External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G03790 239 / 2e-76
C3HZnF
SOM
SOMNUS, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
AT5G44260 222 / 5e-70
C3HZnF
Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
AT4G29190 205 / 9e-64
C3HZnF
AtOZF2
Arabidopsis thaliana oxidation-related zinc finger 2, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
AT2G19810 204 / 2e-63
C3HZnF
AtOZF1
Oxidation-related Zinc Finger 1, CCCH-type zinc finger family protein (.1)
AT2G25900 203 / 2e-63
C3HZnF
ATTZF1, ATCTH
A. THALIANA TANDEM ZINC FINGER PROTEIN 1, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1.2)
AT2G41900 181 / 1e-51
C3HZnF
OXS2
OXIDATIVE STRESS 2, CCCH-type zinc finger protein with ARM repeat domain (.1)
AT5G12850 180 / 2e-51
C3HZnF
PEI1
CCCH-type zinc finger protein with ARM repeat domain (.1)
AT5G58620 176 / 2e-50
C3HZnF
zinc finger (CCCH-type) family protein (.1)
AT5G07500 163 / 9e-49
C3HZnF
PEI1
Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
AT3G55980 162 / 2e-45
C3HZnF
ATSZF1
salt-inducible zinc finger 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017708
355 / 4e-123
AT1G03790 252 / 7e-82
SOMNUS, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
Lus10018621
226 / 5e-72
AT1G03790 245 / 3e-78
SOMNUS, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
Lus10012922
202 / 3e-62
AT2G19810 387 / 9e-134
Oxidation-related Zinc Finger 1, CCCH-type zinc finger family protein (.1)
Lus10035006
202 / 4e-62
AT2G19810 380 / 2e-131
Oxidation-related Zinc Finger 1, CCCH-type zinc finger family protein (.1)
Lus10019709
177 / 4e-50
AT2G41900 759 / 0.0
OXIDATIVE STRESS 2, CCCH-type zinc finger protein with ARM repeat domain (.1)
Lus10028970
168 / 2e-47
AT2G40140 615 / 0.0
\(SALT-INDUCIBLE ZINC FINGER 2, zinc finger (CCCH-type) family protein (.1), zinc finger (CCCH-type) family protein (.2)
Lus10007493
166 / 1e-46
AT2G40140 614 / 0.0
\(SALT-INDUCIBLE ZINC FINGER 2, zinc finger (CCCH-type) family protein (.1), zinc finger (CCCH-type) family protein (.2)
Lus10040662
157 / 4e-45
AT5G58620 341 / 2e-112
zinc finger (CCCH-type) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G138300
237 / 7e-76
AT1G03790 233 / 2e-73
SOMNUS, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
Potri.017G013400
229 / 9e-73
AT1G03790 265 / 2e-85
SOMNUS, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
Potri.006G234300
213 / 2e-66
AT4G29190 306 / 4e-102
Arabidopsis thaliana oxidation-related zinc finger 2, Zinc finger C-x8-C-x5-C-x3-H type family protein (.1)
Potri.018G058600
205 / 1e-63
AT2G19810 314 / 4e-105
