External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G51980 217 / 4e-67
C3HZnF
Transducin/WD40 repeat-like superfamily protein (.1.2)
AT4G25440 208 / 1e-63
C3HZnF
ZFWD1
zinc finger WD40 repeat protein 1 (.1)
AT5G49200 172 / 5e-50
C3HZnF
WD-40 repeat family protein / zfwd4 protein (ZFWD4) (.1)
AT5G40880 166 / 2e-47
WD-40 repeat family protein / zfwd3 protein (ZFWD3) (.1)
AT2G19430 55 / 3e-08
DWA1, AtTHO6
DWD (DDB1-binding WD40 protein) hypersensitive to ABA 1 (.1)
AT3G51930 53 / 1e-07
Transducin/WD40 repeat-like superfamily protein (.1)
AT1G47610 52 / 3e-07
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G08390 52 / 5e-07
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G21040 51 / 7e-07
FBX2
F-box protein 2 (.1.2)
AT5G50230 50 / 2e-06
Transducin/WD40 repeat-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021648
401 / 1e-138
AT4G25440 321 / 1e-105
zinc finger WD40 repeat protein 1 (.1)
Lus10021656
357 / 5e-121
AT4G25440 329 / 9e-109
zinc finger WD40 repeat protein 1 (.1)
Lus10032066
322 / 5e-108
AT4G25440 343 / 5e-115
zinc finger WD40 repeat protein 1 (.1)
Lus10014595
301 / 4e-96
AT5G14250 445 / 3e-150
FUSCA 11, COP9 SIGNALOSOME SUBUNIT 3, CONSTITUTIVE PHOTOMORPHOGENIC 13, Proteasome component (PCI) domain protein (.1), Proteasome component (PCI) domain protein (.2)
Lus10001652
281 / 2e-95
AT5G51980 192 / 2e-59
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10021655
280 / 2e-92
AT4G25440 169 / 3e-48
zinc finger WD40 repeat protein 1 (.1)
Lus10021640
262 / 1e-87
AT4G25440 185 / 7e-57
zinc finger WD40 repeat protein 1 (.1)
Lus10032062
255 / 8e-83
AT4G25440 212 / 3e-65
zinc finger WD40 repeat protein 1 (.1)
Lus10033255
229 / 5e-71
AT5G51980 285 / 3e-91
Transducin/WD40 repeat-like superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G134800
206 / 6e-63
AT4G25440 600 / 0.0
zinc finger WD40 repeat protein 1 (.1)
Potri.015G137200
200 / 2e-60
AT4G25440 587 / 0.0
zinc finger WD40 repeat protein 1 (.1)
Potri.014G077800
198 / 9e-59
AT5G51980 391 / 5e-132
Transducin/WD40 repeat-like superfamily protein (.1.2)
Potri.009G109500
57 / 1e-08
AT3G18950 555 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.002G130400
56 / 1e-08
AT2G26490 632 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.007G009500
52 / 2e-07
AT3G49660 469 / 4e-168
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
