External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G26670 192 / 2e-62
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
AT5G55510 187 / 6e-60
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
AT3G10110 54 / 3e-09
MEE67
maternal effect embryo arrest 67, Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
AT1G18320 53 / 5e-09
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
AT5G24650 41 / 0.0002
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
AT3G49560 40 / 0.0006
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10043170
332 / 4e-117
AT4G26670 220 / 5e-73
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Lus10026848
57 / 4e-10
AT3G10110 224 / 5e-76
maternal effect embryo arrest 67, Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Lus10020223
56 / 2e-09
AT3G10110 229 / 8e-78
maternal effect embryo arrest 67, Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Lus10032913
42 / 0.0002
AT5G24650 234 / 6e-77
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G360300
204 / 9e-67
AT5G55510 254 / 2e-86
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Potri.011G088500
198 / 3e-64
AT5G55510 252 / 1e-85
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Potri.001G281200
54 / 4e-09
AT3G10110 205 / 4e-68
maternal effect embryo arrest 67, Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Potri.009G076400
52 / 2e-08
AT3G10110 194 / 1e-63
maternal effect embryo arrest 67, Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Potri.012G004600
42 / 9e-05
AT5G24650 262 / 2e-87
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
Potri.015G000600
40 / 0.0008
AT5G24650 300 / 6e-103
Mitochondrial import inner membrane translocase subunit Tim17/Tim22/Tim23 family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02466
Tim17
Tim17/Tim22/Tim23/Pmp24 family
Representative CDS sequence
>Lus10032577 pacid=23146257 polypeptide=Lus10032577 locus=Lus10032577.g ID=Lus10032577.BGIv1.0 annot-version=v1.0
ATGGCGACAACGGCGGATTCCCCAAACCCTGATTCCGACCTCCCTGGTTCTAGTGAAAACCCTACCTCTGCTCCTCCATCTACTCCTCCCTCCTCCGCCA
TCATCCCATCCAGTCCTTCCGTCTGCCTATTTGGGTTCGCTACCGAGTCCGCCGGTGGCGCATTGATAGGCTCCATCTTCGGCTTCGGTTCTGGTTTAGT
TCGAAAGAAAGGCTTTAAGGGATCTCTTGCGGAGGCCGGTTCTCATGCCAAGACATTTGCTCTTCTGTCTGGAGTACACAGCTTGGTTATATGCTTTTTG
AGGAGGCTACGTGGGAAGGATGATGTCATCAATGCTGGTGTGGCTGGGTGTTGTACTGGTCTTGCTTTGAGCTTCCCAGGTACACCTCAGGCTCTGTTGC
AGAACTGTGTGACGTTTGGAGCATTCTCGTTTATCACTGAAGGGCTCAACAAGCGGCAGACAGCACTTGCAGCTCAACCATTTTCCGGAAACCATCGCGC
ACCTCGTTCCCTTGCCCTACCTCTCGCGGTTCCACTTCCTGAGGAACTAAACCAGGCCTTCTCTTTCTTCCGTGATTCCTTGAGGAAGCTGAATCATCGG
AAGTCGCCGCCTTGGAGTGCCTGA
AA sequence
>Lus10032577 pacid=23146257 polypeptide=Lus10032577 locus=Lus10032577.g ID=Lus10032577.BGIv1.0 annot-version=v1.0
MATTADSPNPDSDLPGSSENPTSAPPSTPPSSAIIPSSPSVCLFGFATESAGGALIGSIFGFGSGLVRKKGFKGSLAEAGSHAKTFALLSGVHSLVICFL
RRLRGKDDVINAGVAGCCTGLALSFPGTPQALLQNCVTFGAFSFITEGLNKRQTALAAQPFSGNHRAPRSLALPLAVPLPEELNQAFSFFRDSLRKLNHR
KSPPWSA
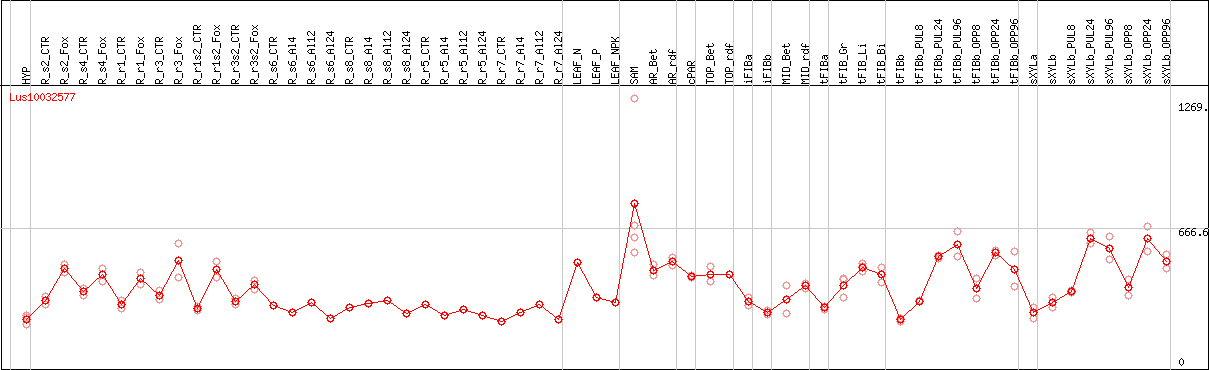
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032577 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.