Lus10032463 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10032463 pacid=23146230 polypeptide=Lus10032463 locus=Lus10032463.g ID=Lus10032463.BGIv1.0 annot-version=v1.0
ATGTCTGGAGCGGAGGAGCTATGGGAGCGGTTAGTTCGCGCTGCACTGAGGAGTGAAAGAACTCGAGCTGCTGCATTTGCTGGTGCTGTTACTGGGATTG
CTGGAAATGTCCCTTCTTCACTTGAGAACAATAGGGATATAGATGGCATTTTGAGAGCTGCTGATGAAATTGAAGACGAAGATCCAAACATCTCTAGAAT
CCTTTGTGAGCATGGGTATTCACTTGCACAAGATTTGGACCCTAATAGTGAAGGAAGAGGCGTTCTACAGTTCAAGACAGGTCTTATGTCTGTTATTAAG
CAAAAACTGACAAAGAGAGAGGGTGGAACAATAGATAGAGGCCAAGACATTGCACGCTTACAGGAATTCTACAAGCTATACAGAGAGCGGAACAGAGTTG
ACACTTTAAAAGAAGAGGAAATGAAACTGCGGGAATCCGGTGCTTTCAGTGGAAATTTGGGCGAGTTGGAGCGAAGAACTATTAGACGGAAAAGAGTATT
TGCAACTCTAAAAGTCTTGGGGTCAGTTCTTCAGCAGTTGAATAAGGATATGCCTGACGAGCTGAGTAGGATGATTAAGTCAGATGCATCTATGACAGAA
GACCTGATTGCTTATAACATTATTCCTCTTGATCTTCAAACTGTAACGAATGCTGTTGTGGCTTTCCCTGAGGTTCGAGCAGCTGTGTCTTCTTTGAGAT
ATTTCCGAGGTCTGCCGATTTTGCCCGAGGATTTTCCTATTCCATCCACGAGAGCTTCTGATATGTTCGATTTTCTGCACTATGTTTTTGGTTTTCAGAA
AGATAATGTCTCTAATCAGCGAGAGAACGTAGTTCACCTCCTCGCAAATGAGCAATCTCGACTTGGCATACCTGATCCAACTGAACCAAAACTAGATGAA
GCTGCTGTACAGAACGTGTTTATGAAGGCACTTGGAAATTATATCAATTGGTGCAGCTACTTGTGCATTCAGCCTGTTTGGAGCAATTCGGAAGCTTTAA
GCACAGAAAAGAAGTTGCTCTATGTTTCTCTGTACTTCCTTATATGGGGTGAGGCTGCCAATATCAGATTTCTTCCAGAATGTTTGTGCTACATATTTCA
CCATATGGCTAGGGAAATGGATGAGATATTGAGGCAGCAAAATGCACAACCTGCGAATAGTTGCATTTCTGAGGATGATGCCCGTGCCCGTGTATCATTC
CTTGATCAAGTTATTCTCCCTTTATATGGGGTGATTGCAGCTGAAGCTTCAAACAATGACAATGGTCGGGCGGCTCATTCTGCTTGGAGAAACTATGATG
ATTTCAATGAGTACTTTTGGTCACTTCACTGCTTTGACCTCAGTTGGCCTTGGCGTCTCAGCTCAACTTTTTTCCAGAAGCCCAAACCAAGGACCAAGGT
TCTCCTCAAAACAGCTGGAAGTCAGAGGAGAGGCAAAACTTCGTTTGTTGAGCATCGGACGTTTCTGCATCTTTACCATAGCTTCCATCGGTTGTGGATT
TTCCTCGTCATGATGTTTCAGGGGCTGGCAATAATTGCTTTCAATGATGGCCGCTTCAACACTAGGACTCTCAGAGAAATTCTAAGTCTGGGTCCAACAT
TTGTTGTGATGAAGTTTTCTGAGAGTGTATTGGATGTCCTCATGATGTATGGTGCTTATTCAACAACACGGCATGTGGCTGTTTCTCGAATTCTTCTTAG
ATTTGTTTGGTTCGCCTGTGCTTCTGTCTTCATATGTTTCCTTTATGTGAAAGCCTTACAGGAGCCAGATACAAGTTCAGTCCTTTTCAAGCTATATGTT
ATCGTTATAGGAATTTATGCTGGCGTCCAGTTTTTCCTAGGTTTCTTAACGCGGATCCCGGCTTGTCACCTCATGACTAATCAATGTGATCAGTGGTCAA
