External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G48420 407 / 3e-143
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT4G39970 161 / 4e-47
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT5G45170 77 / 8e-16
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT4G21470 69 / 7e-13
ATFMN/FHY
riboflavin kinase/FMN hydrolase (.1)
AT1G56500 67 / 7e-12
haloacid dehalogenase-like hydrolase family protein (.1)
AT4G11570 54 / 4e-08
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
AT4G25840 48 / 5e-06
GPP1
glycerol-3-phosphatase 1 (.1)
AT5G57440 42 / 0.0002
GPP2, GS1
GLYCEROL-3-PHOSPHATASE 2, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
AT2G38740 40 / 0.0009
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013663
591 / 0
AT3G48420 427 / 4e-151
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10000633
168 / 6e-50
AT4G39970 427 / 1e-151
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10023149
167 / 2e-49
AT4G39970 423 / 4e-150
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10016820
142 / 9e-43
AT3G48420 105 / 4e-29
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10018280
87 / 1e-18
AT5G45170 410 / 1e-136
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10040631
83 / 1e-17
AT5G45170 427 / 3e-149
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Lus10018906
69 / 6e-13
AT4G21470 556 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Lus10028605
66 / 5e-12
AT4G21470 558 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Lus10029840
65 / 3e-11
AT1G56500 1528 / 0.0
haloacid dehalogenase-like hydrolase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.015G088500
440 / 3e-156
AT3G48420 434 / 5e-154
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.007G095200
167 / 2e-49
AT4G39970 439 / 3e-156
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.014G003916
81 / 2e-19
AT3G48420 87 / 2e-22
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.015G112500
86 / 1e-18
AT5G45170 386 / 2e-133
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.002G077200
70 / 3e-13
AT4G21470 561 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Potri.005G183400
67 / 3e-12
AT4G21470 575 / 0.0
riboflavin kinase/FMN hydrolase (.1)
Potri.013G007800
63 / 8e-11
AT1G56500 1489 / 0.0
haloacid dehalogenase-like hydrolase family protein (.1)
Potri.018G092500
57 / 4e-09
AT5G57440 398 / 2e-142
GLYCEROL-3-PHOSPHATASE 2, Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1)
Potri.001G104366
56 / 1e-08
AT4G11570 494 / 9e-176
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
Potri.003G127100
56 / 1e-08
AT4G11570 493 / 2e-175
Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.1), Haloacid dehalogenase-like hydrolase (HAD) superfamily protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0137
HAD
PF00702
Hydrolase
haloacid dehalogenase-like hydrolase
Representative CDS sequence
>Lus10033034 pacid=23177041 polypeptide=Lus10033034 locus=Lus10033034.g ID=Lus10033034.BGIv1.0 annot-version=v1.0
ATGGCGTCGGCGACTGTCTCTCTTTCATTCCTGCCGTTTGATTCCATTGCTTCCCCAAATGTTAAGCGGACTTCAATTTTCAAATTCATTTGGTCGCATG
AATCCAATTTATCGTCCTCAATGACGGGTAGGAGGATTTGTGTCAGCAGAAGCATCCGCAAGTCAGGTGCAGCTCGGGTAACTTGGTTGACGGCGACTGC
GGCTGCTTCGTCTTCTCCTTCACCTACGGCTTTGCCTTCTGCCCTACTATTCGACTGCGACGGCGTGCTCGTCGATACTGAAAAGGACGGCCACAGGGTT
TCCTTCAACGACACTTTCGATGAAAAGGAATTGGGGGTCACTTGGGACGTGGAGTTGTATGGAGAGTTGTTGAAAATTGGAGGTGGAAAAGAAAGGATGA
CAACATACTTCAATAAAGTTGGGTGGCCAGAGAATGCTCCAAAGAGTGAAGAGGAAAGGAAAGCTTTCATTGCTTCACTTCATAAAAGGAAGACAGAGTT
GTTCATGGCTCTAATTGAGAAGAGATTACTGCCTCTACGTCCTGGGGTTGCAAAGCTGATTGATCAGGCAATGGGAAAAGGGGTGCCTGTTGCTGTATGC
AGCACCTCCAATGAGAAAGCGGTTTCTGCCATAGTCTCCTACTTGTTGGGACCTGAACGTGCACAGAAGATGAAGATATTTGCTGGAGATGTAGTCCCTA
GGAAAAAGCCTGATCCAGCAATTTATACACTAGCAGCCACCACACTAGGGGTAGATCCTCAAAGCTGCGTAGTAATAGAGGACAGTGGAATAGGACTTGC
AGCAGCTAAAGCAGCAGGGATGAAATGTATAATCACAAAGAGCGGGTACACAGCTGAGGAGGACTTCTTGAATGCAGACGCAGTGTTTGATTTCATTGGA
GACCCACCAGAAGAACGATTTGACTTTGACTTCTGTGCTAGCCTTGTTGAGAAGCAATATGTGAGCTAA
AA sequence
>Lus10033034 pacid=23177041 polypeptide=Lus10033034 locus=Lus10033034.g ID=Lus10033034.BGIv1.0 annot-version=v1.0
MASATVSLSFLPFDSIASPNVKRTSIFKFIWSHESNLSSSMTGRRICVSRSIRKSGAARVTWLTATAAASSSPSPTALPSALLFDCDGVLVDTEKDGHRV
SFNDTFDEKELGVTWDVELYGELLKIGGGKERMTTYFNKVGWPENAPKSEEERKAFIASLHKRKTELFMALIEKRLLPLRPGVAKLIDQAMGKGVPVAVC
STSNEKAVSAIVSYLLGPERAQKMKIFAGDVVPRKKPDPAIYTLAATTLGVDPQSCVVIEDSGIGLAAAKAAGMKCIITKSGYTAEEDFLNADAVFDFIG
DPPEERFDFDFCASLVEKQYVS
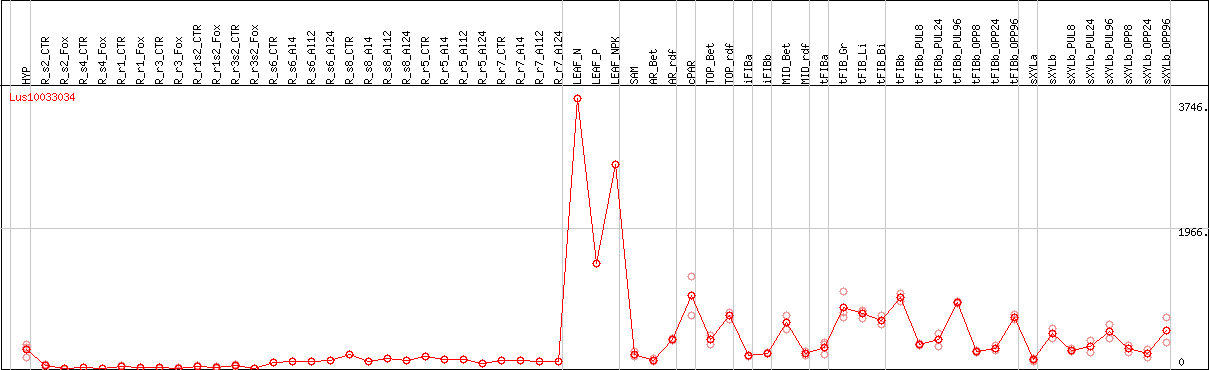
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033034 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.