Lus10033347 [FLAX]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10033347 pacid=23172143 polypeptide=Lus10033347 locus=Lus10033347.g ID=Lus10033347.BGIv1.0 annot-version=v1.0
ATGGATCCAGAGATAGCTAAGACGCAGGAGGAGAGGAAGAGGATGGAGCAGGAGCTCGCCTCCCTTACTTCGCTCAACTACGATAAGGACCTCTACGGCG
GAACAGATAAGGACTCCTACGTCACCTCCATCCCCGTCAACGACGACGAAGACAACGACATCGCCGACAATGAAGTCGCTCGCAAGCTCGCCTCCTATAC
CGCTCCCAAGTCGCTTCTCAAGGAGATGCCTCGCGGCGGCGATGACGATGATGACGGCGGTTTCAAGAAGCCTTCGAGGATTATCGATCGCGAGGATGAT
TACCGGCGCAGGAGATTGAATAGGGTGATATCTCCGGACAGACATGATGCGTTTGCGATGGGGGAGAAGACGCCGGATCCGTCAGTTAGGACTTACGCGG
AAGTTATGAGGGAGGAGGCTCTGAAGAGGGAGAGAGAGGCGACGTTGAGGGAAATTTCAAAGAAGAAGAAGCAGGAGGAGGAAGATGCTAAAGAAGCTAA
GGACTCTGAAGTAACTAAGAAGGAGCCAGCTGCACCTGCTACAACTGCAGCAGCTCCGAAGAGGAGGAATAGATGGGACCAGTCACAGGAAGGTGAAGCT
TCTGCTGGGGCTGCTAAGAAAGCTAAGACGAGCTCTGATTGGGATTTACCAGACGCAACTCCTGGGATTGGAAGGTGGGATGCTGCTACTCCCACACCTG
GTAGAGTTGGGGATGCGACCCCCTCGATGGGGAGGAGGAATAGATGGGATGAAACCCCAACTCCCGGAAGGGTAGCTGATTCGGATGCAACTCCTTCTGG
TGGTGTTACTCCTGGTGCTACACCTGCTGGGGTGACTTGGGATGCAACTCCCAAAGGGTTGGCCACACCGACTCCTAAAAAGCAAAGATCACGATGGGAT
GAAACTCCTGCAACTATGGGAAGTGCGACTCCGATGGCTGGGGCTACACCTGCTGCTGCATATACCCCTGGTGTTACTCCTGTTGGTGGGATTGAATTGG
CCACCCCAACTCCGAGTCTCATTCAACGTGGTGCTCTTACTCCGGAGCAGTATAATCTATTGAGATGGGAGAAGGATATTGAAGAGAGGAATAGACCTTT
GACTGATGAGGAACTAGATGCTATATTTCCTCTGGAAGGATATAAGATTTTGGAGCCACCGGCATCTTATGTTCCCATTAGGACTCCTGCCAGGAAATTG
TTGGCTACACCCACTCCTATGGGTACTCCGCTTTATGCTATACCTGAAGAGAATAGGGGGCAGCAATATGATGTGCCGAAAGAAGCTCCTGGTGGGTTGC
CCCATATGAAGCCGGAGGATTATCAGTACTTTGGAGCCTTGTTAAATGAGGACGATGAGGAAGAGTTGACACCTGATGAACAAAAAGAGCGGAAGATTAT
GAAGCTGTTGTTGAAGGTTAAGAATGGAACACCGCCTCAGCGTAAGACTGCTTTGCGACAGTTGACAGATAAGGCAAGAGAGTTTGGTGCTGGCCCACTG
TTCAACAGGATTCTGCCTTTGCTTATGCAGCCCACTTTGGAGGATCAGGAGAGACATCTTCTAGTCAAGGTGATTGATAGGGTTTTGTATAAGTTGGATG
AGCTGGTTAGGCCTTACGTGCACAAGATTCTTGTTGTTATTGAGCCGTTGCTTATTGATGAAGATTATTATGCCCGTGTTGAAGGAAGGGAGATCATTTC
