Lus10034306 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10034306 pacid=23176053 polypeptide=Lus10034306 locus=Lus10034306.g ID=Lus10034306.BGIv1.0 annot-version=v1.0
ATGGGATCCCTGAAGAACAAAGATGAAGTCGAACTGGCTGGAGATGAATCAGCTCCCAAAGATGGGATTCTTTACGTTAATGGAGTTCGTAAGGTTTTGC
CCGATGGCTTAGCTCACTTGACCCTTCTTGAGTATCTCAGAGATATTGGGCAGACTGGGACAAAGCTGGGATGTGGTGAAGGTGGTTGTGGTGCTTGCAC
TGTTATGGTTTCCCAATATGATATCAGGACAAAGACATGTCTCCACCGTGCTGTCAATGCCTGCTTGGCACCTCTGTATTCCGTTGAGGGAATGCATGTA
ATAACCGTCGAGGGAGTTGGTACTCGAAGAAGTGGGTTGCACCCGGTTCAGGAATCATTGGCAAGTGCTCATGGTTCACAATGTGGATTTTGCACGCCTG
GATTTGTAATGTCGATGTATGCCCTATTAAGGTCGAGTCGAGAGCCTCCTACCGAGGAGCAAATCGAAGAATCCCTTGCTGGAAATTTGTGCCGTTGCAC
TGGTTACAGACCTATTGTTGATGCGTTCCGAGTCTTTGCAAAGACGGATGATGCATTGTACACTGAAGCGAATAGTAAAGAAGGTGAATTTGTGTGCCCT
TCAACCGGTAAACCGTGTTCATGCAAAAAAAATAGTTCACCAGTCGATAATACAGCTTGTGGTGAGAGGTATAGTATTGCATCTTATAGTGAAGTAGATG
GGAGCAAGTACATTGAGAAAGAGCTAATCTTTCCTCCTGAGCTCTTAATGAGGAAACTAACTCCCTTGAATCTGAATGGCTTCAATGGGCTGAAATGGTT
TCGGCCTGTAAGACTTCAGCATGTCCTGGACTTAAAAGCTGAGCACCCAGATGCAAAGTTGGTGATCGGTAATAGTGAGGTCGGGATAGAGATGAGGCTA
AAGAGACTCCAGTATCAGACTTTGATCCATGTGGGTCATGTACCTGAACTTAACGTCTTGAACGTTAAGGAAGATGGTATAGAGATAGGTGCAGCTGTAA
GATTGACTGATCTCATGCAACTTCTTAGAAAGGTTGCAAATGAGCGTGTTGGTCACACAACATCATCTTGTAAGGCCATTGTTGAACAGTTGAAGTGGTT
TGCAGGAACTCAAATAAAGAATGTTGCATCTGTTGGAGGCAACATTTGCACAGCCAGTCCAATATCTGATTTAAACCCACTCTGGATGGCTGCAAGGGCT
AAGTTTCGAATTATTGACTTAAAGGGAAGCATTAGAACTACACAGGCAGATAAGTTCTTCCTGGGTTACAGGAAGGTGGATCTTCAGAATGGCGAGATCT
TACTCTCGGTATTCTTACCATGGACGATGAAGTTTGAGTATGTGAAGGAATTTAAGCAGGCTCATCGTAGGGACGATGACATTGCAATAGTGAATGCTGG
AATGCGTGTTTCTCTTGAAAAGAAGGGTGAAGAATGGGTTGTCTTAGATGCATCAATTGCTTATGGTGGCGTGGCTCCCCTTTCTTTGTCTGCTGTCAAA
GCCAGTGGCTTCCTTGTTGGGAAGAGCTGGCATCAGGAGCTGTTCTTGGGTGCTTTACAGGCCTTAGAGACCGATATTGTGATCAAGGAAGATGCTCCTG
GTGGTATGGTAGAATTCAGGAAATCGTTGACTCGCAGTTTCTTGTTCAAGTTTTTCCTTTGGGTTTCACATCAAATGGAAGGAGATCAATGTGGTGGGAT
AAGTGTACCATTGTCACATCTTTCAGCTGTACAATCCTTCTCTCGACCTTCAGTCATGGGAAGCCAAGATTATGAGATTAGGAAACACGGGACCTCTGTG
