGAACTGCAGCCAGACAGCAAGGTCTTTTTAGAATTAATTCCCATCTTCCCAAAGATTC
TTGGCTTTAAGTTTCAATATTATCAACGAATGGAGATAAACAATCTTGTACCCTTTGGACTCTATCAATTAACAGCTGTGCTGGTGATGAAGGACTTCAT
TGACCTAGACAGCATATACGCCCATATGCTTCCCAGGGATGATGATGCCATACAACATTACTGTGCATTTTCCACCAAGCAATTAGACGAGGCTAACAAG
ATTGGAAAAATAAATTTGGCTGCCACGGGGAAGGACCTTATGGATGAAGGGAGGCAGGGAGTTGAAACTATTGATCTCTTTACGGCTATAGACATGGAAA
GTGATGCGGTAGCTGAGCGGTCTGCTGAACTTGAGAAGAGTCAACCTCTGGGCTTACTTGGTGGTTTCCTTGCAGTAGATGACTGGTATCATGCTCAGAT
GTTGTTTGATCGCCTTTCTATCCTAAATCCCGTGGCACATGTTCAAATATGTAACGGTCTGTTTAGGCTCATAGAAAAGTCTATCTCTTCAGCATATGTC
GCTGCTCGTGAAGCACATGCAGCTCGTGCAGCTGGTAATGATGTTACGGCTCCAGTTTCTCTTCCGAAAGAACTCTTTCAGATGCTGACTGTTGTTGGAC
CTTACCTTTATCGCGATACAATATTACTTCAGAAGGTGTGCAGAGTATTAAAGGAATATTACCTGTCAGCTCGTAAGCTTGTGAATGATGGTGATGGAGC
TGCGAATCCTGATCCTTCATCTGGAAATCAAGCTTCGCGCCAGCTCCTAAGGGATGCTAGGTCAAGAATCGAGGAAGCTTTAGGATCATGTATACTTCCT
TCCCTACAGTTGATTCCTGCAAATCCAACAGTTGGACAAGAGATCTGGAAGGTTATGAGTCTCCTCCCATATGAGACACGTTATCGATTATACGGCGAAT
GGGAAAAGGATGATGAGAGGAATCCGATGGTTTTGGCTGCAAAGCAAACAGCAAAGTTGGATACTAGAAGAATACTGAAAAGACTTGCAAAGGAAAATCT
GAAACAGCTGGGTAGGATGGTTGCCAAACTTGCACATGCAAACCCCATGACCGTACTTCGAACTATTGTTCATCAGATTGAGGCATACAGAGATATGATT
ACTCCAGTTGTAGATGCTTTCAAGTATCTAACACAGGCATGCATTTCTTTCTTTTTCCTTTTACCAAGACCTAGGCTTGAGTATGACGTCTTGGAATATG
TGGTGATCGAGCGTTTGGCTCAGGGTGGTCGCGATAAGCTAAAAGATGATGGCCTTAATTTGTCGGACTGGCTCCAGTCTCTGGCTTCATTCTGGGGTCA
CCTGTGCAAGAAGTATCCATCAATGGAGCTAAGGGGTCTTTTTCAGTACCTTGTGAACCAATTGAAAAAGGGCCAAGGAATTGAGCTTCTTCTTCTCCAG
GAGCTTATTCAGCAAATGGCAAATGTCCAGTACACGGAGAACCTGACAGAGGAACAGTTGGATGCTATGGCAGGGAGTGAAACTCTTCGTTTTCAGGCTA
CTTCATTTGGAGTGCCACGGAATAATAAGGCATTAATTAAATCTACTGACAGGCTCCGAGAATCCTTGCGTCCAAAGGATGAACCAAAGCTAGCCATCCC
TCTGCTATTACTTATTGCTCAGCATCGTTCAGTAATTGTCATAAAGGCTGATGCGCCATATATTAAAATGGTTAGCGAGCAGTTCGACAGATGCCATGGG
