Lus10034021 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10034021 pacid=23154701 polypeptide=Lus10034021 locus=Lus10034021.g ID=Lus10034021.BGIv1.0 annot-version=v1.0
ATGGCTGCTCCACTGAGTGTCTGGGATGGTGTTGTGGGAATAACCCAGTTGGCACAAAAAAAGGTCGGCGATCCTCTGCTATGGGGTCTCCAAATATCTT
CCAAATTGAGCTCCTATGGCATCTCTTTGCCCTCCCCGGAGCTCGCCACTGTATTGGTTTCTTACATTTGTTGGGATAACAATGTCCCCGTTCTGTGGAA
GTTCCTTGAAAAGGCTTTGGTGCTCAACCTTGTTCCTCCTTTCTTGGTTCTTGCCCTTCTCTCTCAAAGGGTTATTCCATGCAGACGATCCAAGCCAGTT
GTGTATAGACTATACATGGAACTTCTCAGAAGGCATGCTTTTGAGCTCAAAGTTCACATAAATGGCCCAAATCATGAAAGGGTCATGAAATCAGTAGATG
ATGTTCTTCATCTTTCCGAGAATTTCGGCTTAGAAGCCAGTGATCCTGGAATCCTTGTGGTTGAGTTCATCTATTCAATTGTGTGGCAGTTGCTAGATGC
GTCTTTAGATGATGAGGGATTGCTCCAACTTACATCTGATAAAATGTCGGTATGGTCAACAAACTCTCAAGATATGGAAATAGACGGCCATGAAGATTTT
GATGAGAGAAGAATGGAACATAATGGGAAGTTGTTCAGCATGAACACAGTCATTGCTGTTGAGATGATTGGTGGATTCCTGAAAGACAAGAAGACTTCAA
GGATTCTTTACCTGGCACGACACAACTTGCCCTCACATTGGCCAAGGTTTATTGAGAGATTGCAGCTACTTGGAGCAAATTCATCAGCAGTTAGAAGCTC
AAAGGTGTTAACTTCTGAGGATCTTTTGCGTTTGGCTTCAGATAATTGCACAGTTCTCACTCGGGAACCGAAGAAGAACTTGCTGCAGAAGTTCAATGCT
GTAACTAGTTTTGAGTCCCCTTCAGTGTCTTCTGCTGTTCTCTGCGACGGAGGCAGTATTTCTTCTCTTTGGCTTCCTCTTGATCTTGCATTAGAGGATG
CCATGGATGGGTATCAAGTCAATGCTACTAGCGCTATTGAAATTGTAAAAGGTTTGGTGAAAACGCTTCAAGCTATGAATGGGACCACATGGCATGACAC
CTTTGTGGGACTTTGGATTGCTGCTCTCCGCCTTGTTCAAAGGGAAAGGGATCCCATTGAAGGTCCGATCCCTCGCTTGGATACGCGTTTGTGCATGCTC
CTGTCCATCATACCTCTTGTGGTTACTGAACTCATTGAGGAAGAGGACAATGCACCTATGGAAGAAGTAGTAGTAGCAAATCGCCGCAATGATGTGGTAT
CTAGCATACAGTTGTTAGGGGAGTACCAGGGTCTGCTGAGTCCACCGCAGTCTGTTGTATCGTTGGCGAATCAGGCTGCTGCAAAAGCTATGTTGCTTGT
TTCAGGCATCAATGTTGGGAGTGATTACCTTGAGTGTGTCAGCATGAAAGACATGCCAGTCAGTTGTTCTGGGAACATGCGCCATCTGATTGTTGAAGCT
TGTATAGCCAGAAATTTGTTAGATACATCTGCATACTTTTGGCCCGGGTATGTGAATGGTCATATTAACCAAATACCTAATGTCCCACCTCAGATGCCTG
GTTGGTCATCATTCTTGAAGGGTGCTCCTCTTACTCCAATTATGATCAATGCAATGGTTTCAACCCCGGCTTCAAGTTTAGCAGAGCTGGAGAAAGTATT