GGATCGCCAGAGATTCATCTATCTGCAAGACTTCAGGTCACAGGGGAAGCAGAGTATGCAGATGATGCTCCGATGCCACCGAATGGTTTGCATGCTGCTC
TTGTATTGAGCCGAAAACCTCATGCTCGGATTGTTGCAATCGATGATTCGGAAGCCAAGTCTTCACCTGGTTTCGCAGGCGCGTTCTTTGCCAAGGATGT
ACCGGGTGACAATCACATTGGAGCAATTGTACACGACGAGGAAGTATATGCTACGGAGTTCGTTACCTGTGTTGGTCAGGTTATCGGGGTGGTTGTTGCT
GATACGCATGAGAATGCAAAGTTAGCTGCGAGAAAAGTTCATGTTGAGTACAAAGAGCTCCCTGCAATATTGACCATACAAGAAGCTGTAAATGCTGGAA
GTTTCCATCCTCACACGGAGAAATGGTTGAAAAGTGGAGATGTGGATGCTTGTTTTGGATCGGGCAAATGTGACAAGATCGTCGAAGGAGAGGTATGCAT
AGGGGGACAGGAACACTTCTACTTGGAACCACAAAGCAGCTACGTTTGGACGTTGGATAGTGGCAATGAAGTTCACATGATTTCGTCGACACAAGCTCCT
CAAAAGCACCAGAAGTATGTGTCCCACGTGCTTGGTATTCCGATGTCGAAAGTAGTATGCAAAACGAAACGAATAGGTGGTGGATTCGGTGGGAAGGAGA
CTCGGTCAGCTTTCCTTGCTGCAGCTGCTTGTGTACCATCTTATCTATTGAATCGTCCTGTGAAGATCACACTAGATCGAGATGTGGACATGATGATAAC
AGGGCAACGTCACAGCTTTCTTGGGAAATACAAGGTTGGCTTCACAAATGAGGGGAAAATTCTTGCACTGGATCTCAAGATCTATAACAATGCTGGAAAC
TCTCTAGACCTTTCGCTTGCTATACTCGAACGAGCGATGTTCCATTCTGACAATGTTTACGAAATTCCAAACGTTCGAATAGTAGGAAATGTTTGCTTGA
CTAATCTTCCGAGTAATACTGCCTTCCGCGGATTTGGAGGTCCCCAAGGTATGCTTATAACTGAGAATTGGATCCAGAGAGTTGCTGTAGAAGTTAACAA
ATCCCCAGAAGAGATACGGGAGATGAATTTTCAAGGTGAAGGAACAATACTGCATTACGGTCAGCAACTACAGCACTGTACACTATCCCCACTGTGGAGT
GAACTGAAGTTGTCTTCCGACTTCTTGAAGGCCCGTGAAGATGTTCATCACTTCAACCGTCATAACCGGTGGAAGAAGCGTGGATTGGCTATGGTTCCAA
CGAAATTCGGGATATCTTTTACTTCGAAGTTTATGAACCAAGCTGGCGCGCTTGTGAATGTCTACACGGATGGAACTGTTTTGGTTACACATGGCGGTGT
AGAAATGGGTCAAGGTTTGCATACCAAGGTCGCTCAAGTTGCTGCTTCGGCCTTCGATATTCCTCTGAGTTCTGTGTTCATTTCTGAGACCAGTACTGAC
AAGGTACCGAACTCATCCCCTACTGCAGCTTCGGCAAGCTCGGATATGTACGGAGCAGCAGTTCTGGATGCTTGTGAGCAGATCAAAGCCCGAATGGAAC
CTGTTGCTTGTAAACATAGCTTCACCTCTTTCGCTGAGTTGGCGGGTGCTTGCTATGCTCAAAGAATAGACCTTTCTGCTCATGGATTCTACATTACACC
CGATATTGGGTTTGACTGGACGACTGGTCAAGGGAACCCCTTTCGTTACTTCACGTATGGTGCTGCATTCGCGGAGGTCGAGATCGACACTCTTACCGGA
GATTTCCATACGAGGAATGTGAATGTTGTTATGGACCTTGGCTATTCTCTCAATCCAGCCATTGATGTCGGACAGATCGAAGGAGCATTCGTGCAAGGAC
TGGGATGGGTAGCGTTAGAAGAACTGAAATGGGGAGATGAAGCACATAAGTGGATCGCACCGGGTTGCTTGTACACTTGTGGCCCTGGAAGTTACAAGAT
