External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G47870 182 / 3e-55
AS2
ASL29, SCP, LBD27
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
AT3G13850 113 / 3e-29
AS2
LBD22
LOB domain-containing protein 22 (.1)
AT1G72980 108 / 4e-28
AS2
LBD7
LOB domain-containing protein 7 (.1)
AT5G63090 104 / 8e-27
AS2
LOBB, LOB
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
AT3G27650 103 / 8e-27
AS2
LBD25
LOB domain-containing protein 25 (.1)
AT5G66870 106 / 1e-26
AS2
LBD36, ASL1
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
AT1G31320 98 / 9e-25
AS2
LBD4
LOB domain-containing protein 4 (.1)
AT2G30130 97 / 3e-24
AS2
PCK1, LBD12, ASL5
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
AT2G30340 99 / 4e-24
AS2
LBD13
LOB domain-containing protein 13 (.1)
AT2G28500 98 / 5e-24
AS2
LBD11
LOB domain-containing protein 11 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036665
481 / 1e-172
AT3G47870 182 / 2e-55
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Lus10008629
341 / 2e-117
AT3G47870 175 / 2e-52
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Lus10042187
150 / 4e-44
AT3G47870 47 / 4e-06
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Lus10003789
115 / 4e-30
AT3G13850 162 / 7e-49
LOB domain-containing protein 22 (.1)
Lus10002513
110 / 1e-28
AT3G13850 161 / 1e-48
LOB domain-containing protein 22 (.1)
Lus10013510
107 / 2e-27
AT3G13850 143 / 1e-41
LOB domain-containing protein 22 (.1)
Lus10039940
105 / 3e-27
AT5G63090 236 / 5e-80
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Lus10031191
104 / 3e-27
AT1G31320 199 / 2e-66
LOB domain-containing protein 4 (.1)
Lus10031769
103 / 8e-27
AT1G31320 193 / 8e-64
LOB domain-containing protein 4 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G072000
278 / 2e-93
AT3G47870 180 / 7e-55
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Potri.015G066700
270 / 3e-90
AT3G47870 191 / 2e-59
SIDECAR POLLEN, ASYMMETRIC LEAVES 2-like 29, LOB domain-containing protein 27 (.1)
Potri.003G039700
119 / 1e-30
AT3G13850 184 / 6e-56
LOB domain-containing protein 22 (.1)
Potri.001G196400
112 / 5e-29
AT3G13850 184 / 3e-57
LOB domain-containing protein 22 (.1)
Potri.005G097800
104 / 3e-27
AT1G31320 193 / 6e-64
LOB domain-containing protein 4 (.1)
