External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G48560 94 / 2e-22
TZP5, IMR1, ALS, AHAS, CSR1
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035359
97 / 4e-26
AT3G48560 219 / 3e-69
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Lus10032041
97 / 5e-24
AT3G48560 565 / 0.0
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Lus10029955
97 / 1e-23
AT3G48560 981 / 0.0
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Lus10035207
95 / 6e-23
AT3G48560 800 / 0.0
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Lus10022446
94 / 7e-23
AT3G48560 702 / 0.0
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Lus10016751
94 / 2e-22
AT3G48560 1048 / 0.0
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Lus10035206
45 / 1e-06
AT5G50460 127 / 1e-40
secE/sec61-gamma protein transport protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G098300
95 / 7e-23
AT3G48560 1052 / 0.0
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
Potri.015G097200
95 / 7e-23
AT3G48560 1058 / 0.0
TRIAZOLOPYRIMIDINE RESISTANT 5, IMIDAZOLE RESISTANT 1, ACETOLACTATE SYNTHASE, ACETOHYDROXY ACID SYNTHASE, chlorsulfuron/imidazolinone resistant 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0254
THDP-binding
PF02775
TPP_enzyme_C
Thiamine pyrophosphate enzyme, C-terminal TPP binding domain
Representative CDS sequence
>Lus10032037 pacid=23161253 polypeptide=Lus10032037 locus=Lus10032037.g ID=Lus10032037.BGIv1.0 annot-version=v1.0
ATGAATGTTCAAGAGCTCGCAACAGTTCGAGTGGAGAACTTGCCAGTGAAGATCATGATACTCAACAATCAGCAGTTGGGGATGGTGGTTCAGTGGGAAG
ATAAGTTTTACAAGGCCAACAGAGCACATACTTTCCTCGGCACAGAGTTTCTTTCGGGATGGATGCTGCTGAAAGAATCACCAGAAAACTATTATCCCGA
GTCCCCCCTGCATCCCTACGATGATATTGGAGCTTACCTTGCAAATCTAACCAGATCCTACGATGATGTTATTGATGGGGATGAAGATGAGCTTGACGAA
GAACCCTACGAATCCCATCACCACGAAACCTATCGCCGTACGCACTGCCACCTTCGAAAACTCTGCCAGATCACCAAATCAATGAGCACACTGAGTCAAC
ACACAGATCGGCGGAGTTTCCCCAAAATCGCATCGGAAGTTTCAGAAACGAAGAATCAGCGAAAGGAAACAGAGATTTACCTTTCCGATCTGGCTTGTGG
CAGCGCTTCACAAGTCTTACACTGTCCTTGGCGAATTCCCTGA
AA sequence
>Lus10032037 pacid=23161253 polypeptide=Lus10032037 locus=Lus10032037.g ID=Lus10032037.BGIv1.0 annot-version=v1.0
MNVQELATVRVENLPVKIMILNNQQLGMVVQWEDKFYKANRAHTFLGTEFLSGWMLLKESPENYYPESPLHPYDDIGAYLANLTRSYDDVIDGDEDELDE
EPYESHHHETYRRTHCHLRKLCQITKSMSTLSQHTDRRSFPKIASEVSETKNQRKETEIYLSDLACGSASQVLHCPWRIP
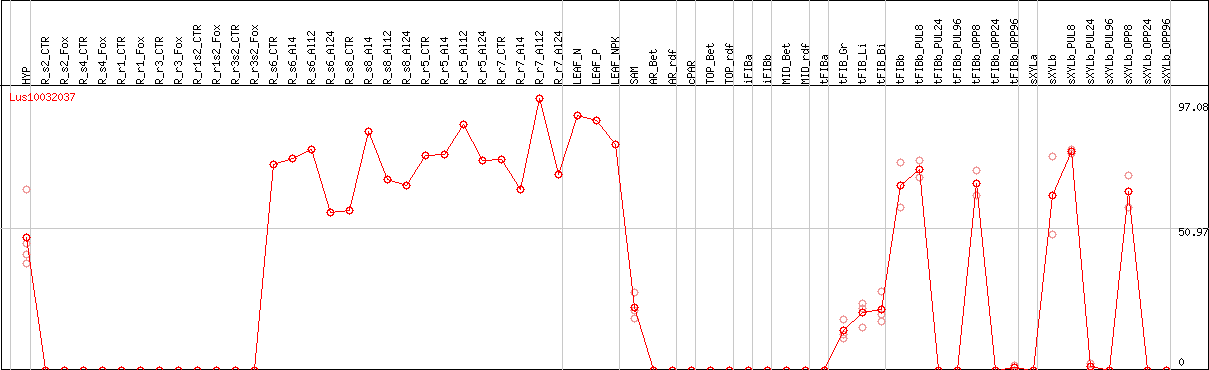
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032037 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.