CCCTTCTTTGAATGATGTCCCATTCAAGTTCAATGTGTCACTTCTCAAGGGCCATCCGAATGTGAAGGCAATCCATTCATCGAAAGCAGTAGGCGAGCCG
CCGTTCTTCTTGGCTTCAGCAGTGTTCTTCGCAATCAAAGATGCAATCAAAGCTGCAAGAGCCGAAGTTGGGCATCACGAGTGGTTTCCTCTGGATAATC
CAGCAACACCGGAGAGGATCAGGATGGCTTGCCTAGATGAGCTCACGGAACCGCTCGTGGCCTCCGATTTTCGTCCTAAGCTTAGTGTCTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10034306 pacid=23176053 polypeptide=Lus10034306 locus=Lus10034306.g ID=Lus10034306.BGIv1.0 annot-version=v1.0
MGSLKNKDEVELAGDESAPKDGILYVNGVRKVLPDGLAHLTLLEYLRDIGQTGTKLGCGEGGCGACTVMVSQYDIRTKTCLHRAVNACLAPLYSVEGMHV
ITVEGVGTRRSGLHPVQESLASAHGSQCGFCTPGFVMSMYALLRSSREPPTEEQIEESLAGNLCRCTGYRPIVDAFRVFAKTDDALYTEANSKEGEFVCP
STGKPCSCKKNSSPVDNTACGERYSIASYSEVDGSKYIEKELIFPPELLMRKLTPLNLNGFNGLKWFRPVRLQHVLDLKAEHPDAKLVIGNSEVGIEMRL
KRLQYQTLIHVGHVPELNVLNVKEDGIEIGAAVRLTDLMQLLRKVANERVGHTTSSCKAIVEQLKWFAGTQIKNVASVGGNICTASPISDLNPLWMAARA
KFRIIDLKGSIRTTQADKFFLGYRKVDLQNGEILLSVFLPWTMKFEYVKEFKQAHRRDDDIAIVNAGMRVSLEKKGEEWVVLDASIAYGGVAPLSLSAVK
ASGFLVGKSWHQELFLGALQALETDIVIKEDAPGGMVEFRKSLTRSFLFKFFLWVSHQMEGDQCGGISVPLSHLSAVQSFSRPSVMGSQDYEIRKHGTSV
GSPEIHLSARLQVTGEAEYADDAPMPPNGLHAALVLSRKPHARIVAIDDSEAKSSPGFAGAFFAKDVPGDNHIGAIVHDEEVYATEFVTCVGQVIGVVVA
DTHENAKLAARKVHVEYKELPAILTIQEAVNAGSFHPHTEKWLKSGDVDACFGSGKCDKIVEGEVCIGGQEHFYLEPQSSYVWTLDSGNEVHMISSTQAP
QKHQKYVSHVLGIPMSKVVCKTKRIGGGFGGKETRSAFLAAAACVPSYLLNRPVKITLDRDVDMMITGQRHSFLGKYKVGFTNEGKILALDLKIYNNAGN
SLDLSLAILERAMFHSDNVYEIPNVRIVGNVCLTNLPSNTAFRGFGGPQGMLITENWIQRVAVEVNKSPEEIREMNFQGEGTILHYGQQLQHCTLSPLWS
ELKLSSDFLKAREDVHHFNRHNRWKKRGLAMVPTKFGISFTSKFMNQAGALVNVYTDGTVLVTHGGVEMGQGLHTKVAQVAASAFDIPLSSVFISETSTD
KVPNSSPTAASASSDMYGAAVLDACEQIKARMEPVACKHSFTSFAELAGACYAQRIDLSAHGFYITPDIGFDWTTGQGNPFRYFTYGAAFAEVEIDTLTG
DFHTRNVNVVMDLGYSLNPAIDVGQIEGAFVQGLGWVALEELKWGDEAHKWIAPGCLYTCGPGSYKIPSLNDVPFKFNVSLLKGHPNVKAIHSSKAVGEP
PFFLASAVFFAIKDAIKAARAEVGHHEWFPLDNPATPERIRMACLDELTEPLVASDFRPKLSV
|
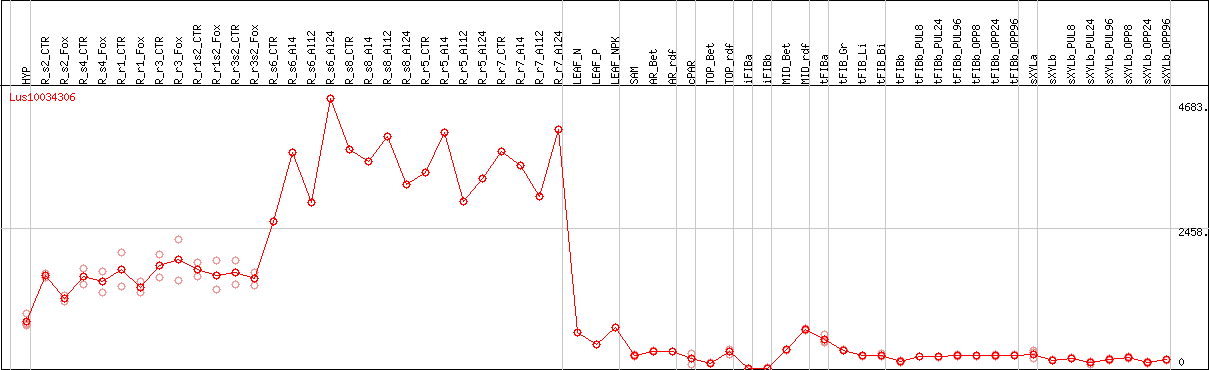
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034306 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
