External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G26782 219 / 8e-69
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G30700 179 / 5e-53
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G57430 177 / 3e-52
OTP84
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G08070 174 / 2e-51
EMB3102, OTP82
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G27610 173 / 6e-51
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G37380 170 / 1e-50
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G02750 171 / 2e-50
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G23330 171 / 2e-50
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G41080 166 / 2e-49
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G22070 168 / 3e-49
pentatricopeptide (PPR) repeat-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031989
300 / 3e-99
AT3G26782 865 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10028837
187 / 2e-58
AT1G11290 650 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10017446
185 / 3e-55
AT1G11290 1152 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10040577
176 / 9e-55
AT1G25360 577 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10038400
178 / 1e-52
AT3G57430 1097 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10001220
178 / 2e-52
AT3G57430 1110 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10015414
177 / 3e-52
AT3G57430 629 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10013991
177 / 4e-52
AT3G57430 632 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10000706
174 / 3e-51
AT3G12770 878 / 0.0
mitochondrial editing factor 22 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G322100
250 / 2e-80
AT3G26782 897 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G075800
194 / 1e-58
AT3G57430 1189 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 84, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.018G040100
182 / 1e-54
AT1G08070 974 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.001G354400
179 / 1e-53
AT5G46460 863 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.017G091600
177 / 8e-53
AT4G37170 924 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.016G075000
175 / 3e-52
AT2G41080 755 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G191000
176 / 4e-52
AT1G11290 1178 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G005700
175 / 5e-52
AT3G49142 912 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.002G220300
175 / 5e-52
AT1G11290 559 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G027800
176 / 6e-52
AT1G25360 1047 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10035157 pacid=23162592 polypeptide=Lus10035157 locus=Lus10035157.g ID=Lus10035157.BGIv1.0 annot-version=v1.0
ATGAGAGCGCTCATGAAAGATAGTGGGCTATTCAAGACGCCTGGTTTCAGTCTAACTGAACTCAAGGGCGACGTGCATGTTTTCCTGGTTGGAGATAAAG
ACCATCCTCAACACAAGAAGATTTATCAGTATTTGGAAAACCTTGATGTAAGGCTGCAGCATGCTGGCTATGTACCTGATATGGCTTCTGTCCCTCATGA
TGTTGACGAGGAAGAGAAGGGAATAAATTTGCGAGGTCATAGTGAGAAATTAGCGGTTGCTTTTGGGATCATTAACTCAGTTCCTGGAGCAACAATTCAT
GTCATGAAGAATCTTCGGGTCTGTGGTGATTGCCACACTGTGATTAAGCTGATATCCAAGATTGAAAACCGGGAAATTACCGTGAGAGATTCAAAGCGGT
TTCATCACTTCAAGGATGGTGTTTGTTCATGTGGAGATTATTGGTAA
AA sequence
>Lus10035157 pacid=23162592 polypeptide=Lus10035157 locus=Lus10035157.g ID=Lus10035157.BGIv1.0 annot-version=v1.0
MRALMKDSGLFKTPGFSLTELKGDVHVFLVGDKDHPQHKKIYQYLENLDVRLQHAGYVPDMASVPHDVDEEEKGINLRGHSEKLAVAFGIINSVPGATIH
VMKNLRVCGDCHTVIKLISKIENREITVRDSKRFHHFKDGVCSCGDYW
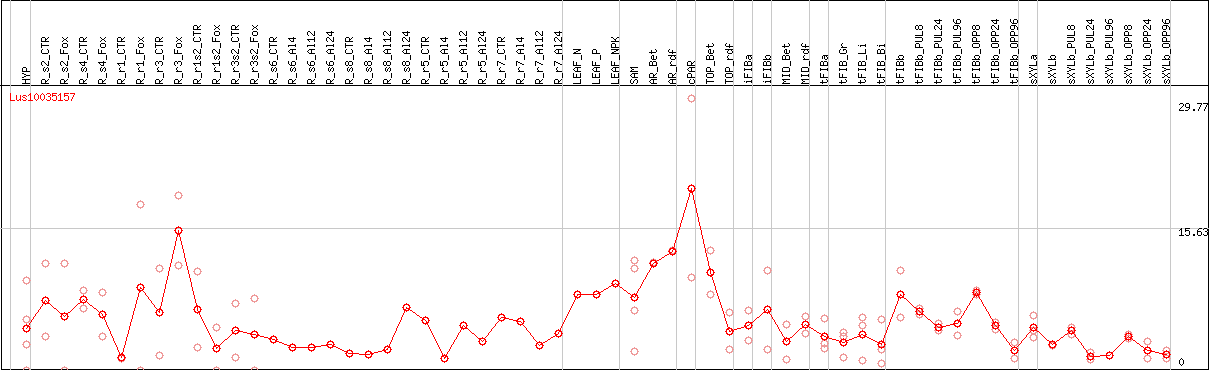
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10035157 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.