External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G02150 124 / 1e-33
TCP
TFPD, PTF1, TCP13
TEOSINTE BRANCHED1, CYCLOIDEA AND PCF TRANSCRIPTION FACTOR 13, plastid transcription factor 1 (.1.2)
AT5G60970 119 / 6e-31
TCP
TCP5
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 5 (.1)
AT5G08070 112 / 2e-29
TCP
TCP17
TCP domain protein 17 (.1)
AT1G30210 96 / 1e-22
TCP
ATTCP24, TCP24
TEOSINTE BRANCHED 1, cycloidea, and PCF family 24 (.1.2)
AT1G53230 94 / 1e-21
TCP
TCP3
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 3 (.1)
AT4G18390 91 / 2e-20
TCP
TCP2
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 2 (.1.2)
AT3G15030 89 / 1e-19
TCP
TCP4, MEE35
maternal effect embryo arrest 35, TCP family transcription factor 4 (.1.2.3)
AT2G31070 86 / 1e-18
TCP
TCP10
TCP domain protein 10 (.1)
AT1G67260 69 / 7e-13
TCP
TCP1
TCP family transcription factor (.1.2)
AT1G68800 67 / 2e-12
TCP
TCP12, BRC2, TCP18
BRANCHED 2, TCP domain protein 12 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017856
115 / 1e-29
AT5G60970 175 / 6e-52
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 5 (.1)
Lus10034679
115 / 1e-29
AT5G60970 169 / 2e-49
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 5 (.1)
Lus10023297
93 / 6e-21
AT4G18390 207 / 8e-63
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 2 (.1.2)
Lus10038512
93 / 8e-21
AT4G18390 204 / 1e-60
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 2 (.1.2)
Lus10000463
85 / 2e-18
AT3G15030 224 / 1e-69
maternal effect embryo arrest 35, TCP family transcription factor 4 (.1.2.3)
Lus10035055
75 / 3e-15
AT1G68800 102 / 7e-25
BRANCHED 2, TCP domain protein 12 (.1)
Lus10021713
74 / 6e-15
AT3G18550 107 / 2e-26
BRANCHED 1, TCP family transcription factor (.1.2)
Lus10003078
66 / 4e-12
AT1G67260 105 / 2e-25
TCP family transcription factor (.1.2)
Lus10034072
66 / 9e-12
AT1G67260 100 / 4e-23
TCP family transcription factor (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G116100
135 / 1e-36
AT3G02150 173 / 1e-50
TEOSINTE BRANCHED1, CYCLOIDEA AND PCF TRANSCRIPTION FACTOR 13, plastid transcription factor 1 (.1.2)
Potri.017G094800
132 / 1e-35
AT3G02150 161 / 4e-47
TEOSINTE BRANCHED1, CYCLOIDEA AND PCF TRANSCRIPTION FACTOR 13, plastid transcription factor 1 (.1.2)
Potri.015G058800
124 / 1e-32
AT5G60970 177 / 6e-52
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 5 (.1)
Potri.011G083100
94 / 3e-21
AT4G18390 150 / 5e-41
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 2 (.1.2)
