Lus10033945 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10033945 pacid=23154722 polypeptide=Lus10033945 locus=Lus10033945.g ID=Lus10033945.BGIv1.0 annot-version=v1.0
ATGACGAAGCATCGCTGGAAGGAGTCTGTCAAATCCTTTTTTGGGAGCCACATTGATCCAGAGAAAGACGAGCAGCTCCAAGGCAGTAAAGAAGAATTCG
ACGATAAATTGAAAAGGATCTTGGAGCTAATTAACGAAGAAGATTCTGAAGTGGAAAGGTCCAAAAGAAAGCCTCTTGTGGACTTAATTGAGGATTTTCA
CAATCAGTATGAATCGCTATATTCACGATACGATCATCTTACAGGAGAGCTGAGGAAGAAATTTTATGGGAAGAAGAGGGGGTCGGACTCCTCTTCTTCA
TCTAGCTCAGACTCGGATTCCGATGACGATTCAAAGGACAAAAGCGGCAAAAATGGACGACTTGAAAGAGGCAAACTAACTGACGGCATTAAGAAAGAAC
TTGAAACCGCCAGTCTTGAAGTTGCTGAACTGAAGGGTAAGGTTGCTGCTGCAACCGAAGAAAAGGATGCGTTATATCTCGAGTATCAATCAGCATTGAG
CAAGATTGAAGAAGCAGAAGCGATTGCCAAAAGCATGAAGGTTGAATTAGAAACAATAAATGCCGAAAAGGTGAAAATTCTCGTCGAGAACGAAGAACTG
ACGCAGAACCTCAATCAGGCTGGCAAGACAGAAGCAGAACTGAATCAAAGATTAGAAGATATAGCCAAAGAGAAAGAAGGTTTGACCGCAGAGAAAACAG
CTCTCTTGGAAAGGATTGAAGAAGAAGAGAAGATCAAAAGAGAATTGAGGATGGCGAGTGATAGGCTGGCAGATGAGAGAGTGGTTCTTTCCAAAGAACT
TGAAAGTCTCAAGGGAGAAATTTCCACATTAAACGAGAAACTGAAATCATCAGAGGTGGTTATGTCGAATTTGACCCAGAATTCGACAGTTGCAGAGGAA
GAGAAAAAATCTATGGTGACCGAAATTTCTGTACTAAACCAGAAGTTGGAAGAGATGAAATTAGAGAAGGATAGTCTCGTTTCCGAGAAGGAGACTGCAA
CACAGAGAATCCTAGAAGAAGAGAAGGTCACAGCCGATCTAAGAGATTTGGCAGACAAGTTCCAAGGTGAGAATTCATCTCTTAAGCATGAACTAGAAAC
CATTACAAATCAACTCATTGACACGAAAAAGCAATTAGAATCTGCGGAGCAAATGGGTTCAGATTTTTCCGCGAAACTGAAGGTTGCCGAGGAAGAGAAT
GCTTCTCTAGCGTCCAGAGTTTCACAGACTCTAGATGAATTGCAGAGTACTCAAAACACCGTACAGGAACTCAATACTCTATCAAGTCAGTTAGAAGAGA
AATTGGATGGGAAGGAAAAAGAGGTTTCATCTCTTACGCAGAAGTACGAGGCACTTGGTGTCGAATCCGCAATCCAGGTGACAGAATTGGAGTCACAGGT
CGCCACCGTGGAATTAAAATTGGCTGGAATGTCTGAACAGAACAGAGAACTGGAATTGGAGGTCCAGACCAAAATTGATGAAGCGAGCCAACTGGTGGGA
GAGAATGGAAAGCTACAGAGCCAAATTCTGGAACTTGAGAAGATATCAAAGGATAGAGAAGTTGAAATCTCAGAACTGGCAAGGAAACTTGAGGAAAACG
AAAGAGAATCTTCAGCTAGATTTGTAAGCTTGACATCAGAGGTTACAACCCTACTGGCTGATTTGGAATACATGGGTGGTCAGAAAGCTGAGCTCGAAAA
ACAGTTGATAAGCAACAGTGATGAAACATCGAACCAGTTGAAAGTCTACATCGATCAAGTCAATGAACTCCAACAGCAACTGCAGACTGTACACAGCCGA