AAATTTGAGTAGAGCTGCTGGTTTGGCAACTATGATTGCTGCTATGCGTCCTGATATTGATAATATTGATGAATATGTTAGGAATACAACTGCTAGAGCT
TTCAGTGTTGTGGCTTCTGCTCTTGGTATTCCAGCACTGCTGCCTTTTCTGAAAGCTGTGTGTCAGAGTAAGAAATCGTGGCAAGCTCGTCACACTGGTA
TCAAGATTGTGCAGCAGATTGCTATTCTCATTGGGTGTGCTGTTCTTCCTCATTTAAGGTCTCTGGTAGAAATTATAGAGCATGGACTAATTGACGAGAA
TCCGAAAGTGAGAACTATCACTGCATTGTCTTTGGCTGCCCTTGCTGAGGCTTCTGCACCTTATGGTATTGAAAGCTTCGACTCTGTTCTGAAACCCTTG
TGGAAAGGAATAAGGTCACACCGCGGAAAGGTCTTGGCTGCCTTCTTGAAGGCAATTGGTTTTATCATTCCTCTGATGGATGCTATGTATGCCTCTTACT
ACACTAAGGAAGTGATGTACATTCTCATCCGTGAGTTTCAGTCTCCTGATGAAGAAATGAAGAAGATTGTTCTGAAAGTGGTGAAGCAATGTGTCAGCAC
TGAGGGTGTGGAACCAGATTACATCCGTAATGATATTCTTCCAGAATTCTTTAAGAACTTCTGGGTTAGAAGGATGGCATTGGATAGAAGAAACTACAAG
CAACTTGTAGAAACCACAGTGGAGATTGCTAACAAGGTTGGTGTTAAAGATATTGTTGGGAGGATTGTTGAGGATCTTAAAGACGAGAGTGAGCCTTATC
GTCGGATGGTTATGGAAACAATCGAGAAGGTGGTTACGAACTTGGGTTCTTCTGACATTGATGCTAGGTTGGAAGAGCTTTTGATAGATGGTATTCTGTA
TGCTTTCCAAGAACAGACTAGTGATGATGCCAATGTCATGCTTAATGGGTTCGGAGCAGTTGTCAACTCTCTTGGGCAGAGAGTGAAACCTTATCTCCCT
CAGATTTGTGGTACCATAAAGTGCCGGCTGAACAACAAAAGTGCGAAGGTGAGACAACAAGCAGCTGATCTTATTTCTAGGATTGCTGTTGTGATGAAGC
AGTGCCATGAGGAGCAGCTTATGGGTCATCTTGGTGTTGTCTTGTACGAGTATTTGGGAGAAGAATACCCTGAAGTGTTGGGATCAATCCTGGGAGCTCT
GAAGGCTATTGTCAATGTGATTGGTATGACAAAAATGACTCCTCCCATCAAGGACTTGCTTCCAAGGCTGACACCAATTTTGAAGAATAGACATGAGAAA
GTCCAGGAGAACTGCATTGATCTTGTTGGCCGTATTGCTGATCGTGGTGCTGAGTTTGTCCCAGCTAGAGAATGGATGAGGATCTGCTTTGAACTGCTTG
AGATGCTGAAGGCGCACAAGAAGGGTATTCGTAGAGCTACCGTGAACACTTTCGGGTATATCGCCAAAGCCATCGGTCCACAAGATGTTCTTGCAACTTT
GCTGAACAATCTCAAGGTCCAGGAGCGCCAGAATCGTGTGTGTACCACTGTTGCTATTGCTATAGTTGCAGAAACTTGCTCGCCCTTCACAGTGCTTCCT
GCCTTGATGAACGAGTACCGTGTTCCAGAGCTCAATGTCCAGAATGGTGTTTTGAAGTCCCTATCTTTTCTGTTCGAGTACATTGGCGAAATGGGTAAAG
ACTACATCTATGCAGTAACGCCATTGCTCGAGGATGCCCTCATGGACAGGGACTTGGTTCACAGGCAGACAGCAGCTTCAGCTGTCAAGCATATGGCACT
TGGCGTTGCCGGGTTGGGTTGCGAGGATGCTCTAGTTCACCTGCTCAACTTTGTGTGGCCGAACATATTTGAGACTTCTCCTCACGTGATAAATGCTGTA