Potri.004G065800
93 / 5e-21
AT4G18390 151 / 2e-41
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 2 (.1.2)
Potri.001G375800
89 / 1e-19
AT1G53230 279 / 1e-90
TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 3 (.1)
Potri.011G096600
89 / 2e-19
AT3G15030 262 / 7e-84
maternal effect embryo arrest 35, TCP family transcription factor 4 (.1.2.3)
Potri.019G091300
86 / 5e-19
AT3G15030 227 / 5e-71
maternal effect embryo arrest 35, TCP family transcription factor 4 (.1.2.3)
Potri.013G119400
85 / 2e-18
AT3G15030 243 / 4e-77
maternal effect embryo arrest 35, TCP family transcription factor 4 (.1.2.3)
Potri.017G112000
69 / 4e-13
AT1G67260 111 / 1e-27
TCP family transcription factor (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03634
TCP
TCP family transcription factor
Representative CDS sequence
>Lus10032984 pacid=23160450 polypeptide=Lus10032984 locus=Lus10032984.g ID=Lus10032984.BGIv1.0 annot-version=v1.0
ATGCATGAAGGTGTTGATCCTAGCGGAAGGTACAGTATTAGAGACGAGAGACAAGAAGAGGAGCAGAAGAATGTTGATATTATAACTAGGAAGAAGCAGG
TGGAGGCTGGAGGGAGGACATGGTTGAGGGTGAAAGACCCGAGGATAGTAAGGGTATCATCATCAGCTGGTAGTGGTAATAAAGATAGGCATAGCAAAGT
GTGCACCATTAGAGGATTAAGGGATAGGAGGGTTAGGCTTTCTGTTCCTACTGCCATCCAGTTGTACGACCTCCAATCTAGACTTGGCCTTAATCAGCCA
AGTAAGGTTGTCGATTGGCTTCTCAATGCTGCCAAGTCCGACATTGATCAACTCCCTCCTCTCCCTTTCTTCCCTTACCCAACTACTACAACCTCCACCC
AATATTCCACTACTAGCACTTTCATCACCAACCATTATCCACAACCACCACATCATGATTTCTGGAATAGCAAATTTGAACAAGACCAAGACCCAAGTCC
TTTTAATTACCAAGCTGCTTCATTCCATCAACATCATCAAAGTAGACCAATTTACGATGATCTCCAGCCAGCGGGGAGTATTAATAATGATGAAACAGGC
GGCTTTCATTTGGGAGCATCAGGAGGAGCGACTCTTAGGTTTCAACCAATAAACCATCATCATCATACCGACACTAATGCAAGGCCGCCTCTGGTGACAT
TGCCATCAGTATTGATGTCCCTATCATCGTCACAACAGCAGCAGCAGCAGCAGCTAGGTGTACTAGGAGGATGTAATATTAGAATGGATCATGATGATGA
AGAAGATCAGGAGAAGGAGGAAGGTGGTTCAAGTAGCCATGAACAACAACAATTTGCTAATACTAACTTCCATACGCTGCTACGCCTTCATTCTACTACC
AGTGTTGTTGCTCCTCCATCATCCTCATCCTCAATCTTATAA
AA sequence
>Lus10032984 pacid=23160450 polypeptide=Lus10032984 locus=Lus10032984.g ID=Lus10032984.BGIv1.0 annot-version=v1.0
MHEGVDPSGRYSIRDERQEEEQKNVDIITRKKQVEAGGRTWLRVKDPRIVRVSSSAGSGNKDRHSKVCTIRGLRDRRVRLSVPTAIQLYDLQSRLGLNQP
SKVVDWLLNAAKSDIDQLPPLPFFPYPTTTTSTQYSTTSTFITNHYPQPPHHDFWNSKFEQDQDPSPFNYQAASFHQHHQSRPIYDDLQPAGSINNDETG
GFHLGASGGATLRFQPINHHHHTDTNARPPLVTLPSVLMSLSSSQQQQQQQLGVLGGCNIRMDHDDEEDQEKEEGGSSSHEQQQFANTNFHTLLRLHSTT
SVVAPPSSSSSIL
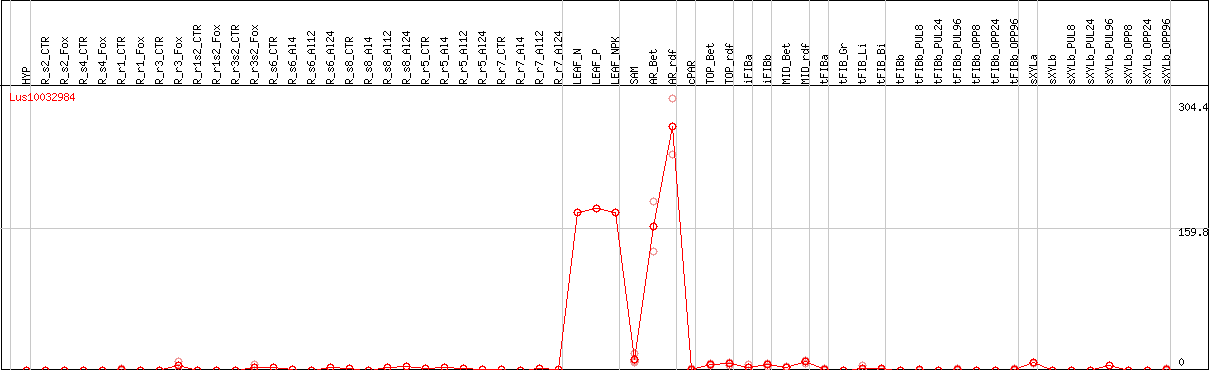
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10032984 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.