AAAGTTGAACTGGAAGAGCAACTCCAGAGCAGAATAGAAGCAAATTCAGAAATTCTGATACAGATGGAAAAGCTGAAAGAAGAAATTGCATGCAAGACCG
AGGATCATCAGAAAGTAGTGGCAGATTTGGAATCAATGAGTGGTCAGAAAGCAGAGCTCGAAAAACAGATGACAAGCAGCAGCGAAGAGGTAGCAAACCA
GTTGAAGGTTTTGATGGATCAAGTCAGTGAACTGCAACAGCAACTGCAATCTGTAGAAAGCCAAAAAGTTGAACTGGAAGTCCAGCTCCAGAACAGGATC
GCTGAAACTTCAGAATATCTGATACAGATAGAAAAACTGAAGGAAGAAGTAGTTAGCAAGGCTGAAGATCATCAGAAAGTACTGGCCGAATTTGAATCAC
TGAGTGGTCTGAAAGTGGAACTCGAAAAACAGATGACCAGCAGCAGCGAAGAAGCATCAAACCGGTTGAAGGTCATTATGGATCAAGTCAACGAACTGCA
GCAACAACTGCAGTCTGTAGACAGCCAAAAAGATGAACTAGAAGTCCAGCTCCAGCACAAGATTGCTGAAAATTCAGAAGCTTCGGCACAGATAGAAAAA
CTGAAAGAAGAAATATCAGGCAAGGCTGAGGATCATCAGAAAGTACTGGCAGATTTGGAATCTATGAGTGGTCAGAAAGCGGAACTGGAAAAACAGATGA
CCAGCAACAGCGAAGAAGCATCAAACCAGTTGAAGGTCTTCATGGATCAAGTCAGTGAACTGCAACTTCAACTGCAGTCTGTAGGCAGCCAAAAGGTTGC
ATTGGAAGTGCAACTCCAGAACAAGACAGAAGCAAATTCAGAATCTCTGATACAGATAGAAAAGCTGAAGGAAGAAGTAGCTGCCAAGGCTGAAGATCTT
GAGAAAGTACAGATGGAGAAAGATCATCTAACGGAAAAGATAAAGGAGCTGATTTTGGAAATAGAGACTCTAGGCAACCAGAAGAACAATCTCGAACAGG
AAGTAAGAAATCACATCGAAGAGATCGCATCTCTGAAGGACGAAATGTCGACTTTACACGACAGAGTTTCCGGAATGGACCAGACAATCTCAGAAAGAGG
GCTGGAGTTCTCTGCCCTCCATGAGAAACACACGAGTAGAGAGACTGAAGTTTCTGCTGAGATAACGTCTCTCAAGGAACAAGCAAACAGTCTACAACAA
GAACTCGATTCGCTGCGGAACGAGAAGAACGAACTACAAACACAACTCGAAAAAGACCAGCGTGAATTTTCATCAAAGTTGACCGAAATGGAAAATCAGA
AGACTGAGTTGGCAACTCAGGTTGCTGATCTGCAGAAATTGCTGGACGAACAAGCTGAATCATACAAGAAACTAAGCGACGACTACAAACAAATAGAAGC
TCTGTACGAGGACAGCAAGGCTAATGCGAAATCGGCAGAAAAGAATGCAAACGATCTCAACAAGCAAGTTGCCGAGTTACTGGAGACAATCGAGAGTCAG
AAGCGAGACATCGAATTCAAAGGCGAAGAGCTACGAAGTCTAGGCGACAAGAACAACAACATCGAAGTGAAACTTCGACTCTCGAGCCAGAAAGGCCGTG
TCACCGACCAATTACTGAGCGAACAAGAGGAAACGTTCAGGAAAGAAGAAGCCAAATACAAGGAGAAGCAGAGAGAGCTGGAAGCAAGAATATCGTCGCT
CTCATCTACTCTTGCTGTAACTAAAGAAGCTCATCGCAAAATGGTGACCGAACTCTCCGACAAACTTCACGGCGCCATTACAGGATTGGACACTCTGTCG
TCCAGATTCGACGAGGATTGCAACCGGTATCAGCACCGCGTCATGGAGATGTCTCGCGAGGTTCAGATCGCCAAAGGATGGGTGGTCGCCATGAATAAGA
