External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G15450 155 / 7e-45
Senescence/dehydration-associated protein-related (.1)
AT3G21600 152 / 1e-43
Senescence/dehydration-associated protein-related (.1.2)
AT4G35985 81 / 3e-17
Senescence/dehydration-associated protein-related (.1)
AT3G51250 66 / 7e-12
Senescence/dehydration-associated protein-related (.1)
AT2G17840 51 / 3e-07
ERD7
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012957
413 / 3e-146
AT4G15450 321 / 1e-108
Senescence/dehydration-associated protein-related (.1)
Lus10011127
174 / 6e-52
AT4G15450 382 / 8e-132
Senescence/dehydration-associated protein-related (.1)
Lus10043028
168 / 1e-49
AT4G15450 373 / 3e-128
Senescence/dehydration-associated protein-related (.1)
Lus10014132
72 / 4e-14
AT2G17840 476 / 1e-166
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
Lus10005067
66 / 6e-12
AT2G17840 459 / 5e-160
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
Lus10028439
66 / 7e-12
AT3G51250 520 / 0.0
Senescence/dehydration-associated protein-related (.1)
Lus10027840
63 / 7e-11
AT2G17840 489 / 4e-169
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
Lus10041892
49 / 2e-06
AT3G51250 358 / 9e-122
Senescence/dehydration-associated protein-related (.1)
Lus10034499
45 / 2e-05
AT2G17840 306 / 2e-103
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G164748
196 / 1e-60
AT4G15450 332 / 3e-112
Senescence/dehydration-associated protein-related (.1)
Potri.018G087500
191 / 1e-58
AT4G15450 401 / 5e-139
Senescence/dehydration-associated protein-related (.1)
Potri.005G111600
78 / 3e-16
AT2G17840 491 / 3e-172
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
Potri.004G174100
70 / 2e-13
AT2G17840 472 / 7e-165
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
Potri.009G133500
60 / 5e-10
AT2G17840 449 / 1e-156
EARLY-RESPONSIVE TO DEHYDRATION 7, Senescence/dehydration-associated protein-related (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF06911
Senescence
Senescence-associated protein
Representative CDS sequence
>Lus10034970 pacid=23181350 polypeptide=Lus10034970 locus=Lus10034970.g ID=Lus10034970.BGIv1.0 annot-version=v1.0
ATGGACGAAGGGGAAACTCTCGAACTCGCTTCCGGCGGGTTTCTAATAGCTCGAGTCGTGGACAATAAGGGAGGATTCCCTATTGCTACAGTGGTTAAAG
TTGGGAAAGACCTTCAGTGGCCACTGACGAGAGACAAGCCTGTCGTAAAGCTGTATTCTTTCCACTACTTGTTCTCTCTGCCCATGAGAGACGGCTGTCC
GTTGAGCTACTGCCTGACGTTTTCGAAGCAGCAAGGGAAGAACTTAGCGGCGTTGGATTCCTTTCTTGACCAAAATTCGTGCTTCTCTGCCTCTGGTTAT
GGTTCTGATTCGGAATTGGTTAGGAATGATGGAGGGAGACAAGAGCAGGGAATTGCGGAGGGTACTCTTCAGATTGTCAGGGGAATCTTCATGTGCAGCA
ACGTTTACAGCAACTTGGTTAAGTGTGCGGAGATGATTCTGAAAGGGGGCGAAATCAACGGCTACAAAAAACCAGTAGTAGCACAAAATGGGAACATTGG
AGCCAAAAAGGGGTTAAAGAGGGCTCGAGAGTTATCAAAGATGACAGAAGATGAGCAAAGCGATTTTGGAAGGAGCGAATATGATAAGATATTGGACGCA
GCGGAGAAGCAAGCGCTTGCTGCCACCACTGCCGCCACGACAAGGGTGGTCAGCCAGAGGTACAGCGAAGGAGCGCCGGAGACAACTGGGGATGTGCTGG
GCACTGCAGGTCACTGCGCTAACACGGCGTGGAATCTTGTTCAATTCAGGAAGGCTATCAATCCAGCTTCCTCTGTGGCCACGGGAGTTCATAGGAACAA
CAACAGAAACTAG
AA sequence
>Lus10034970 pacid=23181350 polypeptide=Lus10034970 locus=Lus10034970.g ID=Lus10034970.BGIv1.0 annot-version=v1.0
MDEGETLELASGGFLIARVVDNKGGFPIATVVKVGKDLQWPLTRDKPVVKLYSFHYLFSLPMRDGCPLSYCLTFSKQQGKNLAALDSFLDQNSCFSASGY
GSDSELVRNDGGRQEQGIAEGTLQIVRGIFMCSNVYSNLVKCAEMILKGGEINGYKKPVVAQNGNIGAKKGLKRARELSKMTEDEQSDFGRSEYDKILDA
AEKQALAATTAATTRVVSQRYSEGAPETTGDVLGTAGHCANTAWNLVQFRKAINPASSVATGVHRNNNRN
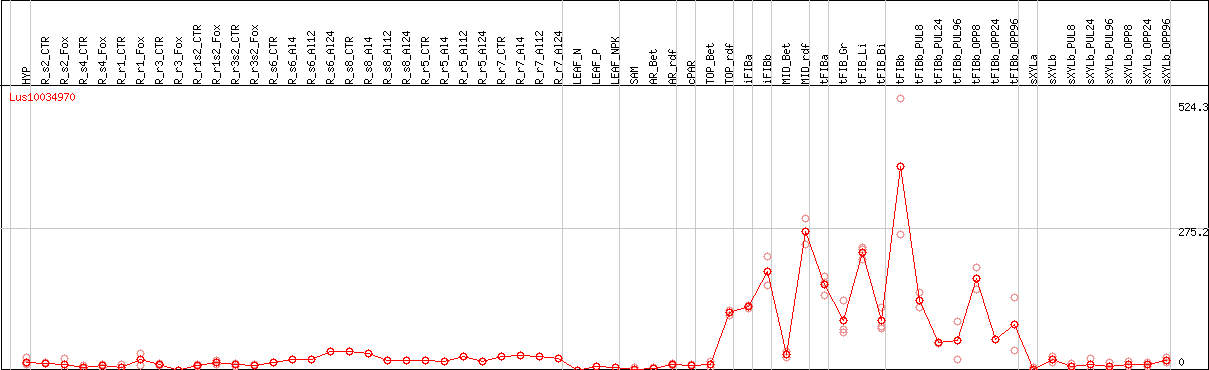
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10034970 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.