Lus10034615 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10034615 pacid=23162740 polypeptide=Lus10034615 locus=Lus10034615.g ID=Lus10034615.BGIv1.0 annot-version=v1.0
ATGGAGGGAACGGAAGCAGAGGAAGAAGGAAACACCAAGTACACGTCATCAACAAACAGGAAGAGCAGCAAGAAGAAAAGGATTGAGCTAGGGATCTCAT
GTATGCTCAACACCGAGGTTGCCTCCGTCCTCGCCGTCCTCCGTCGCCCACCGCCCGATCCCACGGCGGCGCAGTTCCTTTCCCAAGAGGATCAGTACGA
TACCGCCCTCGTCCACTCTTTCAAATCCCTCCGTACATTGATTTTCAACCCTCCTCAGGACTGGCAGACCATGGACCCCGCCGTCTACCTCACGCCATTC
CTCGACGTCATTCAATCCGACGAGATTCCCTCTGCTGCTACCTCCATCGCTCTCTACTCCGTCCTTAAGCTCCTCAGCGTCGGAATCTTCGACGAGCGGA
CCACCGGAGCAAGGGACGCCANNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNN
NNNNNNNNNNNNNNNNNNNNNNNNNNNNNNCGGAATCTTCGACGAGCGGACCACCGGAGCAAGGGACGCCATCAACTCCGTCGTCATGGGGATCACCGGA
TGCCGATTGGAACGAACGGATCCCGTCTCGGAAGACGCCGTTATGATGCGGATCCTCCAGATCTTGACGGCGATCATGAAGCATCAGGCTGCCATTCTGT
TAACGGATCAGGCCGTGTGTACGGTGGTCAACACTTGTTTCCAGGTGGTTCAGCAATCGGCGAGTAGAGGGGATTTGCTTCAACGGAGTGCTAGGTATAC
GATGCACGAATTGGTCCGTATGATTTTTGCGAAATTGCAGGATATTGAAGTGAACCCTGGGGAGGAATCGGATACGGACATGGACGATGCCGATTTCGAT
GGGAATCTGGATTCAGGGTATGGAATTCGTTGCGCTGTTGATATCTTCCATTTCTTGTGTTCGTTGCTCAATGTTGTGGAAGTTGTGGAGAGTGAAGGGA
ATTTGTATCGTACTTCTGATGAGGATGTTCAGCTTTTCGCATTGGTATTGATAAATTCGGCCATCGAGTTGAGTGGAGACGTGGTTGGGAGTCACCCCAA
GCTTCTTCGGATGATCCAAGACGATTTATTCCATCATTTAATCCATTATGGAGCATCTTCCAGCCCCCTCATTTTGTCCATGATCTGTAGCACAGTTTTG
AACATTTACCATTTCCTTCGCAGGTTTGTTCGTCTTCAACTTGAGGCATTCTTCAAGTACGTGTTGTTGAGAGTTGCAGCAAATGGAGTTTCAACTCAGC
TTCAAGAAGTTGCACTTGAGGGAATCGTTGATTTTTGTAGGCAGCCAACTTTTATCATTGAAGCATATGTGAACTATGACTGTGATCCACTATGCATTAA
CATGTTTGAGGAAACTGCAAAGTTGCTCTGCAAGCTGTCATTTCCAGGAGCAAGTCTTGCAACGTCATTACAAGCTCAAGCATTTGAAGGTCTAGTTGGG
ATCATTCACAACATAGCCGAGAACATCAACAGGGAAGGTGATTCCACGCCTTCTGGACCGTACCCGGTTCACTTGACAGAGTACAGTAAGTTCTGGCAAG
ACAGGCCTTTGGAGAATAACGATGCATCTGTGGCATATATGAGGATAAGAAAAGTACAAAAGAGGAAAATTTTGGTTGCTTCAAACCATTACTCTAGAGA
TGAGAAGAAAGGGTTGGAGTATTTGAGGCTAAGTTTGATGTTTCCTAATCCTCCAGATCCTAAAGATTTTGCTTTCTTCTTTCGATTCACACCGGGGTTA
GACAAGACTATGATTGGAGATTACTTGGGTGATCCTGATGATTTCCACATTCAGGTTCTGAAAGAGTTCAGTTGTACTTTTGGGTTCTCAGGAATGGTTC