AGATTGAGAAGTTGAAAAAGGAAGCGAACAGCTCAGCCTTACAATTGAAGCACGCAAGGGAACAAGAATCTGCATTGAAGATAGCAGTGGGGGAGTTGCA
GGCCAAGGCAAGCAGAGAGGAAACAGAGAAGCATAATCTCACCAAAGCCATGGCAGCCATGGAAGCAAGCATGAAGGAGAAAGATGAAGGAATATTCCAC
CTGGGAGAAGAGAAACGAGAAGCCATACGGCAGCTTTGCATCCTGATTGATTATCACCGCGTCCGATACGACGGCATAATCGAAAGCTTACTCTCCAAGG
TTCCAGGCAGACGGCAGAGAGCAGCTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10033945 pacid=23154722 polypeptide=Lus10033945 locus=Lus10033945.g ID=Lus10033945.BGIv1.0 annot-version=v1.0
MTKHRWKESVKSFFGSHIDPEKDEQLQGSKEEFDDKLKRILELINEEDSEVERSKRKPLVDLIEDFHNQYESLYSRYDHLTGELRKKFYGKKRGSDSSSS
SSSDSDSDDDSKDKSGKNGRLERGKLTDGIKKELETASLEVAELKGKVAAATEEKDALYLEYQSALSKIEEAEAIAKSMKVELETINAEKVKILVENEEL
TQNLNQAGKTEAELNQRLEDIAKEKEGLTAEKTALLERIEEEEKIKRELRMASDRLADERVVLSKELESLKGEISTLNEKLKSSEVVMSNLTQNSTVAEE
EKKSMVTEISVLNQKLEEMKLEKDSLVSEKETATQRILEEEKVTADLRDLADKFQGENSSLKHELETITNQLIDTKKQLESAEQMGSDFSAKLKVAEEEN
ASLASRVSQTLDELQSTQNTVQELNTLSSQLEEKLDGKEKEVSSLTQKYEALGVESAIQVTELESQVATVELKLAGMSEQNRELELEVQTKIDEASQLVG
ENGKLQSQILELEKISKDREVEISELARKLEENERESSARFVSLTSEVTTLLADLEYMGGQKAELEKQLISNSDETSNQLKVYIDQVNELQQQLQTVHSR
KVELEEQLQSRIEANSEILIQMEKLKEEIACKTEDHQKVVADLESMSGQKAELEKQMTSSSEEVANQLKVLMDQVSELQQQLQSVESQKVELEVQLQNRI
AETSEYLIQIEKLKEEVVSKAEDHQKVLAEFESLSGLKVELEKQMTSSSEEASNRLKVIMDQVNELQQQLQSVDSQKDELEVQLQHKIAENSEASAQIEK
LKEEISGKAEDHQKVLADLESMSGQKAELEKQMTSNSEEASNQLKVFMDQVSELQLQLQSVGSQKVALEVQLQNKTEANSESLIQIEKLKEEVAAKAEDL
EKVQMEKDHLTEKIKELILEIETLGNQKNNLEQEVRNHIEEIASLKDEMSTLHDRVSGMDQTISERGLEFSALHEKHTSRETEVSAEITSLKEQANSLQQ
ELDSLRNEKNELQTQLEKDQREFSSKLTEMENQKTELATQVADLQKLLDEQAESYKKLSDDYKQIEALYEDSKANAKSAEKNANDLNKQVAELLETIESQ
KRDIEFKGEELRSLGDKNNNIEVKLRLSSQKGRVTDQLLSEQEETFRKEEAKYKEKQRELEARISSLSSTLAVTKEAHRKMVTELSDKLHGAITGLDTLS
SRFDEDCNRYQHRVMEMSREVQIAKGWVVAMNKKIEKLKKEANSSALQLKHAREQESALKIAVGELQAKASREETEKHNLTKAMAAMEASMKEKDEGIFH
LGEEKREAIRQLCILIDYHRVRYDGIIESLLSKVPGRRQRAA
|
DESeq2's median of ratios [FLAX]
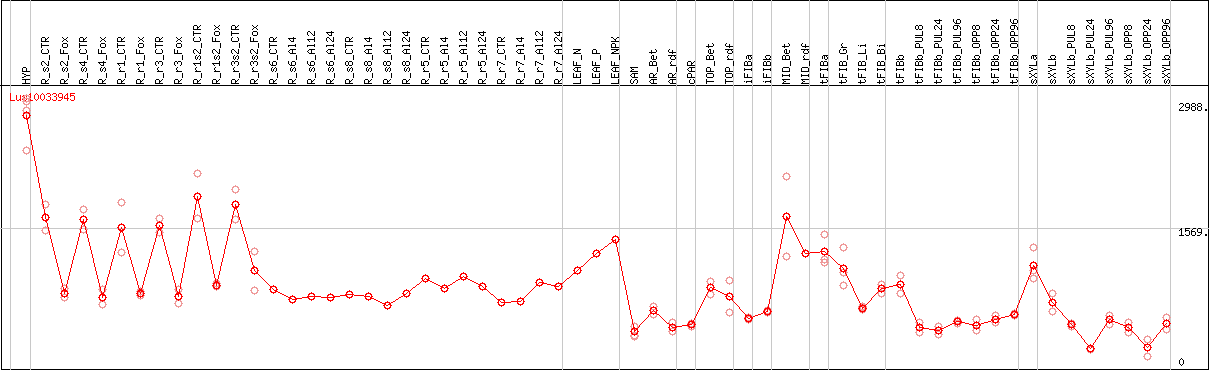
Coexpressed genes
Lus10033945 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