ATGGAAGCCATAGAAGGAATGAGGGTAGCTCTGGGTGCTGCCGTTGTGTTGAACTATTGTCTGCAGGGGCTTTTCCACCCAGCTCGGAAAGTGCGGGAAG
TGTACTGGAAGATCTACAACTCTCTTTACATTGGCGCACAGGACACGCTTGTGGCTGCTTATCCAGTGCTGGATGATGAGGAGAACAACGTATATAGCAG
GCCTGAGTTGGTGATGTTCGTATGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Lus10033347 pacid=23172143 polypeptide=Lus10033347 locus=Lus10033347.g ID=Lus10033347.BGIv1.0 annot-version=v1.0
MDPEIAKTQEERKRMEQELASLTSLNYDKDLYGGTDKDSYVTSIPVNDDEDNDIADNEVARKLASYTAPKSLLKEMPRGGDDDDDGGFKKPSRIIDREDD
YRRRRLNRVISPDRHDAFAMGEKTPDPSVRTYAEVMREEALKREREATLREISKKKKQEEEDAKEAKDSEVTKKEPAAPATTAAAPKRRNRWDQSQEGEA
SAGAAKKAKTSSDWDLPDATPGIGRWDAATPTPGRVGDATPSMGRRNRWDETPTPGRVADSDATPSGGVTPGATPAGVTWDATPKGLATPTPKKQRSRWD
ETPATMGSATPMAGATPAAAYTPGVTPVGGIELATPTPSLIQRGALTPEQYNLLRWEKDIEERNRPLTDEELDAIFPLEGYKILEPPASYVPIRTPARKL
LATPTPMGTPLYAIPEENRGQQYDVPKEAPGGLPHMKPEDYQYFGALLNEDDEEELTPDEQKERKIMKLLLKVKNGTPPQRKTALRQLTDKAREFGAGPL
FNRILPLLMQPTLEDQERHLLVKVIDRVLYKLDELVRPYVHKILVVIEPLLIDEDYYARVEGREIISNLSRAAGLATMIAAMRPDIDNIDEYVRNTTARA
FSVVASALGIPALLPFLKAVCQSKKSWQARHTGIKIVQQIAILIGCAVLPHLRSLVEIIEHGLIDENPKVRTITALSLAALAEASAPYGIESFDSVLKPL
WKGIRSHRGKVLAAFLKAIGFIIPLMDAMYASYYTKEVMYILIREFQSPDEEMKKIVLKVVKQCVSTEGVEPDYIRNDILPEFFKNFWVRRMALDRRNYK
QLVETTVEIANKVGVKDIVGRIVEDLKDESEPYRRMVMETIEKVVTNLGSSDIDARLEELLIDGILYAFQEQTSDDANVMLNGFGAVVNSLGQRVKPYLP
QICGTIKCRLNNKSAKVRQQAADLISRIAVVMKQCHEEQLMGHLGVVLYEYLGEEYPEVLGSILGALKAIVNVIGMTKMTPPIKDLLPRLTPILKNRHEK
VQENCIDLVGRIADRGAEFVPAREWMRICFELLEMLKAHKKGIRRATVNTFGYIAKAIGPQDVLATLLNNLKVQERQNRVCTTVAIAIVAETCSPFTVLP
ALMNEYRVPELNVQNGVLKSLSFLFEYIGEMGKDYIYAVTPLLEDALMDRDLVHRQTAASAVKHMALGVAGLGCEDALVHLLNFVWPNIFETSPHVINAV
MEAIEGMRVALGAAVVLNYCLQGLFHPARKVREVYWKIYNSLYIGAQDTLVAAYPVLDDEENNVYSRPELVMFV
|
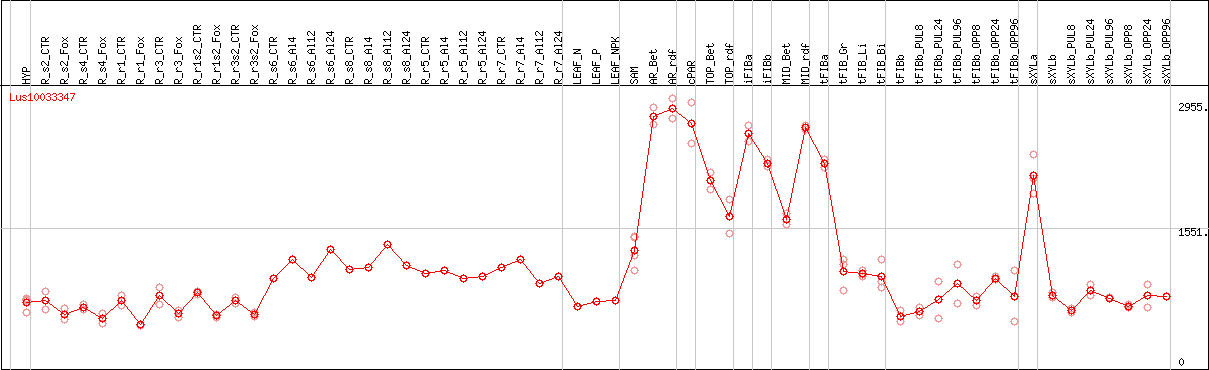
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033347 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
