External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G14190 287 / 3e-94
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
AT1G14185 287 / 3e-94
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
AT1G72970 157 / 1e-43
HTH, EDA17
HOTHEAD, embryo sac development arrest 17, Glucose-methanol-choline (GMC) oxidoreductase family protein (.1), Glucose-methanol-choline (GMC) oxidoreductase family protein (.2)
AT3G56060 155 / 3e-43
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
AT5G51950 155 / 5e-43
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1), Glucose-methanol-choline (GMC) oxidoreductase family protein (.2)
AT1G73050 147 / 3e-40
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
AT1G12570 144 / 4e-39
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
AT5G51930 136 / 3e-36
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026606
471 / 2e-166
AT1G14190 554 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10030460
453 / 8e-159
AT1G14185 555 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10012816
449 / 1e-157
AT1G14190 539 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10030459
425 / 1e-147
AT1G14185 605 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10026607
405 / 9e-140
AT1G14185 605 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10012817
404 / 2e-139
AT1G14185 545 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10030457
403 / 5e-139
AT1G14185 589 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10024728
391 / 3e-136
AT1G14185 391 / 2e-133
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Lus10030458
383 / 9e-132
AT1G14185 481 / 5e-167
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G168100
352 / 4e-119
AT1G14185 654 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Potri.008G087400
339 / 4e-114
AT1G14185 639 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Potri.010G168200
339 / 9e-114
AT1G14190 598 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Potri.003G039000
149 / 1e-40
AT1G72970 808 / 0.0
HOTHEAD, embryo sac development arrest 17, Glucose-methanol-choline (GMC) oxidoreductase family protein (.1), Glucose-methanol-choline (GMC) oxidoreductase family protein (.2)
Potri.001G200800
147 / 4e-40
AT1G72970 831 / 0.0
HOTHEAD, embryo sac development arrest 17, Glucose-methanol-choline (GMC) oxidoreductase family protein (.1), Glucose-methanol-choline (GMC) oxidoreductase family protein (.2)
Potri.008G009600
146 / 9e-40
AT1G73050 785 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Potri.001G111500
144 / 6e-39
AT1G12570 773 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
Potri.012G135200
65 / 1e-11
AT5G51950 525 / 0.0
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1), Glucose-methanol-choline (GMC) oxidoreductase family protein (.2)
Potri.010G249501
60 / 4e-11
AT1G73050 164 / 7e-49
Glucose-methanol-choline (GMC) oxidoreductase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05199
GMC_oxred_C
GMC oxidoreductase
CL0063
NADP_Rossmann
PF00732
GMC_oxred_N
GMC oxidoreductase
Representative CDS sequence
>Lus10032347 pacid=23168638 polypeptide=Lus10032347 locus=Lus10032347.g ID=Lus10032347.BGIv1.0 annot-version=v1.0
ATGGGAAGACATGAGAACATCATAGTTCTGTTGAATGCTACGGTCAAGAGTGTCATATTCAACAACGGAAAATCGGAAAAGACCACTGCTCGAGGCATTC
GATTCATAAGCAGTCACGGTACCGGAGATCAAACCTATGAAGCTTACCTTAAGCAACCAGCAAATCGAACTTGTTCGTCCGGAGAGGTGATATTATCAGC
AGGAGCATTAGGCAGCCCTCAGATCCTCCTACTGAGCGGCATCGGACCAAAACCCCATCTCAACCGACTCAATATCCCACTCGTCCTCGAGTCAAACCAC
GTGGGACTCGAAGTACAAGATAACCCAGGAATTTCCATCTTGGTCGGTTCGAAACCTGGACAGAAGCTCCCAGACCCACCACAAGTAGCAGGGATAACAA
AAGGATTCAACTTCATCATCCAAGGGGTCGTCGGACCAGTAAGCTTCAACGCTACAAGGAACAGAATATCAATCAAGCTAGCATTCCCAGAATCGAAAGG
GAAGTTGGAACTGAACTCAACCGATCCCAGGATGAACCCTGAAGTTTCATACAAATACCTGGACACTGAGAGAGACATGGATGGATGTGTGGAGATGGTC
AAGTTGGCAATGACTGTTGCTAAGTATCTATCTCGGGAGACGAATTTGGTGTCGAATCCTGGTGAGCTGCGGACATACTGCAGGAAGAATGTCGTGAGTT
ACTTCCATTACCATGGTGGGTGTGTTGTTGGATCCGTTGTTGATGAAGATTATAGAGTGGTCGGTGTCGATGGTTTAAGAGTTGTGGATGGTTCGACGTT
CTTCGAAGCGCCTGGGACGAACCCGATGGCTACTGTCCTGATGCTTGGCAGGTATCAAGGGATCAAGATTCTTGCTAGGAGGAATAAGTGTGAAAAATAA
AA sequence
>Lus10032347 pacid=23168638 polypeptide=Lus10032347 locus=Lus10032347.g ID=Lus10032347.BGIv1.0 annot-version=v1.0
MGRHENIIVLLNATVKSVIFNNGKSEKTTARGIRFISSHGTGDQTYEAYLKQPANRTCSSGEVILSAGALGSPQILLLSGIGPKPHLNRLNIPLVLESNH
VGLEVQDNPGISILVGSKPGQKLPDPPQVAGITKGFNFIIQGVVGPVSFNATRNRISIKLAFPESKGKLELNSTDPRMNPEVSYKYLDTERDMDGCVEMV
KLAMTVAKYLSRETNLVSNPGELRTYCRKNVVSYFHYHGGCVVGSVVDEDYRVVGVDGLRVVDGSTFFEAPGTNPMATVLMLGRYQGIKILARRNKCEK
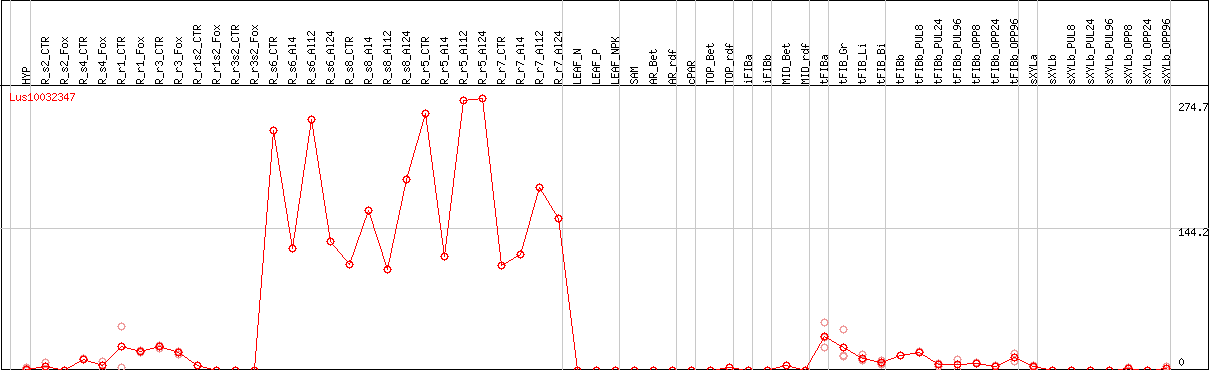
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032347 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.