Oxidation-related Zinc Finger 1, CCCH-type zinc finger family protein (.1)
Potri.001G266700
179 / 6e-51
AT2G41900 708 / 0.0
OXIDATIVE STRESS 2, CCCH-type zinc finger protein with ARM repeat domain (.1)
Potri.006G053900
178 / 8e-51
AT2G41900 791 / 0.0
OXIDATIVE STRESS 2, CCCH-type zinc finger protein with ARM repeat domain (.1)
Potri.006G053800
177 / 2e-50
AT2G41900 817 / 0.0
OXIDATIVE STRESS 2, CCCH-type zinc finger protein with ARM repeat domain (.1)
Potri.016G053700
177 / 3e-50
AT2G41900 820 / 0.0
OXIDATIVE STRESS 2, CCCH-type zinc finger protein with ARM repeat domain (.1)
Potri.009G061000
174 / 3e-49
AT2G41900 672 / 0.0
OXIDATIVE STRESS 2, CCCH-type zinc finger protein with ARM repeat domain (.1)
Potri.001G252600
173 / 8e-49
AT5G58620 572 / 0.0
zinc finger (CCCH-type) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0537
CCCH_zf
PF00642
zf-CCCH
Zinc finger C-x8-C-x5-C-x3-H type (and similar)
Representative CDS sequence
>Lus10033665 pacid=23143386 polypeptide=Lus10033665 locus=Lus10033665.g ID=Lus10033665.BGIv1.0 annot-version=v1.0
ATGACCACCGCCGCCGCCTCCATTTACTCCGATCACCACCCATCTTCCCCCACTGCCAAACTCCCTCCTAGAAAGCTCCTCGTCAACAGAAACAACACCG
TTGACTTCCCTCAGCTGGACGACATGTCGTCTTTGGCGCATTTCTACAAACACCTTCCGTGCAACAAAGCAGACCAGCTCGAGGAAGAAGAAGAAGATGG
CGATCCTTACTCGTCAGATCAATTCAGGATGTATGATTTCAAAGTCAGGAGGTGTACAAGGAGCCGTAGCCACGACTGGACTGACTGCCCCTTTGCCCAC
CCGGGCGAGAAGGCCCGCCGCCGTGATCCGAGGAAGTTTCACTACACGGGCGCGGTTTGCCCTGAGTTCCGACGCGGCGGAGGGTGTAGCCGCGGCGATA
ACTGCGAGTTCGCTCATGGAGTTTTCGAGTGCTGGCTGCATCCGTTGAGGTACAGAACTGAAGCTTGTAAAGATGGGAAGGGCTGTAAGAGGAAGGTCTG
CTTCTTCGCTCACTCCCCCCGCCAGCTCCGCCTCCTTCCGCCGGAGAAGTCGTCGCCGACCCCCCCCCCCCCCCCCCTTCCGCGGGGGGGAGGTCCGCGC
AGGAGAGGGGGGTGGGGTTTTGGTAAATGTCGCCCCCCGCCATCATTCCTGCGGATGCTTGGTCTGCCGTGGCAGCGGCGGCGATTCCTCATCTCCGACA
TCTACTCTCATCGGGCTGTCCCATCTCTCGCCGCCGCTGTCCCCTTCTTTTTCCTCGTCACCGCCTGTGTCACCGGTGAGCCACCACCGTCCTTCAGCGG
CGGGGGCGACCGGGTTTTCTCACCGACTGACGAGGAAGCCGCCGCCGCCGCTGAGGACCCCCCCCCCCGCCTCTGTTGCGGCGGCAGCTGCAGCGGTGTC
GTCGTCTCCGACTTCAGGCCTCACCGTTAA
AA sequence
>Lus10033665 pacid=23143386 polypeptide=Lus10033665 locus=Lus10033665.g ID=Lus10033665.BGIv1.0 annot-version=v1.0
MTTAAASIYSDHHPSSPTAKLPPRKLLVNRNNTVDFPQLDDMSSLAHFYKHLPCNKADQLEEEEEDGDPYSSDQFRMYDFKVRRCTRSRSHDWTDCPFAH
PGEKARRRDPRKFHYTGAVCPEFRRGGGCSRGDNCEFAHGVFECWLHPLRYRTEACKDGKGCKRKVCFFAHSPRQLRLLPPEKSSPTPPPPPLPRGGGPR
RRGGWGFGKCRPPPSFLRMLGLPWQRRRFLISDIYSHRAVPSLAAAVPFFFLVTACVTGEPPPSFSGGGDRVFSPTDEEAAAAAEDPPPRLCCGGSCSGV
VVSDFRPHR
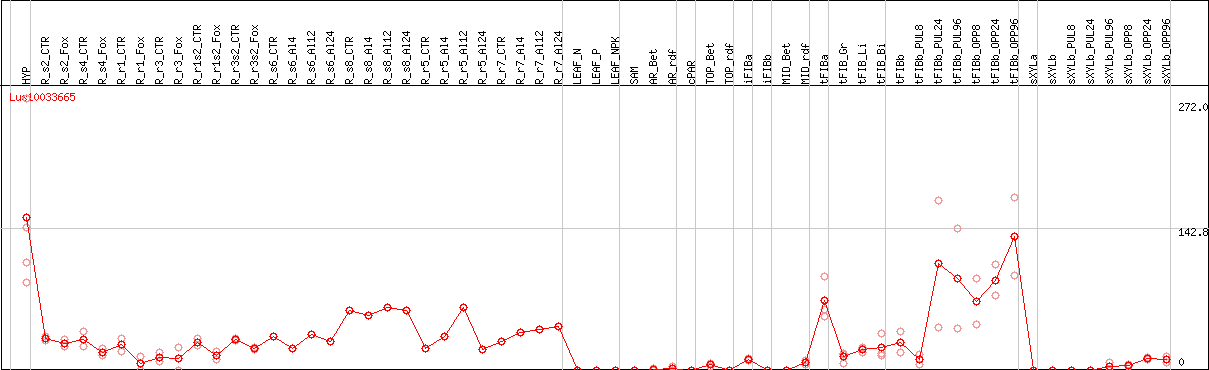
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033665 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.