External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G24010 97 / 5e-24
CSLG2, ATCSLG1
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE LIKE G1, cellulose synthase like G1 (.1)
AT4G24000 97 / 9e-24
ATCSLG2
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE LIKE G2, cellulose synthase like G2 (.1)
AT4G23990 93 / 2e-22
ATCSLG3
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE G3, cellulose synthase like G3 (.1)
AT1G55850 66 / 5e-13
ATCSLE1
cellulose synthase like E1 (.1)
AT4G38190 59 / 8e-11
ATCSLD4
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
AT3G03050 59 / 9e-11
RHD7, ATCSLD3, KJK, CSLD3
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
AT1G02730 59 / 9e-11
SOS6, ATCSLD5
SALT OVERLY SENSITIVE 6, CELLULOSE SYNTHASE LIKE D5, cellulose synthase-like D5 (.1)
AT4G15320 59 / 1e-10
ATCSLB6, ATCSLB06
CELLULOSE SYNTHASE LIKE B6, cellulose synthase-like B6 (.1)
AT2G25540 59 / 1e-10
CESA10
cellulose synthase 10 (.1)
AT2G32540 58 / 3e-10
ATCSLB4, ATCSLB04
CELLULOSE SYNTHASE LIKE B4, cellulose synthase-like B4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023056
170 / 1e-49
AT4G23990 776 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE G3, cellulose synthase like G3 (.1)
Lus10032415
110 / 1e-28
AT4G23990 813 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE G3, cellulose synthase like G3 (.1)
Lus10003196
103 / 5e-26
AT4G23990 815 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE G3, cellulose synthase like G3 (.1)
Lus10023057
87 / 2e-20
AT4G23990 816 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE G3, cellulose synthase like G3 (.1)
Lus10016625
65 / 1e-12
AT1G55850 801 / 0.0
cellulose synthase like E1 (.1)
Lus10026610
64 / 2e-12
AT3G03050 1028 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Lus10013851
63 / 7e-12
AT4G38190 1876 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Lus10026568
62 / 8e-12
AT4G38190 1868 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE D4, cellulose synthase like D4 (.1)
Lus10030453
61 / 3e-11
AT3G03050 1584 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G142201
116 / 6e-33
AT4G24010 200 / 1e-59
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE LIKE G1, cellulose synthase like G1 (.1)
Potri.003G142400
121 / 1e-32
AT4G23990 850 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE G3, cellulose synthase like G3 (.1)
Potri.003G142500
114 / 8e-30
AT4G23990 809 / 0.0
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE G3, cellulose synthase like G3 (.1)
Potri.010G074800
76 / 1e-16
AT4G24000 471 / 2e-156
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE LIKE G2, cellulose synthase like G2 (.1)
Potri.001G449301
69 / 2e-14
AT3G03050 482 / 1e-161
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
Potri.001G369100
69 / 2e-14
AT1G55850 887 / 0.0
cellulose synthase like E1 (.1)
Potri.006G004300
68 / 7e-14
AT1G55850 762 / 0.0
cellulose synthase like E1 (.1)
Potri.010G074700
67 / 1e-13
AT4G24010 469 / 2e-155
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE LIKE G1, cellulose synthase like G1 (.1)
Potri.006G004166
66 / 3e-13
AT1G55850 429 / 1e-145
cellulose synthase like E1 (.1)
Potri.001G136200
62 / 6e-12
AT3G03050 1640 / 0.0
ROOT HAIR DEFECTIVE 7, KOJAK, CELLULOSE SYNTHASE LIKE D3, cellulose synthase-like D3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF03552
Cellulose_synt
Cellulose synthase
Representative CDS sequence
>Lus10032416 pacid=23168670 polypeptide=Lus10032416 locus=Lus10032416.g ID=Lus10032416.BGIv1.0 annot-version=v1.0
ATGGGTTTTATCGCTTCTCCGACCAAGCTCCCACCTCTCCACCAACTCCGGCCACTCCCTCACGCCGTCATCAACCGACGCTTCGCCGCACTCTACGCTT
CCGCCATCTTAGCCTTACTGGACCACCACACCACGTCACTCATCTTCCACCACAAAACCTCCCTATCCATAACCTTACTACACATCGCCTTCCTAATATC
CGACCTCGTACTGGCCTTCATGTGGCTCACCGCTCAGGCCCTCCGCATGTTCCCCGTTCGCCGGGAACAGTTCCCGGCCAACCTCGAGAAGGTCGATCGC
AATGAGTTCCCTGCCCTGGAAGTGTTCATATGCACGGCGGATCCGTACAAGGAGCCGCCGTTGAGGGTGGTGGACACGGACATGTCGGTGCTGGCGTACG
TGTACCCGGCGGAGAAGCTGTCGGTGTACGTGTCGGACGTTGACTCTGTTTGTGTTCATGGATGCTGCTAA
AA sequence
>Lus10032416 pacid=23168670 polypeptide=Lus10032416 locus=Lus10032416.g ID=Lus10032416.BGIv1.0 annot-version=v1.0
MGFIASPTKLPPLHQLRPLPHAVINRRFAALYASAILALLDHHTTSLIFHHKTSLSITLLHIAFLISDLVLAFMWLTAQALRMFPVRREQFPANLEKVDR
NEFPALEVFICTADPYKEPPLRVVDTDMSVLAYVYPAEKLSVYVSDVDSVCVHGCC
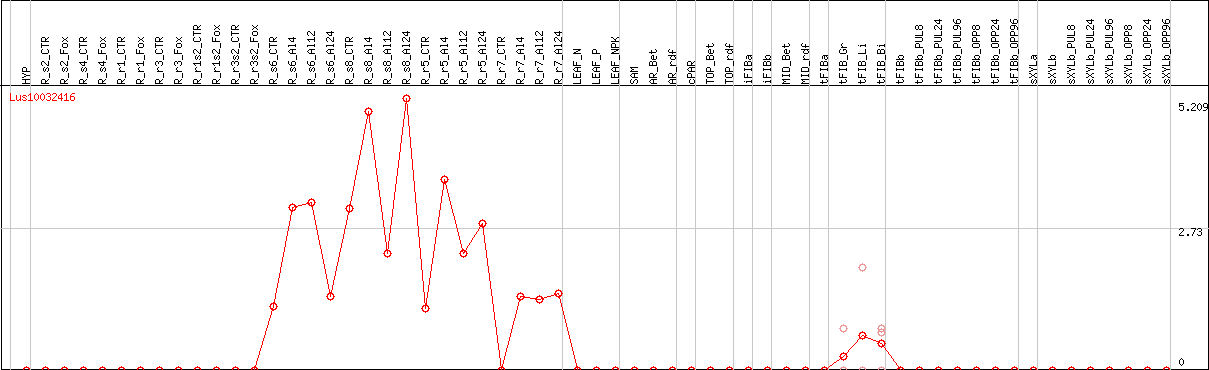
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032416 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.