External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G06240 44 / 7e-06
F-box family protein (.1)
AT4G12560 44 / 9e-06
CPR1, CPR30
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018968
180 / 3e-58
ND 38 / 0.004
Lus10018972
133 / 3e-37
AT5G36930 192 / 3e-51
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10033825
74 / 2e-16
AT5G36930 400 / 3e-123
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10001105
62 / 4e-12
AT3G06240 109 / 3e-26
F-box family protein (.1)
Lus10013988
62 / 4e-12
AT3G49180 471 / 4e-160
ROOT INITIATION DEFECTIVE 3, Transducin/WD40 repeat-like superfamily protein (.1)
Lus10009424
56 / 4e-10
AT3G06240 73 / 9e-14
F-box family protein (.1)
Lus10028961
55 / 1e-09
AT4G12560 81 / 2e-16
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Lus10015408
54 / 2e-09
AT3G16210 64 / 6e-11
F-box family protein (.1)
Lus10027824
53 / 5e-09
AT3G06240 78 / 2e-15
F-box family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G035500
54 / 1e-09
AT4G12560 271 / 3e-87
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.006G012900
50 / 6e-08
AT4G12560 144 / 2e-39
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.003G145950
47 / 4e-07
AT4G12560 113 / 7e-28
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.006G013000
46 / 1e-06
AT4G12560 250 / 5e-79
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.012G099733
43 / 1e-05
AT3G16210 92 / 2e-20
F-box family protein (.1)
Potri.008G006900
43 / 2e-05
AT4G12560 95 / 3e-21
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.006G011400
41 / 5e-05
AT4G12560 142 / 2e-38
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.001G262900
40 / 9e-05
AT4G12560 107 / 5e-26
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.010G207500
39 / 0.0003
AT4G12560 94 / 3e-21
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.010G154500
38 / 0.0006
AT4G12560 139 / 7e-37
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
PFAM info
Representative CDS sequence
>Lus10033824 pacid=23172878 polypeptide=Lus10033824 locus=Lus10033824.g ID=Lus10033824.BGIv1.0 annot-version=v1.0
ATGGATGCATGGGCCTCTGACATCATCCTCTGGAATCCGTCAACTTCCGAAACCATGATCCTGCCGCGTTCTCCCTTAATGGACGCAAAAATGGATGCAA
CAGTGAAGGACAGAGACCGTTTCTTTACAGAACCGATTGGGTTCGGGTTCGACCCACTTACGGATGACTACAAAGTGCTAAGGTGGATTAACTACATTGG
AGATCCTTATTACACAGAAGATGGACATTCTTTTTCCAAAGGATGGGATGCTGCTGAGGTTTACAGCTTGAAGGATCAATCATGGACACCACTTGCGACT
TGTAAGTCCTCCGTGGGTGCTAACCCATTCCGTCAGCACCATACATCTTGGTACGAGATGCTTTACTGA
AA sequence
>Lus10033824 pacid=23172878 polypeptide=Lus10033824 locus=Lus10033824.g ID=Lus10033824.BGIv1.0 annot-version=v1.0
MDAWASDIILWNPSTSETMILPRSPLMDAKMDATVKDRDRFFTEPIGFGFDPLTDDYKVLRWINYIGDPYYTEDGHSFSKGWDAAEVYSLKDQSWTPLAT
CKSSVGANPFRQHHTSWYEMLY
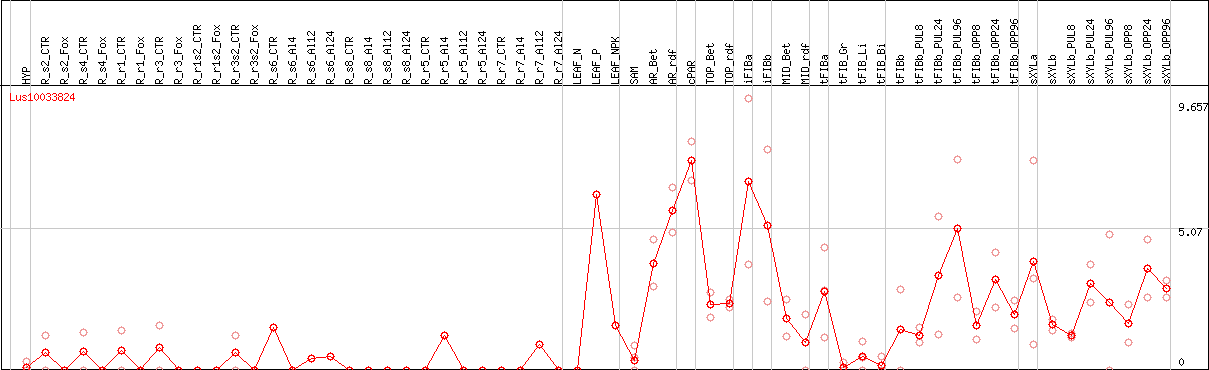
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10033824 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.