TTGACACTGCTCTCCGTACTTATCTTGCCACCTTCAGGTTGCCTGGAGAGTCCCAAAAAATCCAGAGAATAATGGAAGCTTTCTCGGATCAATTCTATGA
GCAACAATCTTCCGATATTTTCGCAACTAAAGATGCAGTTTTCGTCCTTTGTTACTCTCTCATTATGCTCAATACTGACCAGCACAATCCTCAGGTCAAG
AAGAAGATGACAGAGGAAGAGTTCATCAGGAACAATAGGGCTATCAATGCTGGACAGGATCTCCCTAGGGAATACCTCTCCGAGCTTTTCCAATCGATAT
CTTCTAAAGCAATTGCACTCTTCGGACAAACTGGACAGCAAGACATGAATCCTGGACTGTGGATAGAGCTGATGGGTGCTTCCAGATATTTGCAACCCTT
CACTCTTTATGATTTCGACCGTCGTCTCGGGAGAGATATGTTTGCCTCCGTTGCTGGTCCGGCAGTTGCAGCCCTCACCGCATTTTTCGAGCAATCTGAT
GAGGATGAAATGTTCCACGAATGCGTTGAAGCATTGATTTCGGTAGCCAGAATAGTACAGTATGGCCTGGAAGATACGCTAGACGAGATGCTTGCCTCAT
TGTGTAAATGTACCACACTACTGAATCCTTATGCATCGGCAGAAGAGACATTGTTTGCCTTTAGCCATGATCTGAAGCCCAGAATGGCTGTACTTGCAGT
CTTCAGCATTGCGAATAATTTCGGCTGTGGGATTCGAGGAGCGTGGAGGCCAATCTTGGACTGTCTGTTGAAGCTGAAAAAGCTCAAGTTGCTTCCTTCA
TCTGTAGTTGAGATACACAATTCTTTAGCGCCTTCCGTTGATTCGTCATCTGACCATTCTAGAAATGACGGTGGCTTAATTTTCCCATCTCATGAATCAA
AATCATCAGGTGGCAGCAATCGCCAGAGCTCAACCATGATTGGCAGATTCTCTCAATTTTTGTCTCCTGACAGTATGGAAGAATCCTTGTCTCTCGGCCT
GAATGAATTCGAGCAGAATCAAAAAGTTATCAAGCAGTGTCAGATCGGGAGCATATTCACCAACAGCTGGAACTTGTCGGATGAAACTCTAGCATATCTA
GGTCGTACTCTAATCTTTGCATCAGCAGGCAAAGGGCAGAAATTTGCGACGCCAATCGAGGAAGAAGAAACCATAGAATTCTGTTGGGATTTGATAGTTG
CTCTATCTTTTGCCAACATTAACCGGTTACATAACTTCTGGCCAGCCTACCATGAAAACTTGTTTGAAGCTATTCACTTTCCTCTATTCTCCTCAATCCC
ATTTGCAGAGAAGGTCATAATTGCCCTCTTCAGGATCTGCCTCAAGCTACTCTCATCTGCCAGAGTAGAGAAACTGCCAGAGGAACTCATCTTCAAATCC
ATCAACATGATGTGGCAAATAGAGAAGCAGATTCTAGACACATGTTGTGAATCCATAACAAAGTCAGTCAGCAAGATTTTGTTTGAATACCCTGCAAATC
TGCAGAGCTTCCTGGGTTGGAAAACAGTCCTCCATCTCTTATCCGTCTCTGGACGCCATCAGGAAACCTACGATCAGGGAGTAGAGACGCTTATCATGCT
CGTTTCGGACCAAACTCATGTTTCCCCCGTGAATTACCCTTTCTGCATCGACTGTGCATTTGGGTTCGTGGCGCTTAAGAACAGTCCCCTGGAGAAAAAC
CTGAAGATTCTGGACCTTCTGGCGGAATCGGTCAGCTTGCTAATCCATTGGTATAAAAACTTCTCAGACCCTGGTAGCAATTTCAGTACAGCAAGCACAG
GAAGCAATAACTCTTTCCAGGATGCCGCTTCGTTTTCGGTCAGCTTGTTCCTGAAATTAGGAGAAGCCTTCAGGAAAACCAGCTTGGCAAGACGAGAGGA
AATCAGGAACCATGCCATCAAGTCTCTTCATAAGAGCTTTTCCTTAGCAGAGGAGTTGGATTTCACACCTGCCAACTGCCTAAACTTGTTCAACTTAGTG
GTATTTGCCATGGTAGACGATCTGCACGAAAAGCTGATAGAATACTCGAGGCGGGAAGGGGCGGAGAGGGAGATGAGGAGTATGGAAGGGACTTTGAGCT
