External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G23730 368 / 8e-129
XTH16
xyloglucan endotransglucosylase/hydrolase 16 (.1)
AT4G14130 360 / 5e-126
XTR7, XTH15
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
AT5G57560 359 / 2e-125
XTH22, TCH4
xyloglucan endotransglucosylase/hydrolase 22, Touch 4, Xyloglucan endotransglucosylase/hydrolase family protein (.1)
AT4G25810 351 / 3e-122
XTH23, XTR6
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
AT5G57550 348 / 3e-121
XTR3, XTH25, EXGT-A5
xyloglucan endotransglycosylase 3, xyloglucan endotransglucosylase/hydrolase 25 (.1)
AT4G30270 347 / 4e-121
XTH24, SEN4, BRU1, MERI-5, MERI5B
SENESCENCE 4, meristem-5, MERISTEM 5, xyloglucan endotransglucosylase/hydrolase 24 (.1)
AT5G57540 330 / 4e-114
AtXTH13, XTH13
xyloglucan endotransglucosylase/hydrolase 13 (.1)
AT5G57530 329 / 1e-113
AtXTH12, XTH12
xyloglucan endotransglucosylase/hydrolase 12 (.1)
AT4G25820 327 / 1e-112
ATXTH14, XTR9, XTH14
xyloglucan endotransglycosylase 9, xyloglucan endotransglucosylase/hydrolase 14 (.1)
AT5G48070 320 / 6e-110
XTH20, ATXTH20
xyloglucan endotransglucosylase/hydrolase 20 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10003048
552 / 0
AT3G23730 380 / 7e-134
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10010936
531 / 0
AT3G23730 382 / 2e-134
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10010939
517 / 0
AT3G23730 384 / 2e-135
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10010938
513 / 0
AT4G14130 386 / 3e-136
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Lus10031392
511 / 0
AT3G23730 384 / 3e-135
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10031393
508 / 0
AT3G23730 388 / 8e-137
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10007349
431 / 1e-153
AT3G23730 394 / 4e-139
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10020772
430 / 1e-153
AT3G23730 393 / 7e-139
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Lus10020773
430 / 2e-153
AT3G23730 394 / 4e-139
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G201250
427 / 3e-152
AT4G14130 395 / 2e-139
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Potri.005G201200
424 / 4e-151
AT4G14130 405 / 2e-143
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Potri.002G060500
417 / 2e-148
AT4G14130 424 / 3e-151
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Potri.002G060400
395 / 7e-140
AT4G14130 402 / 2e-142
xyloglucan endotransglycosylase 7, xyloglucan endotransglucosylase/hydrolase 15 (.1)