TGAGCTAGCAGTCAAAGGATCAGAGGATGAGAAGATATCAGCTGCTACAATTCTCTGTGGGGCTTCTCTTCTCCGGGGTTGGAACATTCAGGAACACACA
GCTCAGTTCATTACTAGGTTATTGTCTCCACCAAAAGCTGCAGATTATTCAGGGGATGAAAGCCATTTGATAGGCGCTGCTCCGATCCTGAATCTGCTGA
TTGTTGGCATATCATCAGTTGATTGCGTTCAGGTTTTCTCCCTCCATGGCTTGATCCCACATCTTGCGAGCTCGCTGATGCCTATCTGCGAGGTTTTCGG
ATCTTGTGTGCCTGATGTGTCATGGAAGCTTCCCAATGGTGATGAAATCTCTGCCCATGCTGTATTTTCAAATGCATTTGCTGTTCTTTTGAAGCTATGG
CGGTTTAACCACCCCCCTCTCGAGTATGGTGTTGGAGATGTCCCAACTGTCGGATCTCAGTTAACTCCCGAATACCTTCTGCTAGTAAGAAACTCCTACC
TACCCTCATCTGGAATAGTTCTCAAGGATCAAATCAAGAGAAGACTTTCAGAAGTTGCAACTTCATCATCTCCACACCCTGTCTTTCTGGATTCATTCCC
AAGACTCAAATGCTGGTATCGGCAACATCAGAAATGCATAGCGTCGACCCTCTCTGGTCTTGTTCATGGAACTCCAGTTCATCAGACTGTGAACTTGCTA
CTTACTATGATGTTCAGAAAGATGAATAGAGGAAGTCAATCAGTAACTACTATTACCTCTGGAAGCAGTAGTTCATCCGGACCAGGAAATGAGGATTCTT
CTCTAAGACCTAAATTACCTGCCTGGGAGATTTTAGAAGCTGTTCCTTTTGTGGTTGATGCTGCTTTAACAGCTTGTGCTCATGGAAGGCTTTGTCCTCG
TGAGCTCGCCACAGGGCTGAAAGATTTAGCAGATTTTCTCCCTGCATCTTTGGCAACCATTGTTAGTTACTTCTCAGCTGAAGTAAGTAGAGGAGTCTGG
AAACCAGCATTTATGAATGGCACAGACTGGCCAAGTCCTGCAGTGAATTTGTCGAACGTTGAGGACAAGATCAAGAAAATCTTAGCAGCTACTGGTGTCG
ATATACCTAGTTTGGCAGCAGGGGGGAGCTCACCTGCCACACTTCCACTGCCTCTGGCTGCTTTTGTCAGTCTCACAATCACCTATAAAATCGACAAAAC
CTCAGAGAGATTCCTCAATTTGGCTGGTCCAGCGCTGGAATCTCTTGCAGCAGGTTGCCCTTGGCCTTGCATGCCGATTGTGGCTTCCTTATGGACCCTG
AAAGCAAAACGATGGTTCGATTTCCTCGTTTTTTCTGCATCACGCACGGTCTTTCTCCACAACAATGATGCAGTATTTCAGCTCCTCAAAAGCTGCTTTA
CTGCTACACTCGGTGGTGCAGCCATCTCAAGCAATGGTGGGGTGGGAGCACTTCTCGGTCATGGCTTTGGTTCACATTTCTGCGGTGGGATATCTCCTGT
AGCTCCGGGAATCCTATACTTGAGAGTCTACCAGTCCATCAGAGACATTGAGATCATCACAGAAGAGATAATTTCTCTCATGATGCACTCCGTGACACAA
ATTGCGTGCACGGGGCTTCCTAAAGAGAGATTAGGGAGGCTGAAGAGGACAAAGAATGGACTTGAGTGTGGCCAAGTTTCGCTTTCTGCAGCAATGACGA
GGGTGAAGCTTGCAGCTTCACTTGGTGCTTCTCTAGTTTGCTTATCTGGAGGATTAGGCTTGGTTCAGTTGCTGTTCCAAGAAACATTGCCCTCTTGGTT
TATATCTAGCCATCACTCCGAGCAGGAAGAAGATGGCTGCCTCGGAGGGATGGTTGCCATGCTAAGAGGGTATGCCTTGGCATATTTTGCAGTTCTTTGT
GGAGCTTTTGCTTGGGGAATTGACAATTCGTCATCCGCTTCCAGGAGGCGTCCGAAAGTCGTGAGAGCTCACATGGAGTTCCTGGCTAGTGCTCTAGATG