Potri.001G022900
52 / 3e-07
AT3G51930 523 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
Potri.009G058000
51 / 5e-07
AT5G52820 811 / 0.0
WD-40 repeat family protein / notchless protein, putative (.1)
Potri.012G060800
52 / 6e-07
AT1G49040 1782 / 0.0
STOMATAL CYTOKINESIS-DEFECTIVE 1, stomatal cytokinesis defective / SCD1 protein (SCD1) (.1), stomatal cytokinesis defective / SCD1 protein (SCD1) (.2), stomatal cytokinesis defective / SCD1 protein (SCD1) (.3)
Potri.003G202700
51 / 6e-07
AT3G51930 531 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Lus10034716 pacid=23162918 polypeptide=Lus10034716 locus=Lus10034716.g ID=Lus10034716.BGIv1.0 annot-version=v1.0
ATGGTGCTCAAATCGAATGTGTTCGAACGACTTGGGGTTTCTGGACATGATCAGAAGAATAGTCGGACCGTTTGTCGATTCTGGAAAGTGGGAAATTCCA
CTCATCAACCCTGCAGATACTTCCACGGTGAATCTCCTTCTGCGCTCCCGTCTCCATCACCGTACCGTCGTTCTAACGTCTACAATCGAGGAGGATCTCA
GTACTCGTCACAGCACCTCGTTCCAATGTCTACATTCGACCATCTCGATCAAAACCACGGTCGTCTACTCCTGGCTCTCTCTTGCTTCTGGGGTTCTAAG
CTTTTCTGTGGAAGCAGTGATGGGGCTCTTCAAACATGGAATTCCATAAGTGGTGAAGTTGTAGGGTCACTGAAAGTTGGTGGGAAGATTGGGTGTATGA
TTTTGGAGGGTGATTGGCTATATGTTGGATTGACAAGTATGGTCAAGGCATGGAACATTCACTCAGGTGTTCAACACATTCTCGAAGGCCCAGTTGGCCA
AGTTCACGCATTGACTGTTGACTATGAGATGGGGATTGTGTTTGCTGCCGTTGAAGATGGTATCATTTGGGCGTGGAAGACCAGTACCGATGAGCAAGGA
AATCTCTTCCAGTCAGTTGCATCACTGATTGGCCATACAATTGCAGTTATTTCCTTGAGATTTGGAGCAGGTAGTAGGGTGTATTCGGGTTCCATAGATG
GAACCATCAAGGTTTGGGATGGTAGAATTTCTCAATGCATACAAACATTAAAGGGGCATGATGATGCAGTGACGTGTCTCCTCTGCTATAGATGCTACTT
CTTGTCCTGCTCACCGGATAAGAAGATCGAAGTATGGGAAGAGTGTGGGAAAGAGATGTTGTTGTGCTCTTGGAACAATGATTCGGTTGGAATGTATGAG
GTTCCATCATTTGTTGAGAAAGGGAGAATGTACTCGAAATGTGAAGTAAGGAGCTTGGGAGTAGGTACTGAAGGAATGTTCTTCACTGGAGATGCTTCTG
GTGTGGTGACTGTATGA
AA sequence
>Lus10034716 pacid=23162918 polypeptide=Lus10034716 locus=Lus10034716.g ID=Lus10034716.BGIv1.0 annot-version=v1.0
MVLKSNVFERLGVSGHDQKNSRTVCRFWKVGNSTHQPCRYFHGESPSALPSPSPYRRSNVYNRGGSQYSSQHLVPMSTFDHLDQNHGRLLLALSCFWGSK
LFCGSSDGALQTWNSISGEVVGSLKVGGKIGCMILEGDWLYVGLTSMVKAWNIHSGVQHILEGPVGQVHALTVDYEMGIVFAAVEDGIIWAWKTSTDEQG
NLFQSVASLIGHTIAVISLRFGAGSRVYSGSIDGTIKVWDGRISQCIQTLKGHDDAVTCLLCYRCYFLSCSPDKKIEVWEECGKEMLLCSWNNDSVGMYE
VPSFVEKGRMYSKCEVRSLGVGTEGMFFTGDASGVVTV
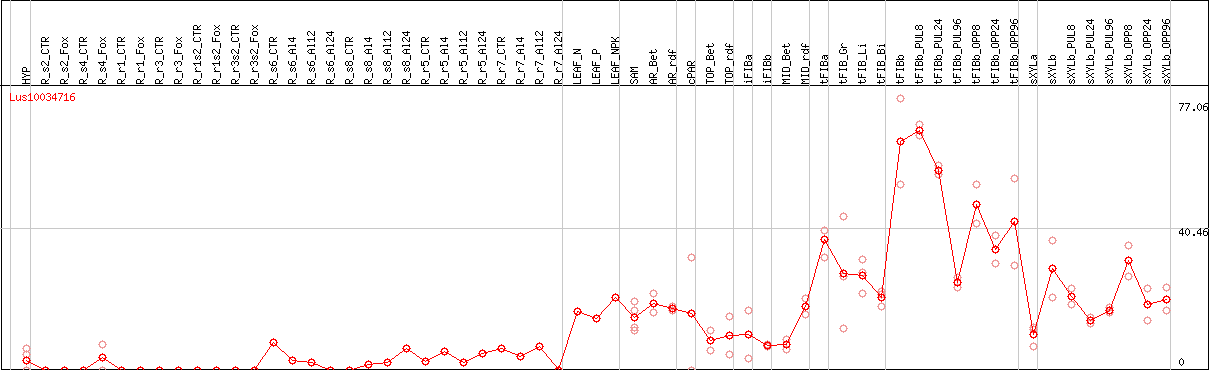
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034716 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.