External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G06120 210 / 7e-69
bHLH
MUTE, bHLH045
MUTE, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G01460 116 / 7e-31
bHLH
bHLH057
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G72210 112 / 1e-29
bHLH
bHLH096
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G46810 112 / 7e-29
bHLH
bHLH070
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G53210 100 / 1e-24
bHLH
SPCH, bHLH098
SPEECHLESS, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT3G61950 98 / 3e-24
bHLH
bHLH067
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT3G24140 96 / 9e-23
bHLH
bHLH097, FMA
FAMA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G22490 92 / 6e-22
bHLH
bHLH094
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT5G46690 90 / 4e-21
bHLH
bHLH071
beta HLH protein 71 (.1)
AT5G65320 70 / 5e-14
bHLH
bHLH099
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010503
265 / 1e-89
AT3G06120 191 / 4e-61
MUTE, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10007139
137 / 9e-38
AT3G24140 308 / 2e-101
FAMA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10016680
135 / 4e-37
AT3G24140 358 / 2e-120
FAMA, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10010101
116 / 3e-32
AT2G46810 199 / 1e-63
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10012603
112 / 8e-31
AT2G46810 202 / 4e-65
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10000332
103 / 8e-26
AT1G22490 221 / 4e-70
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10014938
97 / 6e-24
AT5G53210 264 / 2e-87
SPEECHLESS, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10039328
96 / 4e-23
AT5G53210 257 / 3e-83
SPEECHLESS, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10001066
95 / 2e-22
AT5G53210 268 / 9e-88
SPEECHLESS, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Lus10034031 pacid=23154668 polypeptide=Lus10034031 locus=Lus10034031.g ID=Lus10034031.BGIv1.0 annot-version=v1.0
ATGTCCCACATAGCGGTAGAGAGGAATCGGAGGAGGCAGATGAATGAGAATCTCACAGTCTTGCGATCTCTAACCCCTCGCATCTACATCAAAAGGGGGG
ACCAAGCATCGATCATAGGGGGAGTGATAGAGTTCATAAAAGAGCTGCACCACCTACTTCAAGCACTGGAGTCCAAGAAGCAAAGGCGATTGAGTCTGAG
CCCTAGCAGTGCTAGTAGCCCTAGCCCACGTCGTCATATGTTGCAGCTCATGCCACCTACTGCAGCTACTTGCGATCATCATATTAATAACAACATGAAT
GTAAAGAATGAGGAGTCATCGGAGTCGTTGTCGTCGTCGTCGCAGCAGCAGCAGCAGCAGCTGGGGGCATGTTGCAACTCCCCGGTTGCTGACGTGGAAG
CAAAGCTAACAGGATCCAATGTGGAGTTGAAGGTGATATCGAGCCGGGTAAAGGGTCAGGTTGTGAAGATGATAAGTGTGCTGGAGATGCTGTCGTTCGA
GGTACTCCATCTCAACATCAGTAGCATGGATGATACTGTCCTCTACTCCTTTCTCCTTAAGATAGGGTTGGAATGTAGGGTAAGCATGGAGGAACTTGCA
CTTGAAGTCCAACGTACTTTCTTCTCGACATCTACAGCTCCTGCTCCTGCTCTTGCTAGTAATGCTACTGATATTCATCGTCATCACCATACGAATCATG
AGCTGCTTTGTTTGTAG
AA sequence
>Lus10034031 pacid=23154668 polypeptide=Lus10034031 locus=Lus10034031.g ID=Lus10034031.BGIv1.0 annot-version=v1.0
MSHIAVERNRRRQMNENLTVLRSLTPRIYIKRGDQASIIGGVIEFIKELHHLLQALESKKQRRLSLSPSSASSPSPRRHMLQLMPPTAATCDHHINNNMN
VKNEESSESLSSSSQQQQQQLGACCNSPVADVEAKLTGSNVELKVISSRVKGQVVKMISVLEMLSFEVLHLNISSMDDTVLYSFLLKIGLECRVSMEELA
LEVQRTFFSTSTAPAPALASNATDIHRHHHTNHELLCL
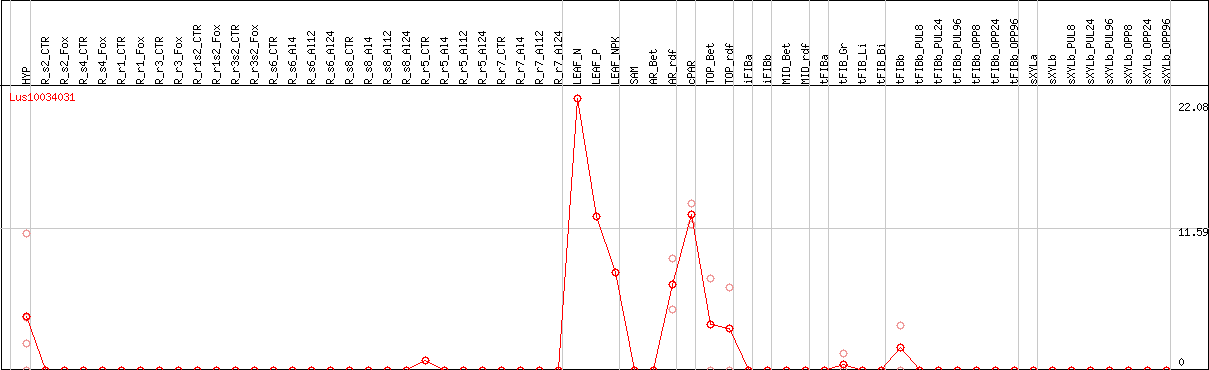
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034031 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.