ACTCTTCTTCAGTATGTGGAATTCCTCGGTAGCGCTATGACCTCAGCCACTACTTATGCAAAAATGATTCCTTCTCTGGATGATCTCGTTCACCAGTACC
ATCTTGATCCTGAGGTTGCTTTTCTGATATATCGTCCGGTAATGAGGCTCTTCAGGTGTGAGGGTAATTCTGAAGTTTTCTGGCCTTTAAAAGACAGTGA
TGGTAATGCCAGTACAGCAACTTCTAATATGGAGTCTGAACCAACAGGATATTCTGGAAATTTGATTCTTGATCTTGGGTCTCCTAAGAAGCCTATAATG
TGGTCGGATCTTCTTGAAACTGTGAAAACCATGTTGCCTGCAAGAGCTTGGAATAGCCTCTCTCCTGATCTTTACGCAACCTTCTGGGGGCTAACACTCT
ATGATCTTTATGTTCCGAGAAGTCGCTATGAGTCAGAAATATCAAAGCTTTATGCTTCTCTCAAAGCTTTGGAAGAACTGTCTGATAATTCAAGCTCAGG
TATCGCTAAAAGGAAAAAAGACAAGGAGCGGATTCAAGAATCTCTAGATCGTTTGACTAGTGAACTCCAAAAACATGAAGAGAACGTTGCATCTGTTCGG
CAGAGGCTTTCCCTTGAAAAAGACAAATGGCTGAGTTCTTGCCCTGATACCTTGAAAATTAATATGGAATTCCTTCAACGCTGTATTTTTCCGCGTTGCA
CGTTCAGCATGCCTGATGCTGTCTATTGTGCTGTGTTTGTGCACACTCTTCATTCCCTTGGTACACCATTCTTTAATACTGTAAATCACATTGATGTTCT
CATATGCAAAACATTGCAACCGATGATATGTTGTTGTACTGAGTATGAAGCTGGTAGGCTCGGGCGATTTTTGTACGAGACGCTAAAGATCGCATACCAT
TGGAAGAGGGAAGAATCTATATATGAACGTGAGTGTGGAAACATGCCTGGTTTCGCTGTTTACTATAGGTTTCCGAACAGCCAACGTGTCACCTATGGCC
AATTTATTAAGGTACACTGGAAATGGAGTCAGAGAATCTCACGGTTGTTGATACAATGCTTAGAATCATCTGAATACATGGAAATTCGGAATGCTCTCAT
TTTGTTAACAAAAATTTCGAGTGTTTTCCCAGTGACCAAAAGGAGTGGGATTAACATTGAAAAGCGAGTATCAAGAATAAGAAACGATGACAGAGAGGAT
CTGAAGGTGCTGGCAACTGGCGTTGCTGCTGCTTTGGCCTCTAGAAAGCCATCATGGGTTTCAGACGAAGAGTTTGGCATGGGATATCTTGAATTAAAGC
CACCATCAAAATCGATATCTGGAACTGCAGCCCCTATACAAAGTAGTTCTGTAGTTAATCTTTCTCAAAATGAACCTGGTAGTTCAAAAGCGCAGACCAC
TACAACACGCTATGGAGATTCTGGAAATTTATCTCGGGATCAGATAAGTAGAGGAAATGGGAAGCTAGACAAAATGGAAAGTGCCTCGCATTCTAAGTCT
GACATTGGGAATAAAAAATTGAAGGGTAGTTCTTCTGCCAATGGGCCTGATGGTCAGTTATCTGCACTACCAGTTGGCACTCAAGATGGGCCATCAAGAT
CATCTTCAAATCAGAAATACACGGAAGAGTCGGGTAATAGAGAACTGGATGAAACGAGTACTCGGGTCACTCAGAGGAGCTCTGCGGAATCAGAGCCAAA
AGGTAATGCAAAACGGGCAATGGCTTCTGGATCCGTTGTTAAGACGCTCAAGCAAGATTCCATTAAGGATGACGGAAAAAGTGGCAAAGCTGTCGGCAGA
