External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G68840 226 / 1e-73
AP2_ERF
EDF2, RAV2, RAP2.8, TEM2, AtRAV2
TEMPRANILLO 2, RELATED TO AP2 8, ETHYLENE RESPONSE DNA BINDING FACTOR 2, related to ABI3/VP1 2 (.1.2)
AT1G25560 219 / 2e-70
AP2_ERF
EDF1, TEM1
TEMPRANILLO 1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, AP2/B3 transcription factor family protein (.1)
AT1G13260 215 / 4e-69
AP2_ERF
EDF4, RAV1
ETHYLENE RESPONSE DNA BINDING FACTOR 4, related to ABI3/VP1 1 (.1)
AT3G25730 174 / 2e-53
AP2_ERF
EDF3
ethylene response DNA binding factor 3 (.1)
AT3G61970 156 / 8e-47
B3
NGA2
NGATHA2, AP2/B3-like transcriptional factor family protein (.1)
AT1G01030 156 / 3e-46
B3
NGA3
NGATHA3, AP2/B3-like transcriptional factor family protein (.1)
AT2G46870 154 / 1e-45
B3
NGA1
NGATHA1, AP2/B3-like transcriptional factor family protein (.1)
AT4G01500 150 / 6e-44
B3
NGA4
NGATHA4, AP2/B3-like transcriptional factor family protein (.1)
AT5G06250 148 / 8e-44
B3
AP2/B3-like transcriptional factor family protein (.1.2)
AT3G11580 146 / 1e-43
B3
AP2/B3-like transcriptional factor family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041487
371 / 4e-130
AT1G13260 416 / 8e-146
ETHYLENE RESPONSE DNA BINDING FACTOR 4, related to ABI3/VP1 1 (.1)
Lus10021707
211 / 1e-67
AT1G13260 369 / 1e-127
ETHYLENE RESPONSE DNA BINDING FACTOR 4, related to ABI3/VP1 1 (.1)
Lus10021326
143 / 1e-41
AT5G06250 202 / 2e-63
AP2/B3-like transcriptional factor family protein (.1.2)
Lus10004226
131 / 2e-36
AT5G06250 183 / 4e-55
AP2/B3-like transcriptional factor family protein (.1.2)
Lus10026212
111 / 2e-30
AT1G51120 150 / 4e-44
AP2/B3 transcription factor family protein (.1)
Lus10036282
102 / 5e-26
AT1G50680 268 / 2e-88
AP2/B3 transcription factor family protein (.1)
Lus10025226
69 / 2e-13
AT4G32010 808 / 0.0
VP1/ABI3-LIKE 2, HSI2-like 1 (.1)
Lus10022741
68 / 5e-13
AT4G32010 748 / 0.0
VP1/ABI3-LIKE 2, HSI2-like 1 (.1)
Lus10029431
59 / 1e-11
AT2G36080 69 / 3e-16
AP2/B3-like transcriptional factor family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G117100
248 / 8e-82
AT1G25560 422 / 6e-148
TEMPRANILLO 1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, AP2/B3 transcription factor family protein (.1)
Potri.010G129200
244 / 8e-80
AT1G25560 441 / 1e-154
