External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G04160 123 / 4e-33
AIR3
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
AT5G59810 119 / 1e-31
ATSBT5.4
Subtilase family protein (.1)
AT4G10530 85 / 1e-19
Subtilase family protein (.1)
AT4G10520 84 / 3e-19
Subtilase family protein (.1)
AT5G11940 83 / 6e-19
Subtilase family protein (.1)
AT3G14067 79 / 1e-17
Subtilase family protein (.1)
AT4G10550 78 / 3e-17
Subtilase family protein (.1.2.3)
AT4G21326 78 / 3e-17
ATSBT3.12
subtilase 3.12 (.1)
AT5G03620 77 / 9e-17
Subtilisin-like serine endopeptidase family protein (.1)
AT1G32960 76 / 2e-16
ATSBT3.3
Subtilase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10023048
284 / 2e-92
AT5G59810 762 / 0.0
Subtilase family protein (.1)
Lus10006505
170 / 1e-49
AT2G04160 745 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Lus10032096
145 / 1e-40
AT1G20160 654 / 0.0
Subtilisin-like serine endopeptidase family protein (.1.2)
Lus10014591
136 / 2e-37
AT2G04160 716 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Lus10016064
132 / 3e-36
AT2G04160 975 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Lus10039087
107 / 1e-27
AT2G04160 810 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Lus10038772
104 / 1e-26
AT5G59810 815 / 0.0
Subtilase family protein (.1)
Lus10039085
104 / 2e-26
AT5G59810 827 / 0.0
Subtilase family protein (.1)
Lus10039086
102 / 1e-25
AT5G59810 850 / 0.0
Subtilase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G050100
169 / 3e-49
AT2G04160 784 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.011G050300
168 / 3e-49
AT2G04160 786 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.011G050000
167 / 8e-49
AT2G04160 806 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.007G102100
147 / 1e-41
AT2G04160 817 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.007G102250
144 / 1e-40
AT2G04160 821 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.011G165900
137 / 3e-38
AT2G04160 882 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.001G468800
137 / 4e-38
AT5G59810 939 / 0.0
Subtilase family protein (.1)
Potri.005G067200
135 / 3e-37
AT2G04160 786 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.014G171600
125 / 5e-34
AT2G04160 986 / 0.0
AUXIN-INDUCED IN ROOT CULTURES 3, Subtilisin-like serine endopeptidase family protein (.1)
Potri.001G469000
111 / 5e-29
AT5G59810 754 / 0.0
Subtilase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0570
PPP-I
PF05922
Inhibitor_I9
Peptidase inhibitor I9
Representative CDS sequence
>Lus10032424 pacid=23168564 polypeptide=Lus10032424 locus=Lus10032424.g ID=Lus10032424.BGIv1.0 annot-version=v1.0
ATGTCTTATAGTATTTCCTCTCTGCTGCTGTTTGTACTACTCCTGCAACCCCTTTGGCTCATCTCGACTGTGTCTGCTGGTGCTGATAAAAAACAGTCAT
ATGTAGTGTATCTTGGGAGACATGACCAACATAACTCTGGCCGGAATCCTGTTTTTACTGCTGATCAAATGAGTCACGCTTATTTACAGCTGTTGGGTTC
TTGCCTGCTGAGCAAGGAGAAGGCTAAAGAAGCTATATTCTACTCATACTCAAGCCACATCAATGGCTTTGCAGCTATGCTTGAAGATGAGGAAGTAGAT
ATGCTCTCCAGTCACCCAGGAGTGGTATCAGTACTCCCAAATAAAGAGCACCACTTGCATACCACAAGGACATGGGGATTCATGGGGCTAGAAGACAGTA
GTGGAACGATCCCAGATGACTCTATCTGGAAACATGCAAACTTTGGCCAAGATGTCATTATCGGCAACCTTGATACAGGCTAG
AA sequence
>Lus10032424 pacid=23168564 polypeptide=Lus10032424 locus=Lus10032424.g ID=Lus10032424.BGIv1.0 annot-version=v1.0
MSYSISSLLLFVLLLQPLWLISTVSAGADKKQSYVVYLGRHDQHNSGRNPVFTADQMSHAYLQLLGSCLLSKEKAKEAIFYSYSSHINGFAAMLEDEEVD
MLSSHPGVVSVLPNKEHHLHTTRTWGFMGLEDSSGTIPDDSIWKHANFGQDVIIGNLDTG
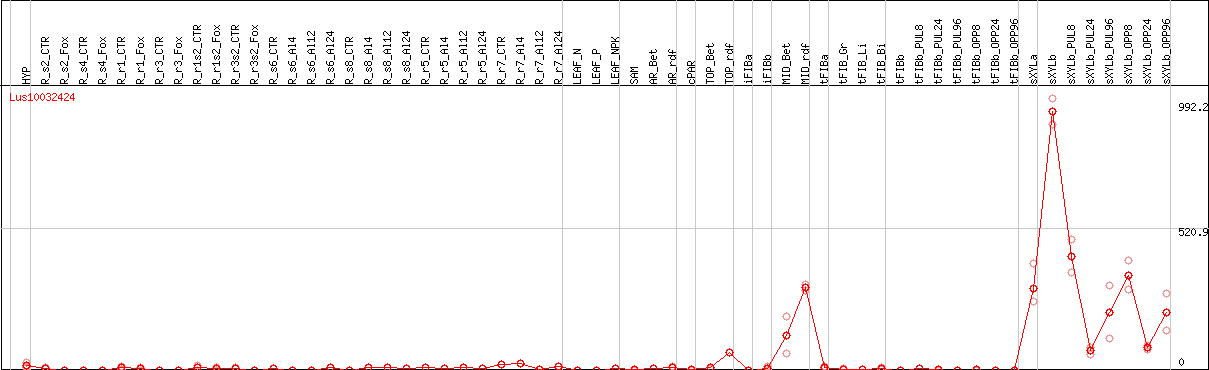
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032424 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.