External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G00120 142 / 1e-42
bHLH
IND1, GT140, bHLH040, IND, EDA33
INDEHISCENT, EMBRYO SAC DEVELOPMENT ARREST 33, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G09750 132 / 1e-38
bHLH
HEC3, bHLH043
HECATE 3, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G21330 123 / 9e-34
bHLH
bHLH087
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G67060 117 / 1e-32
bHLH
HEC1, bHLH088
HECATE 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G50330 117 / 1e-32
bHLH
HEC2, bHLH037
HECATE 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G43175 82 / 5e-19
bHLH
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G01310 84 / 9e-19
bHLH
APTX, bHLH140
APRATAXIN-like (.1)
AT1G27740 81 / 2e-18
bHLH
bHLH054, ROOT HAIR DEFECTIVE 6-LIKE 4(RSL4)
root hair defective 6-like 4 (.1)
AT2G14760 81 / 3e-18
bHLH
bHLH084
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
AT4G33880 81 / 4e-18
bHLH
RSL2, bHLH085
ROOT HAIR DEFECTIVE 6-LIKE 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10042647
141 / 1e-41
AT3G50330 163 / 1e-49
HECATE 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10005710
140 / 2e-41
AT3G50330 160 / 1e-48
HECATE 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10012670
125 / 9e-36
AT3G50330 158 / 3e-48
HECATE 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10003500
107 / 2e-27
AT3G21330 192 / 5e-58
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10009745
103 / 3e-26
AT3G21330 112 / 2e-29
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10002213
87 / 3e-21
AT5G01310 108 / 8e-28
APRATAXIN-like (.1)
Lus10003386
89 / 1e-20
AT5G01310 107 / 1e-24
APRATAXIN-like (.1)
Lus10001068
82 / 3e-18
AT4G33880 179 / 2e-53
ROOT HAIR DEFECTIVE 6-LIKE 2 (.1)
Lus10004315
81 / 6e-18
AT4G33880 179 / 3e-53
ROOT HAIR DEFECTIVE 6-LIKE 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G060900
160 / 1e-49
AT4G00120 151 / 7e-46
INDEHISCENT, EMBRYO SAC DEVELOPMENT ARREST 33, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.007G108000
152 / 2e-46
AT4G00120 150 / 3e-45
INDEHISCENT, EMBRYO SAC DEVELOPMENT ARREST 33, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.007G044600
137 / 2e-40
AT5G67060 150 / 7e-45
HECATE 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.005G138900
119 / 5e-33
AT3G50330 134 / 2e-38
HECATE 2, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.014G027300
117 / 3e-32
AT5G67060 141 / 6e-41
HECATE 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G125000
117 / 3e-32
AT5G67060 137 / 2e-39
HECATE 1, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G191800
120 / 6e-32
AT3G21330 230 / 2e-71
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.006G102600
92 / 1e-23
AT5G01310 112 / 4e-29
APRATAXIN-like (.1)
Potri.016G120800
92 / 2e-23
AT5G01310 109 / 3e-28
APRATAXIN-like (.1)
Potri.001G294300
82 / 3e-18
AT4G33880 164 / 2e-47
ROOT HAIR DEFECTIVE 6-LIKE 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Lus10033488 pacid=23143305 polypeptide=Lus10033488 locus=Lus10033488.g ID=Lus10033488.BGIv1.0 annot-version=v1.0
ATGGACATCAACCAATCCCATCCATGGATCAATCATGATCAACCACAAACCGAGCCAATATTCGATCCTTTCTCATCAACCTTTGGCCTCGACCACAACT
TGATCATCAACCACCAACAAGAAGATGGAGGAGATCAAGAAGAGGAACTGGATGGAGATCAAGAGGAAGAGGAGGAGCTTGGAGCACTCAAGGAGATGAT
GTACAGGATGGCGGCGATGCAACCCGTAGAAATCGACCCGACTTCCATCAGAAAGCCGGGCAGAAGGAACGTCCGGATCAGCGACGACCCGCAGAGCGTG
GCAGCCCGCAACCGCCGCCAGAGGATCAGTGAGAAGATCAGGATCCTCCAGCGGCTTGTCCCTGGTGGGACCAAGATGGACACTGCCTCCATGTTGGAGG
AGGCGGTCCGCTACCTCAAGTTCTTGAAGCGTCAAATTCGCCTCCTCCAGCAGACTGATAATCACATTACCACTAGATTAACCCCTTCTCTCGACGATGG
AGGGGGGTTCGGGATTGCTCCTGCTAATGGAGGTGGGATTCACATCGGTGGTAGCAACGTCGGCGGCGGCGATGAACTCGGGTGTTTTAGTAATCATCAT
GCATGA
AA sequence
>Lus10033488 pacid=23143305 polypeptide=Lus10033488 locus=Lus10033488.g ID=Lus10033488.BGIv1.0 annot-version=v1.0
MDINQSHPWINHDQPQTEPIFDPFSSTFGLDHNLIINHQQEDGGDQEEELDGDQEEEEELGALKEMMYRMAAMQPVEIDPTSIRKPGRRNVRISDDPQSV
AARNRRQRISEKIRILQRLVPGGTKMDTASMLEEAVRYLKFLKRQIRLLQQTDNHITTRLTPSLDDGGGFGIAPANGGGIHIGGSNVGGGDELGCFSNHH
A
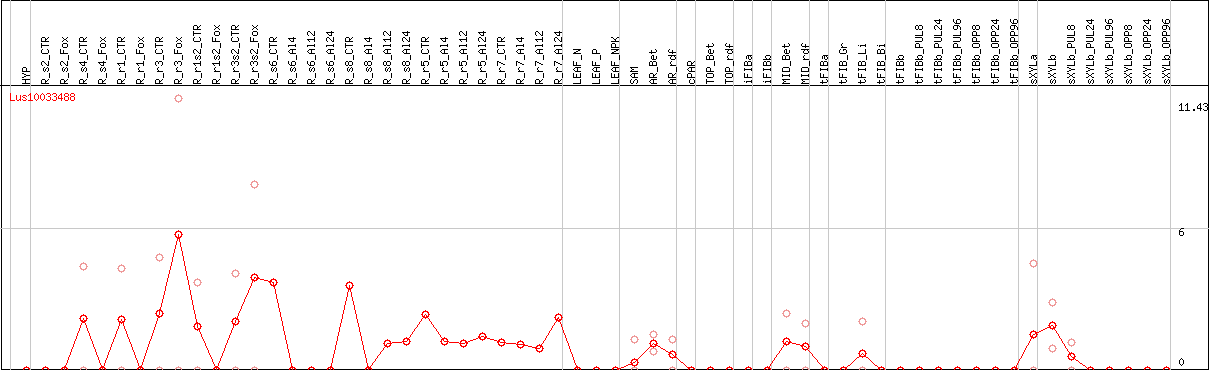
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033488 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.