Potri.015G082200
104 / 5e-27
AT5G63090 223 / 4e-75
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Potri.007G066700
103 / 5e-27
AT1G31320 216 / 3e-73
LOB domain-containing protein 4 (.1)
Potri.005G134900
105 / 2e-26
AT5G66870 254 / 8e-84
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Potri.007G039500
105 / 2e-26
AT5G66870 256 / 1e-84
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
Potri.012G083500
103 / 2e-26
AT5G63090 223 / 4e-75
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03195
LOB
Lateral organ boundaries (LOB) domain
Representative CDS sequence
>Lus10033111 pacid=23177040 polypeptide=Lus10033111 locus=Lus10033111.g ID=Lus10033111.BGIv1.0 annot-version=v1.0
ATGACGTTGAACGGTGGCACAAGCCACGCCTGTGCAGCGTGCAAGTATCAGAGACGGAAGTGCACCTCCGAGTGTCAGCTTGCCCCTTACTTCCCTGCAA
ACAGACCTGAAACCTTCAGGAATGCTCACAAGCTATTCGGCGTCAGGAACATCGTCAAGATACTCGAGAAACTGGACCACCACCAGCGCCAGGAGGCCAT
GCAATCCATCATTTGGCAGGCTAACATCCGGGACATCTACCCTGTCCGCGGCTGCCTCGACTACATCTATGCCCTCCAATACAAGATTCACCAGGCCCAG
GAGGAGCTCCATGCTGTCCACTCTCAGCTCCAGGTCTACCGCAATCAACAAGCTACGGATCGCCCTTTCCCCGAGGACGACATGGTAATTACTTCCTCCC
ACCTGCAACTTGGGATGGCTCCTCCTCCTGCTGCAGCAGCCAATAGTAACGCCCTCTCCCTTTTCGGCCACCACAACGAACACCCTCCGGAACAAGTGGT
GGCTCTCCAGTCCCCGGAACTCCAGCATTCCTACTGTATGAGCATCAATAATAATGTGGTGGGAGGAGGAGGCTACAGCTCTGGTTACTTGGATAACAAA
GATAATGCAGTAGGTAATTCTTTCTGGCTTCACCATAATTATGGCATTAACAACGACGAGGTCATCAATTCTTCCAGCATGGGTCTTCAGTCTCAGTTGG
TAGCAGCACCTAATCATCCACTGGGTATCCAGCAGGAATTGGTTCATGACTACAATGAGATGCATCCGTTTTTTGACACAATTGATGACAGGCAATCTTA
CATTGACTCCACCAAAGAGGCTTATGATACAAGCTCGGAAGAGTCTATCAAGGACACAACGCAGTCCCTCGAACACATTGCGGAGAACGAGCTAAAAAGT
GCGGCAGCATGCTTCAGCATCACCAGTGTCAACTGA
AA sequence
>Lus10033111 pacid=23177040 polypeptide=Lus10033111 locus=Lus10033111.g ID=Lus10033111.BGIv1.0 annot-version=v1.0
MTLNGGTSHACAACKYQRRKCTSECQLAPYFPANRPETFRNAHKLFGVRNIVKILEKLDHHQRQEAMQSIIWQANIRDIYPVRGCLDYIYALQYKIHQAQ
EELHAVHSQLQVYRNQQATDRPFPEDDMVITSSHLQLGMAPPPAAAANSNALSLFGHHNEHPPEQVVALQSPELQHSYCMSINNNVVGGGGYSSGYLDNK
DNAVGNSFWLHHNYGINNDEVINSSSMGLQSQLVAAPNHPLGIQQELVHDYNEMHPFFDTIDDRQSYIDSTKEAYDTSSEESIKDTTQSLEHIAENELKS
AAACFSITSVN
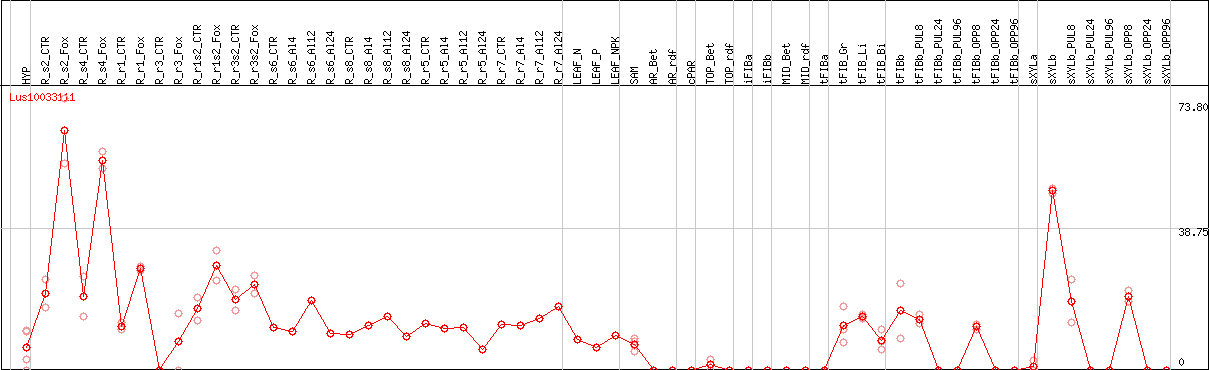
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033111 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.