External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G50720 173 / 4e-57
ATHVA22E
ARABIDOPSIS THALIANA HVA22 HOMOLOGUE E, HVA22 homologue E (.1)
AT4G24960 171 / 4e-56
ATHVA22D
ARABIDOPSIS THALIANA HVA22 HOMOLOGUE D, HVA22 homologue D (.1.2.3)
AT2G42820 117 / 3e-34
HVA22F
HVA22-like protein F (.1)
AT1G74520 114 / 3e-33
ATHVA22A
HVA22 homologue A (.1)
AT1G69700 101 / 5e-28
ATHVA22C
HVA22 homologue C (.1)
AT5G62490 100 / 8e-28
ATHVA22B
ARABIDOPSIS THALIANA HVA22 HOMOLOGUE B, HVA22 homologue B (.1)
AT4G36720 61 / 5e-12
HVA22K
HVA22-like protein K (.1)
AT1G75700 56 / 3e-10
HVA22G
HVA22-like protein G (.1)
AT1G19950 56 / 6e-10
HVA22H
HVA22-like protein H (ATHVA22H) (.1)
AT5G42560 54 / 3e-09
Abscisic acid-responsive (TB2/DP1, HVA22) family protein (.1), Abscisic acid-responsive (TB2/DP1, HVA22) family protein (.2), Abscisic acid-responsive (TB2/DP1, HVA22) family protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10043186
258 / 2e-90
AT5G50720 169 / 1e-55
ARABIDOPSIS THALIANA HVA22 HOMOLOGUE E, HVA22 homologue E (.1)
Lus10023605
120 / 4e-35
AT1G74520 246 / 7e-84
HVA22 homologue A (.1)
Lus10024234
120 / 5e-35
AT1G74520 244 / 5e-83
HVA22 homologue A (.1)
Lus10031386
111 / 4e-32
AT2G42820 246 / 4e-85
HVA22-like protein F (.1)
Lus10042193
116 / 3e-31
AT5G39850 327 / 2e-108
Ribosomal protein S4 (.1)
Lus10010944
109 / 3e-31
AT2G42820 246 / 7e-85
HVA22-like protein F (.1)
Lus10037191
107 / 5e-30
AT1G69700 231 / 6e-78
HVA22 homologue C (.1)
Lus10008623
96 / 7e-26
AT1G74520 234 / 5e-80
HVA22 homologue A (.1)
Lus10033152
60 / 3e-11
AT5G42560 328 / 7e-113
Abscisic acid-responsive (TB2/DP1, HVA22) family protein (.1), Abscisic acid-responsive (TB2/DP1, HVA22) family protein (.2), Abscisic acid-responsive (TB2/DP1, HVA22) family protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03134
TB2_DP1_HVA22
TB2/DP1, HVA22 family
Representative CDS sequence
>Lus10032557 pacid=23146340 polypeptide=Lus10032557 locus=Lus10032557.g ID=Lus10032557.BGIv1.0 annot-version=v1.0
ATGAGCCGTTTCTGGACTTTCCTCACTCATGTTCACGCACTTGCTGGGCCGGTTGTGATGTTGCTGTACCCACTGTATGCATCGGTGATAGCAATAGAGA
GTCCATCAAAAGAGGACGACGAGCAGTGGCTGGCGTATTGGATACTATACTCCTTCTTAACCCTATCGGAGATGCTCCTCCAGTCGATTCTTGAGTGGAT
ACCGATTTGGTACACGTTGAAGCTGATGCTGGTGGCGTGGCTGGTGCTCCCGCAGTTCAGAGGGGCGGCTTTTATCTATGACCGATTTGTTAGGCAACAC
ATCAGGAAGCATCTCACCCACAACAGTACTGCTAATGGCTCCTCCAAGGCCACCAACAAGTTTGTTCACTTTGTTTCCCCCAAAAAAGCGGAATGA
AA sequence
>Lus10032557 pacid=23146340 polypeptide=Lus10032557 locus=Lus10032557.g ID=Lus10032557.BGIv1.0 annot-version=v1.0
MSRFWTFLTHVHALAGPVVMLLYPLYASVIAIESPSKEDDEQWLAYWILYSFLTLSEMLLQSILEWIPIWYTLKLMLVAWLVLPQFRGAAFIYDRFVRQH
IRKHLTHNSTANGSSKATNKFVHFVSPKKAE
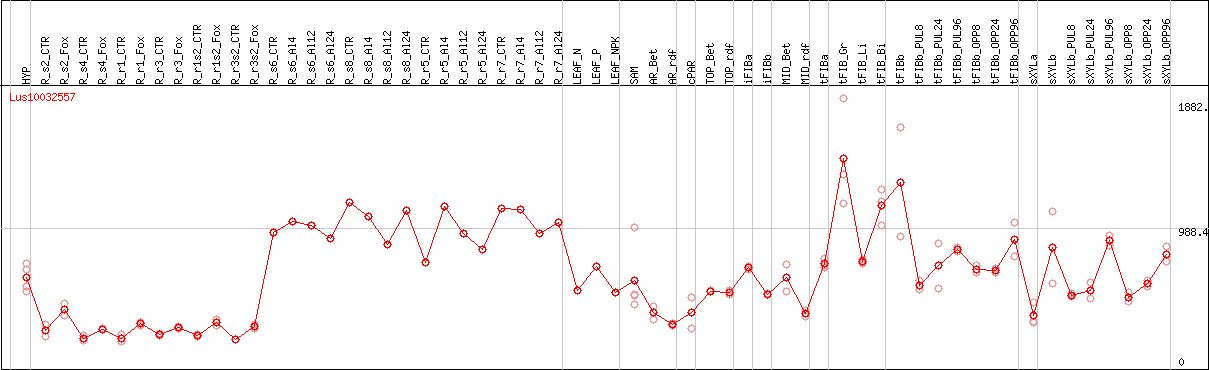
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032557 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.