TGGCGATGGAACTTATGACGGAGGTTTATCTACAGTTCCTGAAGCCAATAGCATCAAGCCCCGGGTTCAGGACGTTCTGGTTGGGTCTATTGAGAAGGAT
GGATACATGTATGAAAGCTGATCTTGGGGAGTATGGTGAGACCAAGTTGCAGGAGGTGATTCCAGGTCTCTTGAAGAAAATGATTATTAAGATGAAAGAA
GATGAAATTTTGGTACAGAAGGAAGATGATGATCTGTTTGAAATAGCTTACATTCAAATTCAGTGGATTGCTCCTGCCTTGAAGGAAGAACTTTTTCCTG
AATTGGAAATTTAG
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10034615 pacid=23162740 polypeptide=Lus10034615 locus=Lus10034615.g ID=Lus10034615.BGIv1.0 annot-version=v1.0
MEGTEAEEEGNTKYTSSTNRKSSKKKRIELGISCMLNTEVASVLAVLRRPPPDPTAAQFLSQEDQYDTALVHSFKSLRTLIFNPPQDWQTMDPAVYLTPF
LDVIQSDEIPSAATSIALYSVLKLLSVGIFDERTTGARDAXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX
XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXGIFDERTTGARDAINSVVMGITG
CRLERTDPVSEDAVMMRILQILTAIMKHQAAILLTDQAVCTVVNTCFQVVQQSASRGDLLQRSARYTMHELVRMIFAKLQDIEVNPGEESDTDMDDADFD
GNLDSGYGIRCAVDIFHFLCSLLNVVEVVESEGNLYRTSDEDVQLFALVLINSAIELSGDVVGSHPKLLRMIQDDLFHHLIHYGASSSPLILSMICSTVL
NIYHFLRRFVRLQLEAFFKYVLLRVAANGVSTQLQEVALEGIVDFCRQPTFIIEAYVNYDCDPLCINMFEETAKLLCKLSFPGASLATSLQAQAFEGLVG
IIHNIAENINREGDSTPSGPYPVHLTEYSKFWQDRPLENNDASVAYMRIRKVQKRKILVASNHYSRDEKKGLEYLRLSLMFPNPPDPKDFAFFFRFTPGL
DKTMIGDYLGDPDDFHIQVLKEFSCTFGFSGMVLDTALRTYLATFRLPGESQKIQRIMEAFSDQFYEQQSSDIFATKDAVFVLCYSLIMLNTDQHNPQVK
KKMTEEEFIRNNRAINAGQDLPREYLSELFQSISSKAIALFGQTGQQDMNPGLWIELMGASRYLQPFTLYDFDRRLGRDMFASVAGPAVAALTAFFEQSD
EDEMFHECVEALISVARIVQYGLEDTLDEMLASLCKCTTLLNPYASAEETLFAFSHDLKPRMAVLAVFSIANNFGCGIRGAWRPILDCLLKLKKLKLLPS
SVVEIHNSLAPSVDSSSDHSRNDGGLIFPSHESKSSGGSNRQSSTMIGRFSQFLSPDSMEESLSLGLNEFEQNQKVIKQCQIGSIFTNSWNLSDETLAYL
GRTLIFASAGKGQKFATPIEEEETIEFCWDLIVALSFANINRLHNFWPAYHENLFEAIHFPLFSSIPFAEKVIIALFRICLKLLSSARVEKLPEELIFKS
INMMWQIEKQILDTCCESITKSVSKILFEYPANLQSFLGWKTVLHLLSVSGRHQETYDQGVETLIMLVSDQTHVSPVNYPFCIDCAFGFVALKNSPLEKN
LKILDLLAESVSLLIHWYKNFSDPGSNFSTASTGSNNSFQDAASFSVSLFLKLGEAFRKTSLARREEIRNHAIKSLHKSFSLAEELDFTPANCLNLFNLV
VFAMVDDLHEKLIEYSRREGAEREMRSMEGTLSLAMELMTEVYLQFLKPIASSPGFRTFWLGLLRRMDTCMKADLGEYGETKLQEVIPGLLKKMIIKMKE
DEILVQKEDDDLFEIAYIQIQWIAPALKEELFPELEI
|
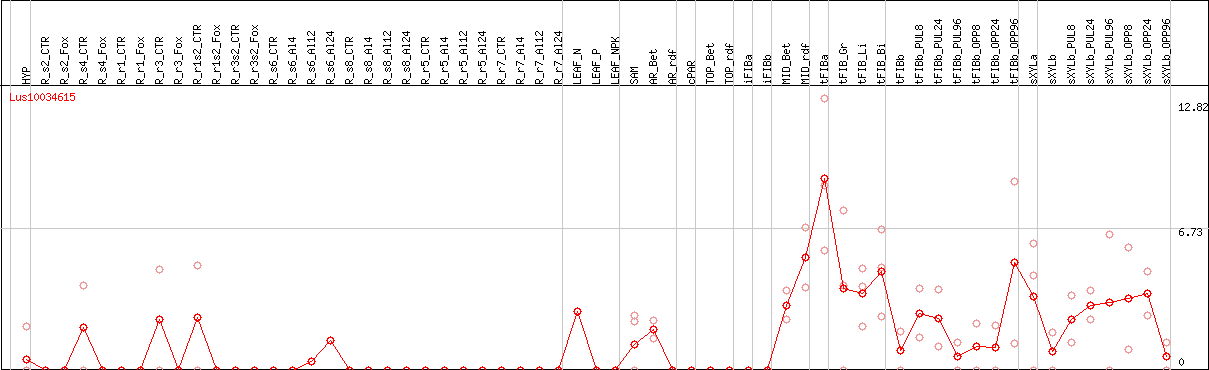
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034615 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
