External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G28010 112 / 3e-30
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G53700 85 / 1e-20
MEE40
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G22470 77 / 6e-18
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G05670 77 / 1e-17
Pentatricopeptide repeat (PPR-like) superfamily protein (.1), Pentatricopeptide repeat (PPR-like) superfamily protein (.2)
AT3G60050 77 / 1e-17
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G13040 77 / 1e-17
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT5G41170 77 / 1e-17
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT3G22670 76 / 2e-17
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G01110 76 / 3e-17
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G64320 75 / 6e-17
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017684
197 / 7e-61
AT4G28010 596 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10032648
84 / 6e-21
AT3G16890 225 / 1e-69
pentatricopeptide (PPR) domain protein 40 (.1)
Lus10043104
82 / 1e-19
AT3G16890 720 / 0.0
pentatricopeptide (PPR) domain protein 40 (.1)
Lus10000364
82 / 1e-19
AT1G22960 619 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10039056
81 / 5e-19
AT5G64320 832 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10034616
80 / 9e-19
AT1G22960 617 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10006373
80 / 9e-19
AT3G53700 1012 / 0.0
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10012328
80 / 1e-18
AT3G53700 1010 / 0.0
maternal effect embryo arrest 40, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10003433
80 / 1e-18
AT1G12700 396 / 2e-129
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G069600
135 / 2e-38
AT4G28010 661 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G271200
82 / 2e-19
AT1G62930 475 / 3e-161
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.006G049400
80 / 1e-18
AT1G79540 787 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.019G021200
80 / 1e-18
AT1G12700 491 / 2e-166
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.010G098400
79 / 2e-18
AT2G02150 924 / 0.0
EMBRYO DEFECTIVE 2794, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.015G105400
78 / 4e-18
AT5G61990 773 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G050500
78 / 5e-18
AT1G12700 523 / 2e-178
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.005G046100
77 / 7e-18
AT1G12700 512 / 1e-173
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.006G242200
77 / 9e-18
AT1G62930 472 / 4e-160
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G047400
77 / 1e-17
AT1G63080 341 / 1e-110
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10033639 pacid=23143436 polypeptide=Lus10033639 locus=Lus10033639.g ID=Lus10033639.BGIv1.0 annot-version=v1.0
ATGCGAGACTCGAACTATGTCCCTGATGTTGTCTCATTTAATACGATGATTGATGGATTGTTGAAAGCCGGAGACGTTCAGTATGCCAAAGAGCTGCTTA
ATGAAATGGTGCAAATGGGTCAGATGCCGGATGTTTTTACATATACTGTGTTGATTAATCGATTGTCGAAACTCGGACAGGTGGACGAGGCTAAGGGAAT
CTTTGAGAGGATGGTTTCGTTAGGTGTTAAACCAGACAGCCACGTATATGATTCGTTGTTAAAACGAGTGCTTTCGAAAGGGGAAGCTGAAGAGACCATT
GATTTGCTGCGTTGA
AA sequence
>Lus10033639 pacid=23143436 polypeptide=Lus10033639 locus=Lus10033639.g ID=Lus10033639.BGIv1.0 annot-version=v1.0
MRDSNYVPDVVSFNTMIDGLLKAGDVQYAKELLNEMVQMGQMPDVFTYTVLINRLSKLGQVDEAKGIFERMVSLGVKPDSHVYDSLLKRVLSKGEAEETI
DLLR
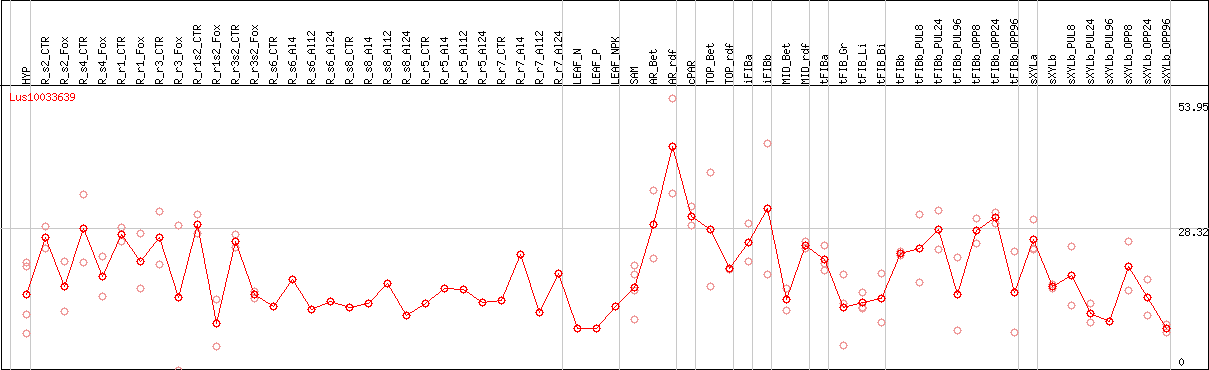
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033639 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.