TEMPRANILLO 1, ETHYLENE RESPONSE DNA BINDING FACTOR 1, AP2/B3 transcription factor family protein (.1)
Potri.014G107200
160 / 3e-47
AT2G46870 241 / 6e-77
NGATHA1, AP2/B3-like transcriptional factor family protein (.1)
Potri.011G149700
159 / 1e-46
AT2G46870 218 / 2e-67
NGATHA1, AP2/B3-like transcriptional factor family protein (.1)
Potri.002G181600
158 / 2e-46
AT2G46870 264 / 6e-86
NGATHA1, AP2/B3-like transcriptional factor family protein (.1)
Potri.001G452200
158 / 4e-46
AT2G46870 198 / 4e-60
NGATHA1, AP2/B3-like transcriptional factor family protein (.1)
Potri.006G208100
149 / 2e-43
AT2G36080 228 / 3e-73
AP2/B3-like transcriptional factor family protein (.1.2)
Potri.016G074500
147 / 9e-43
AT2G36080 219 / 9e-70
AP2/B3-like transcriptional factor family protein (.1.2)
Potri.018G109201
118 / 7e-32
AT1G51120 286 / 8e-95
AP2/B3 transcription factor family protein (.1)
Potri.006G186301
115 / 1e-30
AT1G51120 287 / 3e-95
AP2/B3 transcription factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0405
DNA_b-psBarrel
PF02362
B3
B3 DNA binding domain
Representative CDS sequence
>Lus10034276 pacid=23175834 polypeptide=Lus10034276 locus=Lus10034276.g ID=Lus10034276.BGIv1.0 annot-version=v1.0
ATGCTGAGGAAACACACTTACAACGACGAGCTCCAGCAGAGCCGCCGTGGCCGCCACCACAACCACCACCGTGGGAAACAGAGCGGCACACTCGATTCCT
CCGACAATATTGCTGCCCGGGGAGCCAGGGAGCAGCTGTTCGAGAAGGCGGTGACTCCGAGCGACGTCGGGAAATTGAACCGGCTAGTGATCCCCAAACA
GCACGCGGAGAAGCATTTCCCACTCCAGTTTGGCGGCGCCGCCGCCGGAGGAGGAAGCGGCAGCACGAAAGGCGTGCTGCTGAATTTCGAGGACCCCGGA
GGGAAAGTGTGGCGGTTCCGGTACTCATACTGGAACAGCAGCCAGAGCTACGTGCTGACGAAAGGATGGAGTCGGTTTGTGAAGGAGAAGAATCTGAAAG
CCGGAGATACTGTGAGCTTCCATAGATCCGTTACAGGGCAGGATCAGAAGCAGCTATACATTGACTGGAAGGTCAGGAGTGATGAAGGAGTAGAGTCGAG
CCGGTTGGTTTGCAGGGTCCAACCGGTTCAGATGGTTCGGTTATTTGGGGTGAATATATTTAAACCGGTGGACGGTGGGTGTAATGGGAAAAGAACGATG
GAGGAGATGGTGGGTTTTCACTGCAGTAAAAAGCAAAGGATCATTGGAGCCTTGTAG
AA sequence
>Lus10034276 pacid=23175834 polypeptide=Lus10034276 locus=Lus10034276.g ID=Lus10034276.BGIv1.0 annot-version=v1.0
MLRKHTYNDELQQSRRGRHHNHHRGKQSGTLDSSDNIAARGAREQLFEKAVTPSDVGKLNRLVIPKQHAEKHFPLQFGGAAAGGGSGSTKGVLLNFEDPG
GKVWRFRYSYWNSSQSYVLTKGWSRFVKEKNLKAGDTVSFHRSVTGQDQKQLYIDWKVRSDEGVESSRLVCRVQPVQMVRLFGVNIFKPVDGGCNGKRTM
EEMVGFHCSKKQRIIGAL
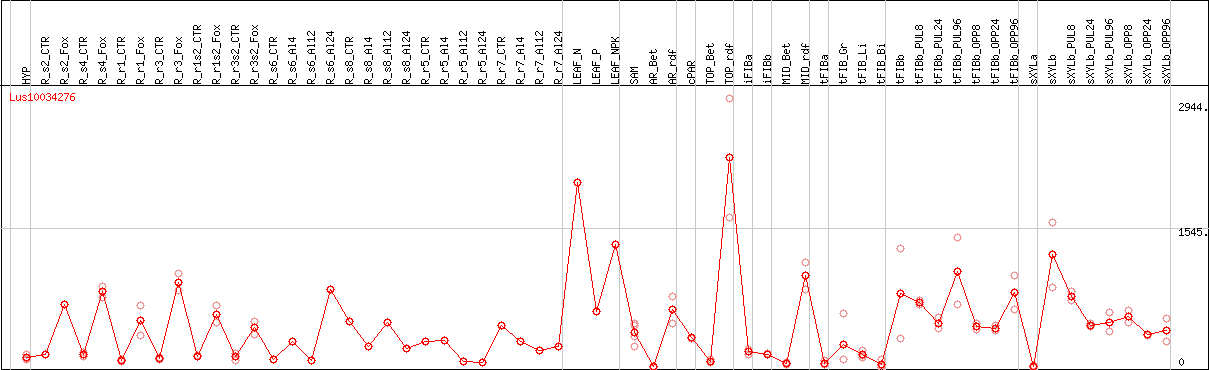
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034276 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.