External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G23980 154 / 9e-43
ARF
ARF9
auxin response factor 9 (.1.2)
AT2G46530 134 / 5e-36
ARF
ARF11
auxin response factor 11 (.1.2.3)
AT3G61830 133 / 1e-35
ARF
ARF18
auxin response factor 18 (.1)
AT1G35540 122 / 2e-31
ARF
ARF14
auxin response factor 14 (.1)
AT1G34170 117 / 7e-30
ARF
ARF13
AUXIN RESPONSE FACTOR 13 (.1.2.3)
AT1G34410 115 / 4e-29
ARF
ARF21
auxin response factor 21 (.1)
AT1G34310 113 / 2e-28
ARF
ARF12
auxin response factor 12 (.1)
AT1G35240 112 / 5e-28
ARF
ARF20
auxin response factor 20 (.1)
AT1G34390 111 / 8e-28
ARF
ARF22
auxin response factor 22 (.1)
AT1G35520 111 / 1e-27
ARF
ARF15
auxin response factor 15 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023058
435 / 2e-150
AT4G23980 743 / 0.0
auxin response factor 9 (.1.2)
Lus10024455
119 / 2e-30
AT1G59750 905 / 0.0
auxin response factor 1 (.1.2.3.4)
Lus10007440
117 / 2e-29
AT1G59750 917 / 0.0
auxin response factor 1 (.1.2.3.4)
Lus10043200
107 / 3e-26
AT5G62000 1091 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Lus10032541
107 / 4e-26
AT5G62000 1095 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Lus10022943
87 / 5e-19
AT1G19220 546 / 4e-177
indole-3-acetic acid inducible 22, AUXIN RESPONSE FACTOR11, auxin response factor 19 (.1)
Lus10027600
86 / 6e-19
AT1G19220 670 / 0.0
indole-3-acetic acid inducible 22, AUXIN RESPONSE FACTOR11, auxin response factor 19 (.1)
Lus10031354
82 / 3e-17
AT5G37020 1013 / 0.0
auxin response factor 8 (.1.2)
Lus10012421
77 / 9e-16
AT1G19850 1026 / 0.0
MONOPTEROS, indole-3-acetic acid inducible 24, AUXIN RESPONSE FACTOR 5, Transcriptional factor B3 family protein / auxin-responsive factor AUX/IAA-related (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G088600
267 / 7e-85
AT4G23980 710 / 0.0
auxin response factor 9 (.1.2)
Potri.003G142100
243 / 8e-76
AT4G23980 706 / 0.0
auxin response factor 9 (.1.2)
Potri.014G100100
141 / 4e-38
AT4G23980 694 / 0.0
auxin response factor 9 (.1.2)
Potri.002G172800
135 / 5e-36
AT4G23980 662 / 0.0
auxin response factor 9 (.1.2)
Potri.004G228800