GCAAAATCTCACTTGGTTGCGACAGTGCAACTTGGAAATCCTACGTGTCGGGATTCATTAGCCTAATGGTGGGTTGTACACCTCCATGGGTTCTCGAGCT
CGAAGCAGATGTTTTGAAGAGACTAAGTAAAGGGTTGAGACAGTGGAACGAGGAGGAATTGGCATTGGCTTTGCTAGGCATTGGTGGGGTGGAAACCATG
GGTGCAGCTGCTGAATTGATAATTGAATTGCATTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10034021 pacid=23154701 polypeptide=Lus10034021 locus=Lus10034021.g ID=Lus10034021.BGIv1.0 annot-version=v1.0
MAAPLSVWDGVVGITQLAQKKVGDPLLWGLQISSKLSSYGISLPSPELATVLVSYICWDNNVPVLWKFLEKALVLNLVPPFLVLALLSQRVIPCRRSKPV
VYRLYMELLRRHAFELKVHINGPNHERVMKSVDDVLHLSENFGLEASDPGILVVEFIYSIVWQLLDASLDDEGLLQLTSDKMSVWSTNSQDMEIDGHEDF
DERRMEHNGKLFSMNTVIAVEMIGGFLKDKKTSRILYLARHNLPSHWPRFIERLQLLGANSSAVRSSKVLTSEDLLRLASDNCTVLTREPKKNLLQKFNA
VTSFESPSVSSAVLCDGGSISSLWLPLDLALEDAMDGYQVNATSAIEIVKGLVKTLQAMNGTTWHDTFVGLWIAALRLVQRERDPIEGPIPRLDTRLCML
LSIIPLVVTELIEEEDNAPMEEVVVANRRNDVVSSIQLLGEYQGLLSPPQSVVSLANQAAAKAMLLVSGINVGSDYLECVSMKDMPVSCSGNMRHLIVEA
CIARNLLDTSAYFWPGYVNGHINQIPNVPPQMPGWSSFLKGAPLTPIMINAMVSTPASSLAELEKVFELAVKGSEDEKISAATILCGASLLRGWNIQEHT
AQFITRLLSPPKAADYSGDESHLIGAAPILNLLIVGISSVDCVQVFSLHGLIPHLASSLMPICEVFGSCVPDVSWKLPNGDEISAHAVFSNAFAVLLKLW
RFNHPPLEYGVGDVPTVGSQLTPEYLLLVRNSYLPSSGIVLKDQIKRRLSEVATSSSPHPVFLDSFPRLKCWYRQHQKCIASTLSGLVHGTPVHQTVNLL
LTMMFRKMNRGSQSVTTITSGSSSSSGPGNEDSSLRPKLPAWEILEAVPFVVDAALTACAHGRLCPRELATGLKDLADFLPASLATIVSYFSAEVSRGVW
KPAFMNGTDWPSPAVNLSNVEDKIKKILAATGVDIPSLAAGGSSPATLPLPLAAFVSLTITYKIDKTSERFLNLAGPALESLAAGCPWPCMPIVASLWTL
KAKRWFDFLVFSASRTVFLHNNDAVFQLLKSCFTATLGGAAISSNGGVGALLGHGFGSHFCGGISPVAPGILYLRVYQSIRDIEIITEEIISLMMHSVTQ
IACTGLPKERLGRLKRTKNGLECGQVSLSAAMTRVKLAASLGASLVCLSGGLGLVQLLFQETLPSWFISSHHSEQEEDGCLGGMVAMLRGYALAYFAVLC
GAFAWGIDNSSSASRRRPKVVRAHMEFLASALDGKISLGCDSATWKSYVSGFISLMVGCTPPWVLELEADVLKRLSKGLRQWNEEELALALLGIGGVETM
GAAAELIIELH
|
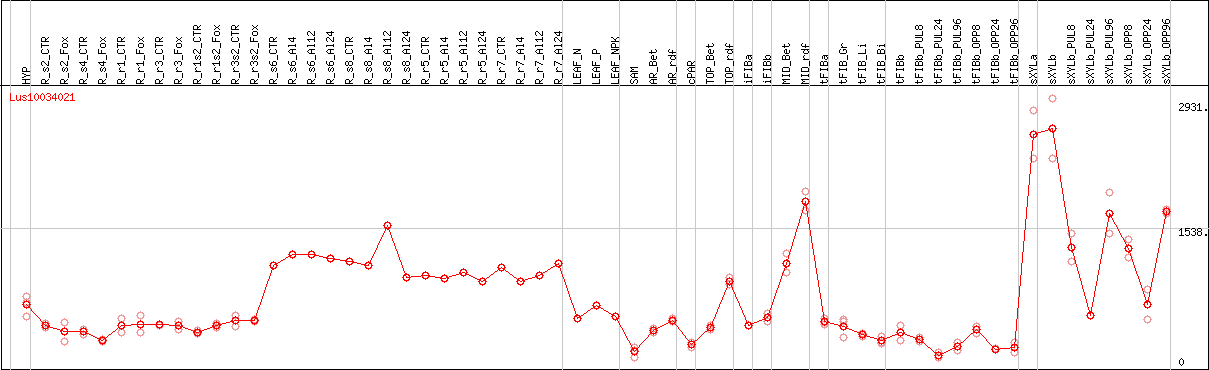
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034021 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