ACTTCTGCCAGTGATAGAGATGTTTCTGGTTTAGTTGTTCTCACTTCAGCAGTCGCATCTAATGGCAATGCAGTTCCTACATCTTCAAAATCAACTCAGT
CTGCTAGATTATCTGATGCTCATCATAATGAGTTGAAATCAGATGGTGGGGCTTCTAAATCTGTGCTCAAAGACAATGCATCTGAAACTGCTGATGTGCA
GAAGCTTTCTTCTAGGCTTGTTCATTCTCCTAGGAATGAAGCCTCTATTTCTACTTCAAGGTCTAGTGATAAATTGCTAAAGCGAGCAAGTCCTTCGGAA
GAACCTGACAGATTAAGTAAACGCCGGAGAGGTGACACCGACTCCAAAGATTTAGAAAGTGAAGCTCGGTTTTCTGATCGAGACAAATCTATTGATACTC
GACCTTTGGATCTTGATAAGATGGGCAATGATGAACAGACTCTGTATAGGACGAGTGATAAACCCTCAGAAAGGTCCAAAGATAAGCTTGGTGATAGACA
CGAAAGAGATTATCGCGAAAGATCAGACCGCCTTGACAAGTCTGTCAGAGATGATAGTCATAGTTTAGCTGAAAGACAGAGAGATAGGTCAATTGAAAGA
TATCGAGAGAGATCAGTTGATAGGGGACAAGAAAGAGGTACTGATAGAGGCCATGACAGACTATCAGAGAAAGCAAAAGATGACCGGAGCAAAGATGATA
GGAATAAATTGCGGTATAGCGATTCGTCTGCAGACAAATCTCACGATGATCGTTTTCGTGGTCAAAGTTTGCCTCCACCGCCACCCTTACCTCCTCACGT
AGTTCCTCAATCGGTTGCTGCAAGTAGAAAGGACGACGATGCTGATAGAAGGCTTGTAACTACCAGACATGGCCAAAGGCTTTCTCCGAGAATTGATGAG
AAAGAACGAAGACGGTCGGAAGAAAATTCTACTGTCTCACCGGATGATGCAAAGCGTAGAAGAGAAGACGATTTTCGGGAGAGAAAGCGAGAGGACCGTG
AGGTGTCTACGGTTAAGGCGGAAGAGAGAGAAAGGGAAAGGGAAAGAGAGAAGGCGAATCTATTGAAGGAGGAGATGGATGCAGCTGCAACGTCCAAGAG
GCGAAAGCTTAAAAGAGAGCATTTGCCCCCAGGAGGAGTCGAAGCTGGTGAATATTCTCCTGTGGCGGCCCAACCACCTTCCCTTGCAATGGGAATGTCT
CAATCGTATGACCTAAGAGAGAGGGGAGATCGCAAGGCCTCCATGGTCCAACGTACAGGATATGTCGAGGAACCTCCTGCCAGAGCTCATGGTAAAGAAG
TGGCCAGCAAGGCTGCCCGGCGCGATGTCGATCCAATGTATGAACGAGATTGGGACGACGAAAAGAGGCAAAGAGCTGAACAGAAGAGGAGGCATCGAAA
GTAA
|
|||||||||||||||
|
AA sequence
|
>Lus10031945 pacid=23161100 polypeptide=Lus10031945 locus=Lus10031945.g ID=Lus10031945.BGIv1.0 annot-version=v1.0
MSLPPLECVHITEDHLREWKNGNANYRVTEPVPLLRFLYELCWTMVRGELPFQKCKLALDSVVFPDTVSKGELSSTFADIITQMALDLTMPGEYRARLIK
LAKWLVESAQVPLRLLQERCEEEFLFEGEMIKIKVQDLKGKEVRVNTRLLYQQTKFNLLREESEGYAKLITLLYRGSEDTTANASAATIGIIKSLIGHFD
