External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G25970 85 / 5e-20
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT4G31350 72 / 2e-15
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2)
AT5G16170 70 / 1e-14
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G68380 67 / 1e-13
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT4G30060 66 / 2e-13
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT1G10280 66 / 2e-13
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT5G57270 66 / 2e-13
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1.2.3)
AT5G11730 64 / 1e-12
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT4G25870 63 / 2e-12
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
AT2G19160 63 / 3e-12
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005108
105 / 7e-28
AT1G10880 330 / 4e-109
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10034349
87 / 1e-20
AT1G68390 387 / 3e-131
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10035828
85 / 5e-20
AT1G68390 401 / 7e-138
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10036610
84 / 1e-19
AT1G68390 404 / 3e-139
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10005107
73 / 6e-16
AT1G68390 241 / 4e-89
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10031958
70 / 1e-14
AT1G68390 403 / 2e-138
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10013427
70 / 1e-14
AT1G10280 528 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10035145
70 / 1e-14
AT1G68390 393 / 6e-137
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Lus10040972
69 / 3e-14
AT1G10280 529 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G233400
92 / 1e-22
AT5G11730 557 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.015G045400
89 / 2e-21
AT1G68390 390 / 2e-134
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.018G059100
88 / 2e-21
AT5G11730 548 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.004G097700
81 / 9e-19
AT5G16170 452 / 6e-159
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.010G121900
75 / 1e-16
AT5G11730 409 / 1e-141
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.015G045500
73 / 9e-16
AT5G16170 394 / 2e-135
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.018G143900
71 / 7e-15
AT2G19160 564 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.008G123500
70 / 8e-15
AT1G68380 254 / 1e-82
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.010G121800
69 / 3e-14
AT1G68390 399 / 1e-137
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
Potri.003G001700
68 / 5e-14
AT1G10280 543 / 0.0
Core-2/I-branching beta-1,6-N-acetylglucosaminyltransferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF02485
Branch
Core-2/I-Branching enzyme
Representative CDS sequence
>Lus10034348 pacid=23175839 polypeptide=Lus10034348 locus=Lus10034348.g ID=Lus10034348.BGIv1.0 annot-version=v1.0
ATGGTCGATGCCGAAAGGAGGCTATTCGCTAACGCGCTCCTCGACTTCTCCAACCAACGGTTCATCCTCCTTTCCGAGTCTTGCATCCTTCTATTCGATT
TCACCACCATCTACAACTACCTAGTTTTCAATTTCTCCAACACGAGCTTCTTGGCTCGATTGATGATCCTCGACCCATGGCCCGAGGCAGATACAACAAA
CTCATGCAGCCAATCATCATGCTATCAGACTGGCGCAAAGGTGGGTCACATCCTACCATGGTTTTTGAGGAAGGATGTGAGCCAAGGGTTCTTGATGAGG
ATCAGAAACATGTCCGACTGTGATTATAGTGATCAAAACCATGGAACAATTTGTTTCCTCTTTGCTAGGAAGTTTCACCCTAGTTCTTTGGAGCCTTTGT
TGAGGATAGTTCCATCTTTGTTGGTTTTCAGCACTTAA
AA sequence
>Lus10034348 pacid=23175839 polypeptide=Lus10034348 locus=Lus10034348.g ID=Lus10034348.BGIv1.0 annot-version=v1.0
MVDAERRLFANALLDFSNQRFILLSESCILLFDFTTIYNYLVFNFSNTSFLARLMILDPWPEADTTNSCSQSSCYQTGAKVGHILPWFLRKDVSQGFLMR
IRNMSDCDYSDQNHGTICFLFARKFHPSSLEPLLRIVPSLLVFST
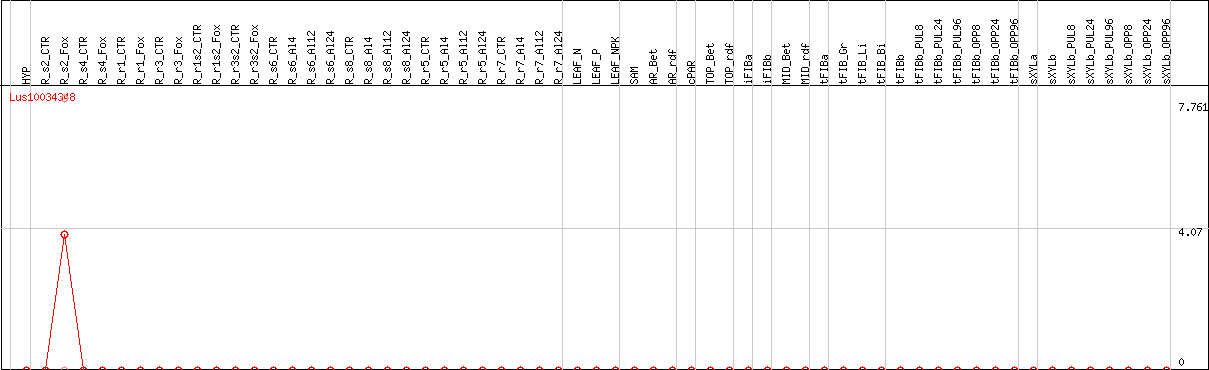
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034348 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.