TAGTTCGATTTGTAAAATGGATGCGCCAGGAACGATACTATGTCGGCCGTGGAATGTATGAACGAACTAGTGATTTCATCAAGTACATGGTTTTTTGGCT
TGTCATCCTGAGTGCAAAATTTTCATTCGCATATTTCCTTCAGATCAAACCATTGGTGGAACCAACCAAGATTATAGTCAAGATGACTGATAATATCCAA
TATTCTTGGCATGACCTTGTTTCTAAAAATAATCATAATGCCCTTACAATTGTGAGCCTTTGGGCTCCCGTGGTTGCTATCTATCTTCTAGATATCCATG
TGTTTTACACTCTTACCTCTGCTGTGTGGGGGTTCCTGCTTGGAGCTAGAGATCGCCTTGGAGAAATCCGGTCATTGGAATCTGTGCACAAACTTTTTGA
GGAGTTCCCTGCAGCTTTTATGAGAACTCTCCATTCTGATAGGACTGTTGGCAATGCACTCGAGCCTGTTGAGAAGAAAAAGATTGATGCAGCCCAATTC
TCCCCATTTTGGAACGAGATCATAAAGAACCTGAGGGAAGAAGATTATATAGCAAATTTCGAATTGGAATTGCTTCAAATGCCCAGAAATTCGGGGAATC
TTCCATTGGTTCAGTGGCCACTTTTTCTTCTTGCCAACAAGATATTTTTGGCCAGGGATATTGCTGCAGAAAGTAGAGATTCCCAACTAGAGTTGTGGGA
GAGGATTTCACGCGATGAGTATATGAAATATGCTGTTGAGGAGTGTTACCATGCTCTTAGATATATTCTTACGGAAATATTTGAAGGTGAAGGGAGGATG
TGGGTTGAAAGGGTGTACGAGGACATACAGGCAAGTATACAGAACAAGAGTATTCATGTTGACTTTCAGTTGACTAAGCTGGCACTCGTTATTCAACGAG
TAACTGCTTTATTGGGAGTTTTAAAAGAGGCAGAAACTTCTGATATGGAAAAAGGTGCAATCAAGGCTGTTCAAGATCTATATGATGTTATTCAGCATGA
TGTCCTTTCTATCGACAAAAGGGAACATTATGACACTTGGAATTTGCTGTCAAAGGCGAGAACCGAAGGACGATTATTTACAAACTTGAAGTGGCCTCGT
GATCCGGAATTGAGAACACAAATTAAAAGACTACATTCACTTTTAACAATCAAAGATTCTGCTGCAAACGTTCCCAACAATATTGAAGCCAGGCGTAGGC
TCGAGTTCTTCACAAATTCACTTTTTATGGACATGCCCCTTGCAAAGCCCGTACGGGAGATGCTCTCCTTCAGTGTCTTTACTCCATATTATTCAGAGAT
TGTTCTTTATAGCATGGCTGAGCTTCTTAAGAAAAACGAGGATGGAATATCAATTCTGTTTTACCTTCAGAAAATATATCCAGATGAGTGGAAGAATTTT
CTTGCTCGAATTGGACGTGATGAAAATTCTGTCGATACAGAACTGTTCGATAGTCCTACAGACATCCTTGAGCTTCGGTTTTGGGCTTCCTATAGGGGTC
AAACACTGGCTCGCACAGTGCGGGGGATGATGTACTACAGGAAGGCTATCATGCTTCAAAGTTACTTGGAAAGAGGGACTGCTCAAGATGTGGAATCTGC
TATTGGTAGTAAAGATGCTACCGATACCCAAGGTTTTGAGTTGTCTCCTGAAGCCAGAGCGCAGGCAGATATAAAGTTTACATACGTGGTGACTTGCCAA
ATATATGGAAAACAAAAGGAAGAGCAGAAACCTGAGGCTGCTGATATTGCACTGCTTATGCAAAGAAATGAAGCTCTTCGTGTTGCCTTTATCGATGAAG
TTGAAACGTTGAAAGAGGGTCATGTGCAGAGAGAATTTTTCTCCAAGCTCGTCAAAGCTGATATCAATGGGAAAGATAAGGAAATATACTCAATAAAGTT
ACCAGGTAATCCAAAACTTGGAGAAGGAAAACCTGAGAATCAAAATCATGCTATTGTTTTTACACGTGGAAATGCAGTGCAAACGATAGATATGAATCAG
