Lus10035417 [FLAX]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10035417 pacid=23170503 polypeptide=Lus10035417 locus=Lus10035417.g ID=Lus10035417.BGIv1.0 annot-version=v1.0
ATGAGCCCAGACAAGGTTGAATCTGGGAATGCTACAGACTCTGATCAGTTCTCTCTCAACTCTGACACCTTGTCTGTCACCTCAGATCAATCCTCATCCT
CATCTTCTTCTTCGTCTTCAACAATTCCCCACCAGAAACCACCAAATTACATGAAGACCACCACTGCCTCTGTTGCAAAGAACACCAACGATTCCCAGAG
ATCAAAGGGTTTGGAAAATCCAAAATTCAGGATGAGGAGATCTTTCAGAAGAAGACCAGATTCCAGCATCAAGATAGTTGATAATCCAGCATCAGATGAT
GATTCATCTTCTTCTGGTTCTGAGGGTGATGCTAAATCTGATTCCAAAACATCTGTTGCGATGAGAAGAACATCTGGTTCAAGACTAACAAAGGTAGCAA
GTTTGAGAATCAGGAGGCAATCCACTGTAGCGAGATCCACTTGTTCATCGGCTCTAAGAGATTCCAAACTTGCCACCAAGCATGAAGAAAACAACAGCAA
CTCCAATGAAGGAACTGCAACAGAAAAGGTTTGTCCATATAGCTACTGTTCATTGCACGGACATCACCATTCCAGCGATCAGGTTCCTATTACATTGAAG
CGATTCGTCTCGATGAGGCGAAGATTGTTGAAGCCACAAAAGAAGACCAACAACTCATCAGATTCCAGGAAAAAGAAGCTAACGAATCAAACAAAGAAAG
GGGTTGTCGTTTCTGGAAAAAGTGGGAAAGCAGCTAATGGCGGCAGACAAAAATCTTGTGTTGTTAAGGCAAATGGTGGATTATTAGAGACTGATAGCTC
CGATGTTTTTCAAGATAAGGACTTGGTCGTTTCCGCAGATTATAAGCCTGATGAGGATGGTGGGATTGTACATCTTTCATCTGAAAACTCAGTGGAGAAG
AACAAGAAGAACATGGGATTATGGGGAGTGATCTACCAGCACATGGCATCAGGCATTGCTGCAGAAGGTGATGAATCCAGCCAGGCTGAGAAGACCGGAA
TGGATCGAAACAAAGATCCGGAAGAGGGGAACGGAGGAATCAGGAATGTCGATTCGCACCAGCTTGATGCAATCAAGCTTGTCCAGGAAGCATTTGACAA
GATCCTCTCAGAAATCCCAGATCCTTTGTCTGATAATCAGTCAGTTGATTATAGTGATGCCGGTTCAGATGCTAGTAGTGCTCCAACTTGTGATGATTCG
CAAAAAGTCAGAATCGAACAAGAACAAATGGAGCTGAAAAAGGCTGAAACTTTCAATGATTCAGAATCAAAGGAGGGAAACAAATCTGCTGCTAAGAAGA
AGGCCAACGGTTGGAGCAACCTGAAGAAGATTATGATAATGAAGAGATTTGTCAAGGCGATGGATAATGTCAAGAACTTCAATCTACGTAAGCGTCAACA
AGATCATCTTCGGTGTACGGAGCAAGAGGGGGGAGCTGAAAAGGTTAATCTGCGGCCACAGACGATTGGGGAGAAGAAGAACTCAGAGGAATGGATGCTT
GATTATGCTCTTCAAAAGGTGATTTCAACACTGGCTCCAGCTCAGAAGAGGAAAGTGGCTCTGATTGTACAAGCATTCGAAACAGTCGCGCCTCAGGGAG
ATACTGATGCTGCAGCTTCTTCTACTCCCAAAACTCCAGTTCATAAGGAACCTGAGCCAGAACAGAACATAATATTCGGGGTCTTACTTCGAAAAGTATC
CACTGTTCCGAAAGAGAGTCGTAATAATGCTGATCGCGGTGTCAGTGATTCTTACTTGATGAAGAGCAGATTGAAGGCAATTGATCAGATTGCTGACGAC
AATAGTATTGATTATGTTACCACTGTTCCGAAAGAGAGTCGTAATAATGCAGATGGTACAGATTGCGATGTCGGTGGTTCTGATTTGATGAAGAGCAGAT
