External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G04960 218 / 2e-70
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
AT1G27000 134 / 5e-38
Protein of unknown function (DUF1664) (.1)
AT2G02730 120 / 9e-33
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
AT1G24265 70 / 8e-14
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2), Protein of unknown function (DUF1664) (.3)
AT1G24267 56 / 5e-09
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014626
332 / 5e-115
AT1G04960 318 / 3e-108
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Lus10036718
146 / 1e-42
AT1G27000 272 / 1e-90
Protein of unknown function (DUF1664) (.1)
Lus10037210
144 / 8e-41
AT1G27000 252 / 8e-82
Protein of unknown function (DUF1664) (.1)
Lus10013269
136 / 1e-38
AT1G27000 254 / 8e-84
Protein of unknown function (DUF1664) (.1)
Lus10030793
123 / 8e-32
AT1G69830 1173 / 0.0
alpha-amylase-like 3 (.1)
Lus10003127
75 / 1e-15
AT1G24265 202 / 6e-63
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2), Protein of unknown function (DUF1664) (.3)
Lus10011347
73 / 9e-15
AT1G24265 226 / 2e-71
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2), Protein of unknown function (DUF1664) (.3)
Lus10027272
52 / 8e-08
AT2G27410 66 / 9e-13
Domain of unknown function (DUF313) (.1)
Lus10027284
50 / 3e-07
AT2G27410 67 / 2e-12
Domain of unknown function (DUF313) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G219300
243 / 5e-80
AT1G04960 325 / 7e-111
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Potri.014G160800
238 / 2e-78
AT1G04960 309 / 6e-105
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Potri.008G149000
131 / 9e-37
AT1G27000 327 / 3e-112
Protein of unknown function (DUF1664) (.1)
Potri.010G092700
127 / 2e-35
AT1G27000 331 / 1e-113
Protein of unknown function (DUF1664) (.1)
Potri.003G169200
77 / 3e-16
AT1G24267 214 / 5e-67
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
Potri.001G058700
77 / 4e-16
AT1G24267 211 / 1e-65
Protein of unknown function (DUF1664) (.1), Protein of unknown function (DUF1664) (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF07889
DUF1664
Protein of unknown function (DUF1664)
Representative CDS sequence
>Lus10033807 pacid=23172863 polypeptide=Lus10033807 locus=Lus10033807.g ID=Lus10033807.BGIv1.0 annot-version=v1.0
ATGTGGTGGAGCTGTGGAGTTTCAGGTCCCCTAATTCAGGAGAGCTCTGTTTTACATTGGTTACTTTGGGAGGAAGACAAGCAGCTGCTGGTGAGAAGGA
ACAACAAGGATATGATCAAATATGGAGCACGCCCTAATCTTCAGAATGGGGCCATTGGTCATCTCCTTGGAGCAGCGGGTGCTGCTGAAGCAATCTTCAG
TATTCGCCAATTGGCTCAAGAAATTAAGGAGATATCCATGTCCAATCCTGTAACCATCTTCAATGGCAATTCTGAAAGCGGAAGCTTAGCATCTTATATC
ATGCCAGCTGCAGCAGTGGGAGCAATGGGTTATTGTTACATGTGGTGGAAGGCAAAGCAGGGTTGGTCATTATCAGATGTAATGTTTGTGACTAAGAACA
ATATGAAAAATGCAGTCGAATCTGTCTCAAAACAATTGGAGCATCTATCTGATTCACTTGCTAATACAAAGAAGCATCTGAGTAATAGGCTCGAAAACTT
GGACTGGAAAGTGGACGAGCAAATAGAGACATCTAAGCAAATTGCTAGTAATGTGGATGATTTGAATTCAAACATTTCCCAAATTGGATTTGATGTTGGA
ACAATTCATCATATGATAGCTGGCCTGGATGTAACAAACTCAGGTCTTTGGTACCTATGCCAGGTCGCTGATGGAATGAAAGATGGGTTTGGAAAAATAG
TTCCCAAGGTCAGATAG
AA sequence
>Lus10033807 pacid=23172863 polypeptide=Lus10033807 locus=Lus10033807.g ID=Lus10033807.BGIv1.0 annot-version=v1.0
MWWSCGVSGPLIQESSVLHWLLWEEDKQLLVRRNNKDMIKYGARPNLQNGAIGHLLGAAGAAEAIFSIRQLAQEIKEISMSNPVTIFNGNSESGSLASYI
MPAAAVGAMGYCYMWWKAKQGWSLSDVMFVTKNNMKNAVESVSKQLEHLSDSLANTKKHLSNRLENLDWKVDEQIETSKQIASNVDDLNSNISQIGFDVG
TIHHMIAGLDVTNSGLWYLCQVADGMKDGFGKIVPKVR
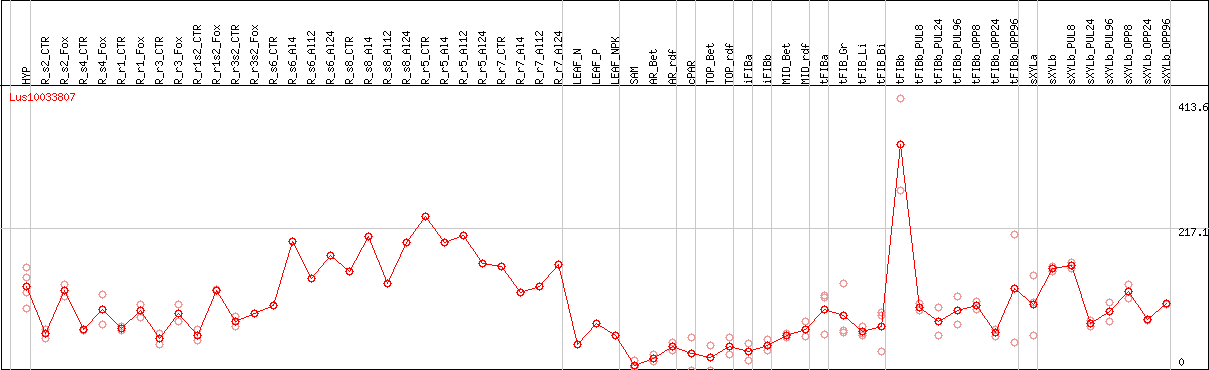
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033807 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.