External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G45940 39 / 0.0003
Glycosyl hydrolases family 31 protein (.1)
AT1G68560 38 / 0.0007
AXY3, TRG1, XYL1, ATXYL1
thermoinhibition resistant germination 1, altered xyloglucan 3, alpha-xylosidase 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031003
162 / 4e-47
AT3G45940 97 / 3e-20
Glycosyl hydrolases family 31 protein (.1)
Lus10008067
41 / 0.0001
AT5G11720 1072 / 0.0
Glycosyl hydrolases family 31 protein (.1)
Lus10041457
40 / 0.0002
AT1G68560 1455 / 0.0
thermoinhibition resistant germination 1, altered xyloglucan 3, alpha-xylosidase 1 (.1)
Lus10035020
40 / 0.0003
AT1G68560 1238 / 0.0
thermoinhibition resistant germination 1, altered xyloglucan 3, alpha-xylosidase 1 (.1)
Lus10038408
39 / 0.0004
AT5G11720 1150 / 0.0
Glycosyl hydrolases family 31 protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G039500
106 / 1e-27
AT5G63840 74 / 3e-13
RADIAL SWELLING 3, PRIORITY IN SWEET LIFE 5, Glycosyl hydrolases family 31 protein (.1)
Potri.011G154500
40 / 0.0001
AT5G11720 1215 / 0.0
Glycosyl hydrolases family 31 protein (.1)
Potri.011G154300
40 / 0.0001
AT5G11720 1194 / 0.0
Glycosyl hydrolases family 31 protein (.1)
Potri.008G120000
39 / 0.0003
AT1G68560 1465 / 0.0
thermoinhibition resistant germination 1, altered xyloglucan 3, alpha-xylosidase 1 (.1)
Potri.001G442800
39 / 0.0003
AT5G11720 1167 / 0.0
Glycosyl hydrolases family 31 protein (.1)
Potri.010G125800
39 / 0.0004
AT1G68560 1422 / 0.0
thermoinhibition resistant germination 1, altered xyloglucan 3, alpha-xylosidase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0058
Glyco_hydro_tim
PF01055
Glyco_hydro_31
Glycosyl hydrolases family 31
Representative CDS sequence
>Lus10035399 pacid=23170596 polypeptide=Lus10035399 locus=Lus10035399.g ID=Lus10035399.BGIv1.0 annot-version=v1.0
ATGATTGGAGTGGAACTTCTGGGAGTGACAACCATTTCAGTGACGTTTAGATCTCGATCGGAGGCTCTGATACCATTTCAGATGAGCTCAATCGAGCGAG
AGGGAGGGAGGAAGGAAGAAAGAGAAGCGTCAGAGAAAGGGCTGCCGATATGCCGACACATGGTCCTCCACTACCCGAGCGACAAGACCGTGGGGAAGCT
TAGCTACGAGCAGTTCCTGGTCGGAACAGAGATCCTGGTGGTGCCTGTTTTGGATAAAGAGAAACAGGAAGTGAAGGCCTATTTCCCAAAGGCTGATCAA
CGTTTTTCGACATCCTCTTCTTGGTTGCATGTTTGGTCGGGGAAAGTGTATGATGAAGTGGGTTGTGATGCTTTGGTTTAA
AA sequence
>Lus10035399 pacid=23170596 polypeptide=Lus10035399 locus=Lus10035399.g ID=Lus10035399.BGIv1.0 annot-version=v1.0
MIGVELLGVTTISVTFRSRSEALIPFQMSSIEREGGRKEEREASEKGLPICRHMVLHYPSDKTVGKLSYEQFLVGTEILVVPVLDKEKQEVKAYFPKADQ
RFSTSSSWLHVWSGKVYDEVGCDALV
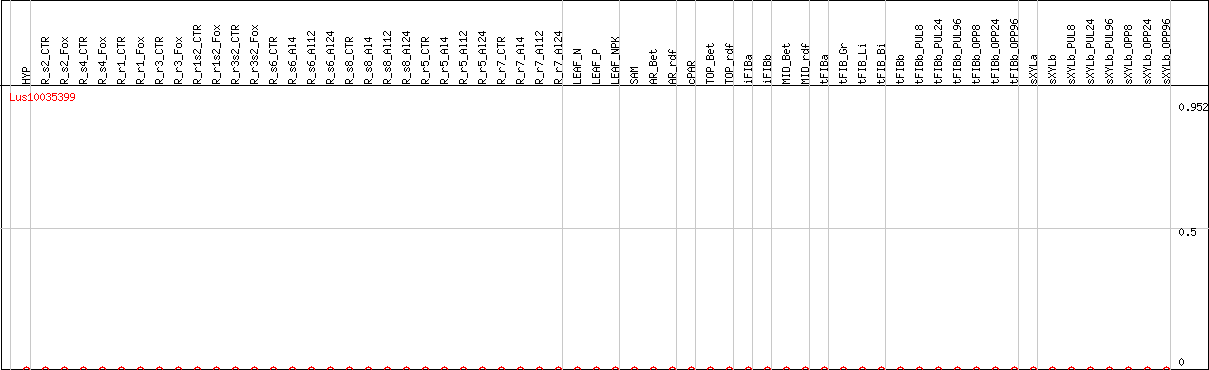
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10035399 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.