External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G15050 231 / 7e-75
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT5G39990 198 / 7e-62
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT4G27480 166 / 1e-49
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT1G53100 165 / 4e-49
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT3G15350 162 / 3e-48
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT4G03340 152 / 4e-44
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G03520 147 / 6e-43
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT2G37585 141 / 2e-40
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT3G03690 138 / 3e-39
UNE7
unfertilized embryo sac 7, Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G71070 113 / 9e-30
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032189
346 / 1e-119
AT5G15050 648 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10014495
342 / 7e-118
AT5G15050 649 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10043324
173 / 4e-52
AT3G15350 641 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Lus10019477
172 / 6e-52
AT3G15350 639 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Lus10030084
160 / 3e-47
AT5G15050 507 / 4e-179
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10001076
158 / 2e-46
AT5G15050 512 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10031626
154 / 1e-44
AT4G03340 659 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10033714
153 / 1e-44
AT4G03340 660 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10020316
151 / 8e-44
AT3G03690 457 / 1e-160
unfertilized embryo sac 7, Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G130300
255 / 3e-84
AT5G15050 654 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.017G075600
236 / 2e-76
AT5G15050 671 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.001G399500
176 / 2e-53
AT3G15350 622 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Potri.011G119600
170 / 5e-51
AT3G15350 614 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Potri.009G003300
158 / 2e-46
AT5G39990 447 / 1e-155
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.019G104600
152 / 5e-44
AT4G03340 639 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.013G145500
152 / 6e-44
AT1G03520 667 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
Potri.013G066200
142 / 1e-40
AT3G03690 486 / 2e-172
unfertilized embryo sac 7, Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.006G263000
139 / 6e-39
AT2G37585 500 / 7e-177
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.008G128000
115 / 2e-30
AT1G71070 574 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF02485
Branch
Core-2/I-Branching enzyme
Representative CDS sequence
>Lus10032190 pacid=23161279 polypeptide=Lus10032190 locus=Lus10032190.g ID=Lus10032190.BGIv1.0 annot-version=v1.0
ATGAAGCGGCTGAAGAACTACTACATGCATTTCCGGCACCACCACCAGGCGGCGGATAGGCGGCGCTGGATCTTCCCTCTTGCAATTGGCTCAATATTCT
CTCTCTTTCTCCTCTTCCTAACTACTACCGTCTATTCCCCTGCCGGTGTCCCAATCTTCCCCTTCTACCGCACAATGTACTCATCCTTCAGCTCCAAATT
CGTCGAGACCAAGCTGCAGCCCATCCCGCTCTCCGATCTCCCTCCGCCGCCGCGATTCGCCTACTTGATCTCCGGCTCCAAGGGCGACGGCGACATGCTC
AAGCGAACCCTACTTGCTCTCTACCACCCTAATAACCGCTACGTGGTGCATTTGGACGCCGAGTCCTCCCCTGACGAGCGGACGGATCTGGCCGATTTCG
TGAGGGACCATAGGGTTTTTGTCAAGTTCGGGAATGTGAAGATGATTGCCAAGGCTAACCTCGTCACTTACAGAGGTCCCACCATGGTTGCCAATACGCT
TCACGCTGCGGCGATCTTGCTGAGGGAAGGATCCGATTGGGATTGGTTCATTAACCTCAGTGCTTCCGACTACCCGCTCGTGACTCAGGACGGTGAGATT
TTTTAA
AA sequence
>Lus10032190 pacid=23161279 polypeptide=Lus10032190 locus=Lus10032190.g ID=Lus10032190.BGIv1.0 annot-version=v1.0
MKRLKNYYMHFRHHHQAADRRRWIFPLAIGSIFSLFLLFLTTTVYSPAGVPIFPFYRTMYSSFSSKFVETKLQPIPLSDLPPPPRFAYLISGSKGDGDML
KRTLLALYHPNNRYVVHLDAESSPDERTDLADFVRDHRVFVKFGNVKMIAKANLVTYRGPTMVANTLHAAAILLREGSDWDWFINLSASDYPLVTQDGEI
F
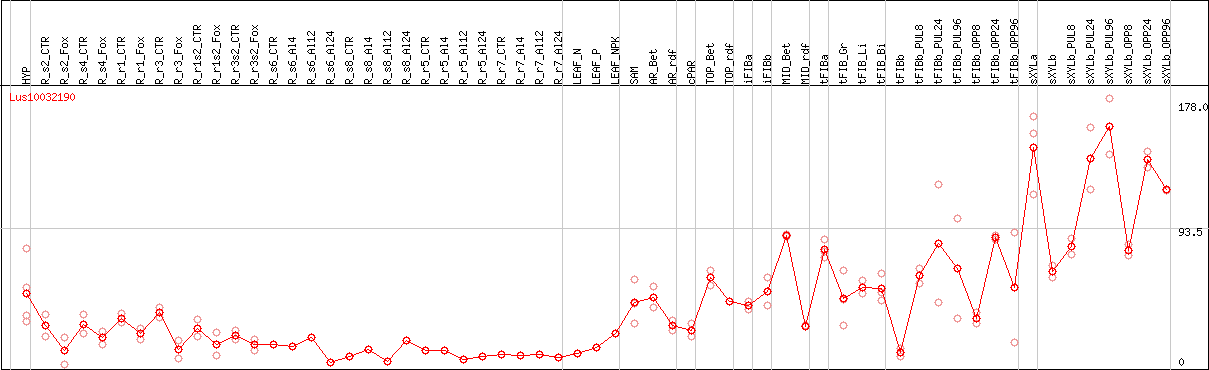
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032190 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.