114 / 8e-29
AT1G59750 906 / 0.0
auxin response factor 1 (.1.2.3.4)
Potri.003G001000
111 / 1e-27
AT1G59750 898 / 0.0
auxin response factor 1 (.1.2.3.4)
Potri.001G066200
104 / 4e-25
AT5G62000 765 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Potri.012G106100
103 / 7e-25
AT5G62000 1035 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Potri.003G163600
100 / 7e-24
AT5G62000 691 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Potri.015G105300
100 / 1e-23
AT5G62000 1065 / 0.0
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0072
Ubiquitin
PF02309
AUX_IAA
AUX/IAA family
Representative CDS sequence
>Lus10032414 pacid=23168533 polypeptide=Lus10032414 locus=Lus10032414.g ID=Lus10032414.BGIv1.0 annot-version=v1.0
ATGATGTGGCAACATAAACAGGCTGATATTATCAGCCATGGAAGCTCTATTTCGAGAACTCAGACTGATGCAGGCTGGCTGTCTCCGCACATGAATGTCA
ATCTGCATAACTTCCAGGACACCACTGAGGATACCAAGAGCGTCTCGAATTGGTCAGTTTCAAACTTTGCATCTCCCCAAGCTGCAAAACCGAAGAGTGA
AGTTGCCAGTAGCTACAGGTTGTTTGGTATTGAGCTCATGAATAGTCCGATGAGTGCACCGATGACTGAGAAGACATTATCTAGTGGCACAACAGAAGTA
CATGTACATAGTACCATCTCAGCTGCTGATCCTGACCAAAAAGCAGACTCTCCCAAGGAAAAGAAAACCGACCAATCTCAAGTGTCGCCAAAGGATACCC
AGAGCAGGCAAAGTTCTGTATCGACCAGAAGCCGCACCAAGGGAGTTGCTGTTGGAAGAGCGGTAGACCTGACGACAATAAAAGGGTATCCTCAGCTGAT
AAGTGAGCTGGAGAGTTTGTTTGATATCGAGGGACAGCTTCAGTTTCGCGACAAGTGGGAAATCGTCTACACTGACGACGAAGGAGACATGATGCTCGTC
GGAGATGATCCATGGGTTGAATTCTGCAACATGGTGAGGAGAATCATCATATGTTCGAGTCAGGATGTGAAAAGGATGAGGAATCCAGGGAGCAAGAATC
CCATGTTTGGAGTAGGAGGAGATGGAGGAGGAGCAGCTGTAGCAGTAAGCTCTGATTCAGCAGTGGAAAACTCCAAACAGTAG
AA sequence
>Lus10032414 pacid=23168533 polypeptide=Lus10032414 locus=Lus10032414.g ID=Lus10032414.BGIv1.0 annot-version=v1.0
MMWQHKQADIISHGSSISRTQTDAGWLSPHMNVNLHNFQDTTEDTKSVSNWSVSNFASPQAAKPKSEVASSYRLFGIELMNSPMSAPMTEKTLSSGTTEV
HVHSTISAADPDQKADSPKEKKTDQSQVSPKDTQSRQSSVSTRSRTKGVAVGRAVDLTTIKGYPQLISELESLFDIEGQLQFRDKWEIVYTDDEGDMMLV
GDDPWVEFCNMVRRIIICSSQDVKRMRNPGSKNPMFGVGGDGGGAAVAVSSDSAVENSKQ
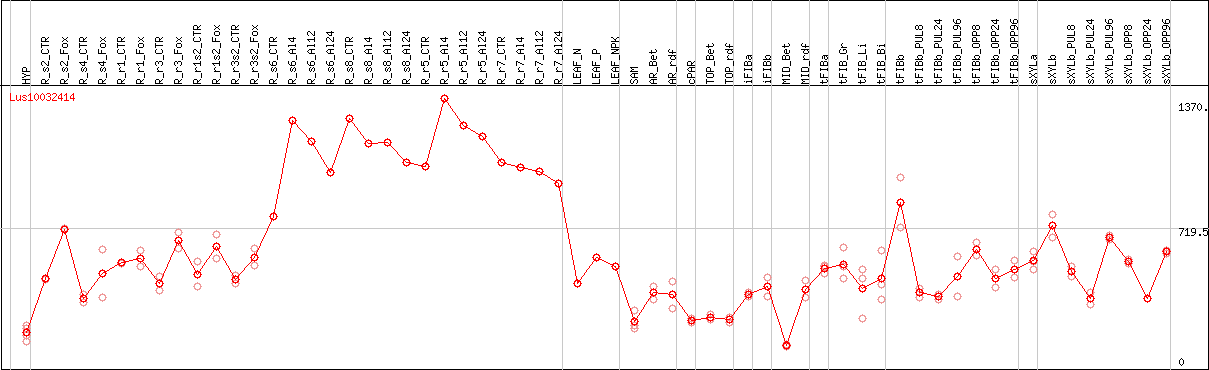
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032414 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.