Potri.002G236200
384 / 2e-135
AT3G23730 436 / 1e-155
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Potri.014G146100
383 / 7e-135
AT3G23730 451 / 1e-161
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Potri.011G077320
376 / 4e-132
AT3G23730 386 / 7e-136
xyloglucan endotransglucosylase/hydrolase 16 (.1)
Potri.013G005700
360 / 3e-125
AT4G25810 431 / 5e-153
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.018G095100
355 / 6e-124
AT4G25810 420 / 1e-149
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
Potri.018G095200
352 / 1e-122
AT4G25810 414 / 3e-147
xyloglucan endotransglucosylase/hydrolase 23, xyloglucan endotransglycosylase 6 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0004
Concanavalin
PF00722
Glyco_hydro_16
Glycosyl hydrolases family 16
CL0004
Concanavalin
PF06955
XET_C
Xyloglucan endo-transglycosylase (XET) C-terminus
Representative CDS sequence
>Lus10034098 pacid=23154693 polypeptide=Lus10034098 locus=Lus10034098.g ID=Lus10034098.BGIv1.0 annot-version=v1.0
ATGGCGACAACTCTGCCTCTGGTGATGCTACTAGTGATGGCGTTGAGTACTCATCTAATGCCAATCTCGAATGCCAACTTCCACCAGCATTTTGATGTCA
CTTGGGGGCATCACCATGCTCAGATCACCAACGGCGGGCAGCAACTCGCCCTCTCCCTCGACAAGGATTCGGGCGCTGGCTTCAAATCCAAGAACGAGTA
TCTCTTTGGCCGTATCGATATGCAACTCAAGCTCATCGCTGGCAATTCTGCTGGGAGTGTCACCACCTTTTACTTGTCGTCGGAAGAAGGACCGAATCAC
GACGAAATCGACTTCGAGTTTCTTGGCAATTCCAGTGGTAATCCTTACACTCTGCACACCAATGTGTTTAGTCAAGGAAAAGGCAACCGAGAGCAACAAT
TTCATCTCTGGTTCGACCCAACTGCGGCTTTCCACACCTACACCATCCTTTGGAATTCCCAAAGAATCATATTCTTAGTGGACAACATACCAATCAGGGT
GTTCAACAACTTGGAATCAATGGGGGTTCCATTCCCAAGCAAGAAGCCGATGAGCATTCACTCGAGCTTATGGAACGCCGATGACTGGGCCACCCAAGGC
GGAAGAGTCAAGGCCGATTGGACCAACGCTCCATTCGTCGCTTCTTACAGAAACTTCAAGGCAGAAGCTTGCGTCTGGTTGCCGTCCTCGGCCTCGTCAG
CCTGCTCTCCGGGAGGATGGCAGACTCAGGGGATTGATGCCGCCGGCCGGAACAGGATCCGGTGGGTGCAGAGCAAATATATGATCTACAATTACTGCAC
TGACTTCCAACGGTTTCCTCAGGGTGTCCCTGTTGAATGCAAGCAGTCCCGATTCCTTTGA
AA sequence
>Lus10034098 pacid=23154693 polypeptide=Lus10034098 locus=Lus10034098.g ID=Lus10034098.BGIv1.0 annot-version=v1.0
MATTLPLVMLLVMALSTHLMPISNANFHQHFDVTWGHHHAQITNGGQQLALSLDKDSGAGFKSKNEYLFGRIDMQLKLIAGNSAGSVTTFYLSSEEGPNH
DEIDFEFLGNSSGNPYTLHTNVFSQGKGNREQQFHLWFDPTAAFHTYTILWNSQRIIFLVDNIPIRVFNNLESMGVPFPSKKPMSIHSSLWNADDWATQG
GRVKADWTNAPFVASYRNFKAEACVWLPSSASSACSPGGWQTQGIDAAGRNRIRWVQSKYMIYNYCTDFQRFPQGVPVECKQSRFL
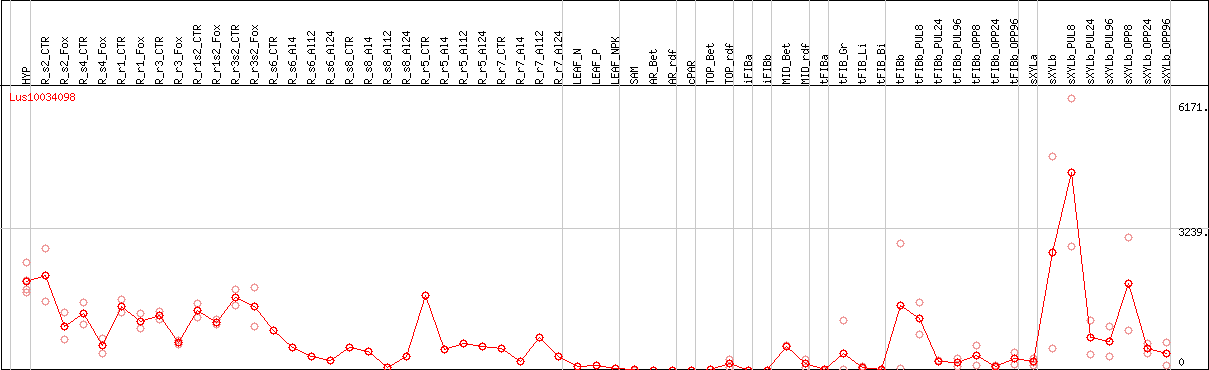
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034098 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.