LDPNRVFDVVLECFELQPDSKVFLELIPIFPKILGFKFQYYQRMEINNLVPFGLYQLTAVLVMKDFIDLDSIYAHMLPRDDDAIQHYCAFSTKQLDEANK
IGKINLAATGKDLMDEGRQGVETIDLFTAIDMESDAVAERSAELEKSQPLGLLGGFLAVDDWYHAQMLFDRLSILNPVAHVQICNGLFRLIEKSISSAYV
AAREAHAARAAGNDVTAPVSLPKELFQMLTVVGPYLYRDTILLQKVCRVLKEYYLSARKLVNDGDGAANPDPSSGNQASRQLLRDARSRIEEALGSCILP
SLQLIPANPTVGQEIWKVMSLLPYETRYRLYGEWEKDDERNPMVLAAKQTAKLDTRRILKRLAKENLKQLGRMVAKLAHANPMTVLRTIVHQIEAYRDMI
TPVVDAFKYLTQACISFFFLLPRPRLEYDVLEYVVIERLAQGGRDKLKDDGLNLSDWLQSLASFWGHLCKKYPSMELRGLFQYLVNQLKKGQGIELLLLQ
ELIQQMANVQYTENLTEEQLDAMAGSETLRFQATSFGVPRNNKALIKSTDRLRESLRPKDEPKLAIPLLLLIAQHRSVIVIKADAPYIKMVSEQFDRCHG
TLLQYVEFLGSAMTSATTYAKMIPSLDDLVHQYHLDPEVAFLIYRPVMRLFRCEGNSEVFWPLKDSDGNASTATSNMESEPTGYSGNLILDLGSPKKPIM
WSDLLETVKTMLPARAWNSLSPDLYATFWGLTLYDLYVPRSRYESEISKLYASLKALEELSDNSSSGIAKRKKDKERIQESLDRLTSELQKHEENVASVR
QRLSLEKDKWLSSCPDTLKINMEFLQRCIFPRCTFSMPDAVYCAVFVHTLHSLGTPFFNTVNHIDVLICKTLQPMICCCTEYEAGRLGRFLYETLKIAYH
WKREESIYERECGNMPGFAVYYRFPNSQRVTYGQFIKVHWKWSQRISRLLIQCLESSEYMEIRNALILLTKISSVFPVTKRSGINIEKRVSRIRNDDRED
LKVLATGVAAALASRKPSWVSDEEFGMGYLELKPPSKSISGTAAPIQSSSVVNLSQNEPGSSKAQTTTTRYGDSGNLSRDQISRGNGKLDKMESASHSKS
DIGNKKLKGSSSANGPDGQLSALPVGTQDGPSRSSSNQKYTEESGNRELDETSTRVTQRSSAESEPKGNAKRAMASGSVVKTLKQDSIKDDGKSGKAVGR
TSASDRDVSGLVVLTSAVASNGNAVPTSSKSTQSARLSDAHHNELKSDGGASKSVLKDNASETADVQKLSSRLVHSPRNEASISTSRSSDKLLKRASPSE
EPDRLSKRRRGDTDSKDLESEARFSDRDKSIDTRPLDLDKMGNDEQTLYRTSDKPSERSKDKLGDRHERDYRERSDRLDKSVRDDSHSLAERQRDRSIER
YRERSVDRGQERGTDRGHDRLSEKAKDDRSKDDRNKLRYSDSSADKSHDDRFRGQSLPPPPPLPPHVVPQSVAASRKDDDADRRLVTTRHGQRLSPRIDE
KERRRSEENSTVSPDDAKRRREDDFRERKREDREVSTVKAEEREREREREKANLLKEEMDAAATSKRRKLKREHLPPGGVEAGEYSPVAAQPPSLAMGMS
QSYDLRERGDRKASMVQRTGYVEEPPARAHGKEVASKAARRDVDPMYERDWDDEKRQRAEQKRRHRK
|
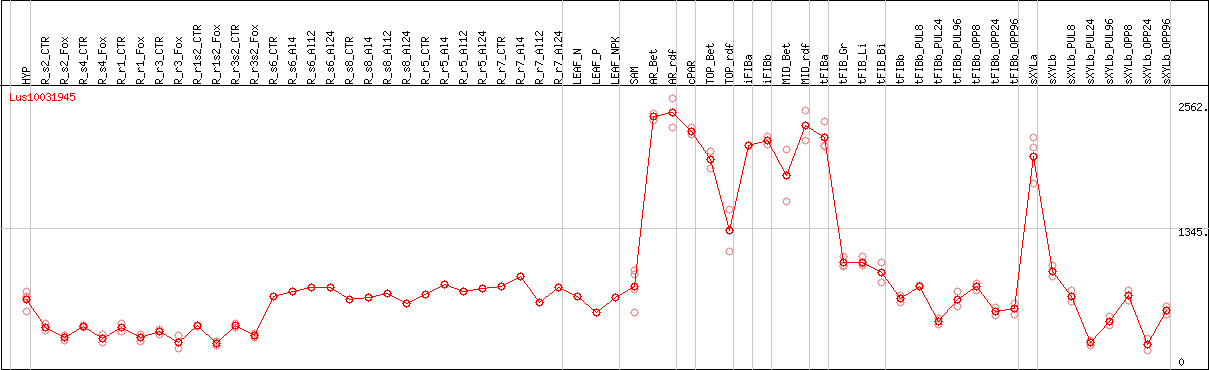
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10031945 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