TGAAGTCAATTGATCAGATTGCTGAAGAACCAATTGCTGAAGACAATAGTGTTGATAATGTTACCTTAGCTCCTGCAGCAGAAAAAGAAGAAGAACCTCC
GGTCGAAGACGTTCAGCCATCCATTGAAGCTGAAAATGATGAGGAGCTTGATGTTGATAGAAATGCAGCGGAACCTGAAGCAGAGATGGGGGGGAACCAG
AAACATAAGAATCTGTGGTTTCTGATATACAAGCACATGGTTTCAGGAACAACACAACCACTTCAGGAATCCGGTGAAGAAGTCAAAGGAGGAAGAAACG
AGTCCCGGTCACAGACTGAAGCCATAAGGCTTGTAGAGGAGGCGATCGACGAAATTCCTCTTCCGGAGACTCAAGATGATGCACCACCTGCTGATGACGA
TCATCCATCAGTAGCCACCAGAGACATAAATTCCCAAGTAGAAGATAAACATGATGAAAGAACAGAGATAGAAACCACAGTTGAAGACCAGAAAGAAGAA
GCAAGGCCAGCAGCGAAGAAGAGCTGGAGCAATCTGAAGAAGGTGATCTTACTGAAGAGATTCATCAAGTCATTGGAGAAAGTGAACGACACCTGCTTGA
CTCCGTCGTCGAGAAAGTTCATCAACCCCAAGGAACCGAAGTTTCTTCCTTCGGTGGATGGGCAAGAAGCTGAGAAGGTTTTGCTGAGGCGCCAAGATGT
TGGAGACAGGAAAAGTGGTGATGAATGGATGCTTGATCATGCTCTTCGACAGGTTGTAGCCCAGCTGACACCGGTTCGTCGAAGAAAAGTTGAGCTGCTT
GTAGAGGCATTTGAGACTGTCAGTCCCACCATTGGCAACTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Lus10035417 pacid=23170503 polypeptide=Lus10035417 locus=Lus10035417.g ID=Lus10035417.BGIv1.0 annot-version=v1.0
MSPDKVESGNATDSDQFSLNSDTLSVTSDQSSSSSSSSSSTIPHQKPPNYMKTTTASVAKNTNDSQRSKGLENPKFRMRRSFRRRPDSSIKIVDNPASDD
DSSSSGSEGDAKSDSKTSVAMRRTSGSRLTKVASLRIRRQSTVARSTCSSALRDSKLATKHEENNSNSNEGTATEKVCPYSYCSLHGHHHSSDQVPITLK
RFVSMRRRLLKPQKKTNNSSDSRKKKLTNQTKKGVVVSGKSGKAANGGRQKSCVVKANGGLLETDSSDVFQDKDLVVSADYKPDEDGGIVHLSSENSVEK
NKKNMGLWGVIYQHMASGIAAEGDESSQAEKTGMDRNKDPEEGNGGIRNVDSHQLDAIKLVQEAFDKILSEIPDPLSDNQSVDYSDAGSDASSAPTCDDS
QKVRIEQEQMELKKAETFNDSESKEGNKSAAKKKANGWSNLKKIMIMKRFVKAMDNVKNFNLRKRQQDHLRCTEQEGGAEKVNLRPQTIGEKKNSEEWML
DYALQKVISTLAPAQKRKVALIVQAFETVAPQGDTDAAASSTPKTPVHKEPEPEQNIIFGVLLRKVSTVPKESRNNADRGVSDSYLMKSRLKAIDQIADD
NSIDYVTTVPKESRNNADGTDCDVGGSDLMKSRLKSIDQIAEEPIAEDNSVDNVTLAPAAEKEEEPPVEDVQPSIEAENDEELDVDRNAAEPEAEMGGNQ
KHKNLWFLIYKHMVSGTTQPLQESGEEVKGGRNESRSQTEAIRLVEEAIDEIPLPETQDDAPPADDDHPSVATRDINSQVEDKHDERTEIETTVEDQKEE
ARPAAKKSWSNLKKVILLKRFIKSLEKVNDTCLTPSSRKFINPKEPKFLPSVDGQEAEKVLLRRQDVGDRKSGDEWMLDHALRQVVAQLTPVRRRKVELL
VEAFETVSPTIGN
|
DESeq2's median of ratios [FLAX]
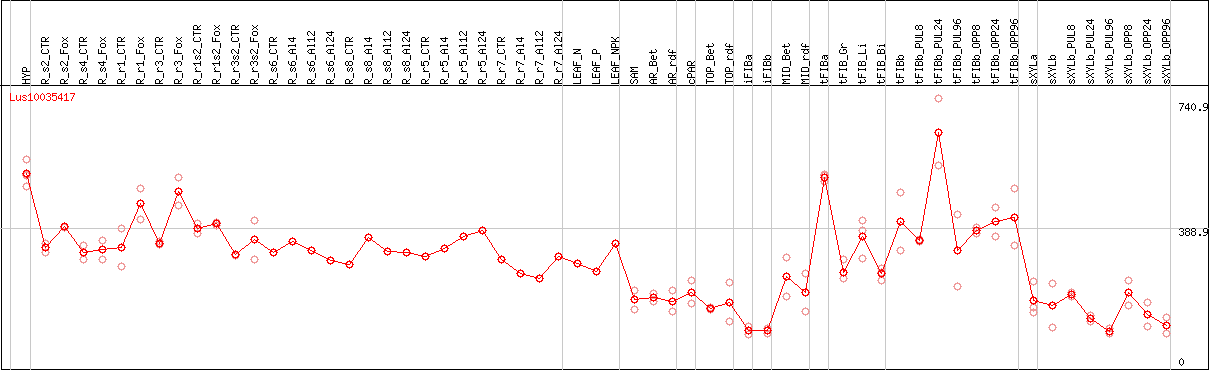
Coexpressed genes
Lus10035417 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