GATAATTATTTTGAAGAAGCCTTGAAAATGAGAAATCTGCTCGAAGAATTTCATCGTGATCACGGTATTCGGCCTGCAACAATTCTTGGTGTCAGGGAAC
ATGTATTTACTGGAAGTGTATCTTCTTTAGCCTCTTTTATGTCAAACCAAGAAACAAGCTTTGTAACTCTTGGTCAGCGAGTTCTGTCTAATCCTTTGAA
GGTCCGAATGCATTATGGTCATCCAGATGTTTTCGACAGAGTTTTCCACATTACACGGGGTGGTATAAGCAAGGCTTCCCGCGTTATTAATATAAGTGAA
GATATATTTGCAGGCTTCAATTCGACCTTGCGCCAAGGAAACATTACACACCATGAGTACATACAGGTTGGAAAAGGACGGGATGTTGGTCTCAATCAGA
TTGCTGTATTCGAAGGGAAGGTAGCTGGTGGCAATGGCGAGCAGGTTCTTAGCCGGGATGTTTTCAGGCTTGGGCAGCTGTTTGATTTCTTCAGAATGAT
GTCTTTTTACTTCACTACTGTGGGATATTACTTCTGCACGATGTTAACTGTGCTGACCGTGTACATGTTCTTGTATGGGAAGGCATATTTGGCGCTTTCT
GGAGTTGGTGAAACCATCCAAGAGAGAGCCCAAATATTGCAGAATACGGCCTTGAGTGCAGCTCTAAATACACAGTTTCTCTTTCAGATTGGTATTTTTA
CTGCTGTGCCAATGGTCTTGGGCTTCATATTGGAACAGGGATTCCTAAGGGCAGTTGTCAGTTTTATCACAATGCAACTTCAACTTTGTTCCGTCTTCTT
TACATTCTCTTTGGGCACACGGACACATTACTTCGGACGAACTATTCTTCATGGTGGCGCAAGGTACCAAGCTACTGGTAGGGGATTCGTGGTTCGCCAT
ATTAAGTTCTCGGAAAACTACCGCCTTTATTCCCGGAGTCATTTTGTCAAGGGACTAGAAGTTGCAGTTTTGCTGATTGTATACCTTGCTTATGGCTACA
ATGAAGGTGGAGCGCTCTCTTATATTCTCCTTACAGTTAGCAGTTGGTTCATGGCCCTCTCCTGGCTGTACGCCCCCTATTTGTTCAATCCCTCAGGATT
TGAATGGCAGAAAACGGTCGAGGACTTCAGAGATTGGACAAATTGGCTTCTTTACAGAGGCGGAATTGGGGTGAAGGGTGAAGAAAGCTGGGAAGCTTGG
TGGGAGGAAGAACTGGCACATATCCGAACTTTTAGCGGGAGGATCATAGAGACAATTCTGAGTCTCCGATTTTTCATTTTCCAGTATGGCATTATATACA
AGCTTGACGTTCAAAGAAATGATACTTCTTTGACCGTGTACGGTATATCATGGGCTGTGTTGGCAGTGCTTATAGTACTCTTCAAGGTTTTCACTTTCAG
TCAGAAGATATCTGTAAACTTCCAGCTTCTACTTCGATTCATCCAAGGGGTTGCCTTTTTGCTGGCTTTGGCGGGTTTAGCTGTTGCAGTTGTGTTCACA
AACCTTTCGGTTCCAGACATATTTGCTTGCATTCTAGCTTTCATCCCTACTGGATGGGGAATCTTATCGATAGCTGCAGCTTGGAAACCATTGATGAAGA
AAGTTGGTCTATGGAAGTCGATCCGTTCAATTGCCCGTCTGTATGATGCCGGAATGGGGATGATCATCTTCATACCAATTGCATTCTTTTCGTGGTTCCC
ATTCATGTCGACATTCCAAACCCGGCTCATGTTCAACCAAGCATTCAGCCGGGGCTTGGAAATTTCTCTGATTCTGGCTGGAAACAACCCCAATACCAGG
TTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10032463 pacid=23146230 polypeptide=Lus10032463 locus=Lus10032463.g ID=Lus10032463.BGIv1.0 annot-version=v1.0
MSGAEELWERLVRAALRSERTRAAAFAGAVTGIAGNVPSSLENNRDIDGILRAADEIEDEDPNISRILCEHGYSLAQDLDPNSEGRGVLQFKTGLMSVIK
QKLTKREGGTIDRGQDIARLQEFYKLYRERNRVDTLKEEEMKLRESGAFSGNLGELERRTIRRKRVFATLKVLGSVLQQLNKDMPDELSRMIKSDASMTE
DLIAYNIIPLDLQTVTNAVVAFPEVRAAVSSLRYFRGLPILPEDFPIPSTRASDMFDFLHYVFGFQKDNVSNQRENVVHLLANEQSRLGIPDPTEPKLDE
AAVQNVFMKALGNYINWCSYLCIQPVWSNSEALSTEKKLLYVSLYFLIWGEAANIRFLPECLCYIFHHMAREMDEILRQQNAQPANSCISEDDARARVSF
LDQVILPLYGVIAAEASNNDNGRAAHSAWRNYDDFNEYFWSLHCFDLSWPWRLSSTFFQKPKPRTKVLLKTAGSQRRGKTSFVEHRTFLHLYHSFHRLWI
FLVMMFQGLAIIAFNDGRFNTRTLREILSLGPTFVVMKFSESVLDVLMMYGAYSTTRHVAVSRILLRFVWFACASVFICFLYVKALQEPDTSSVLFKLYV
IVIGIYAGVQFFLGFLTRIPACHLMTNQCDQWSIVRFVKWMRQERYYVGRGMYERTSDFIKYMVFWLVILSAKFSFAYFLQIKPLVEPTKIIVKMTDNIQ
YSWHDLVSKNNHNALTIVSLWAPVVAIYLLDIHVFYTLTSAVWGFLLGARDRLGEIRSLESVHKLFEEFPAAFMRTLHSDRTVGNALEPVEKKKIDAAQF
SPFWNEIIKNLREEDYIANFELELLQMPRNSGNLPLVQWPLFLLANKIFLARDIAAESRDSQLELWERISRDEYMKYAVEECYHALRYILTEIFEGEGRM
WVERVYEDIQASIQNKSIHVDFQLTKLALVIQRVTALLGVLKEAETSDMEKGAIKAVQDLYDVIQHDVLSIDKREHYDTWNLLSKARTEGRLFTNLKWPR
DPELRTQIKRLHSLLTIKDSAANVPNNIEARRRLEFFTNSLFMDMPLAKPVREMLSFSVFTPYYSEIVLYSMAELLKKNEDGISILFYLQKIYPDEWKNF
LARIGRDENSVDTELFDSPTDILELRFWASYRGQTLARTVRGMMYYRKAIMLQSYLERGTAQDVESAIGSKDATDTQGFELSPEARAQADIKFTYVVTCQ
IYGKQKEEQKPEAADIALLMQRNEALRVAFIDEVETLKEGHVQREFFSKLVKADINGKDKEIYSIKLPGNPKLGEGKPENQNHAIVFTRGNAVQTIDMNQ
DNYFEEALKMRNLLEEFHRDHGIRPATILGVREHVFTGSVSSLASFMSNQETSFVTLGQRVLSNPLKVRMHYGHPDVFDRVFHITRGGISKASRVINISE
DIFAGFNSTLRQGNITHHEYIQVGKGRDVGLNQIAVFEGKVAGGNGEQVLSRDVFRLGQLFDFFRMMSFYFTTVGYYFCTMLTVLTVYMFLYGKAYLALS
GVGETIQERAQILQNTALSAALNTQFLFQIGIFTAVPMVLGFILEQGFLRAVVSFITMQLQLCSVFFTFSLGTRTHYFGRTILHGGARYQATGRGFVVRH
IKFSENYRLYSRSHFVKGLEVAVLLIVYLAYGYNEGGALSYILLTVSSWFMALSWLYAPYLFNPSGFEWQKTVEDFRDWTNWLLYRGGIGVKGEESWEAW
WEEELAHIRTFSGRIIETILSLRFFIFQYGIIYKLDVQRNDTSLTVYGISWAVLAVLIVLFKVFTFSQKISVNFQLLLRFIQGVAFLLALAGLAVAVVFT
NLSVPDIFACILAFIPTGWGILSIAAAWKPLMKKVGLWKSIRSIARLYDAGMGMIIFIPIAFFSWFPFMSTFQTRLMFNQAFSRGLEISLILAGNNPNTR
L
|
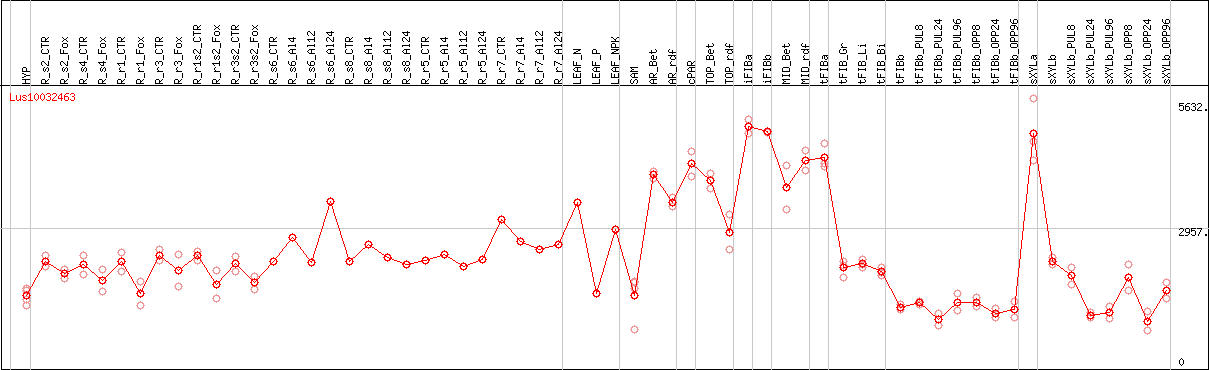
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032463 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
