External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G64530 207 / 1e-68
NAC
ANAC104, XND1
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
AT1G61110 105 / 3e-27
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT1G52890 101 / 6e-26
NAC
ANAC019
NAC domain containing protein 19 (.1)
AT3G15500 100 / 2e-25
NAC
ATNAC3, ANAC055
NAC domain containing protein 55, NAC domain containing protein 3 (.1)
AT4G27410 97 / 2e-24
NAC
RD26, ANAC072
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT3G15510 97 / 5e-24
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
AT1G69490 92 / 8e-23
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT3G04070 93 / 1e-22
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT1G01720 91 / 4e-22
NAC
ATAF1, ANAC002
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT1G77450 87 / 4e-21
NAC
ANAC032
NAC domain containing protein 32 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000206
290 / 3e-101
AT5G64530 264 / 5e-91
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
Lus10035648
290 / 3e-101
AT5G64530 264 / 5e-91
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
Lus10010747
275 / 3e-95
AT5G64530 259 / 4e-89
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
Lus10011215
110 / 4e-29
AT1G61110 305 / 1e-102
NAC domain containing protein 25 (.1)
Lus10018469
109 / 2e-28
AT1G61110 301 / 5e-101
NAC domain containing protein 25 (.1)
Lus10032657
108 / 4e-28
AT3G15510 353 / 1e-120
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10043095
107 / 7e-28
AT3G15510 351 / 1e-119
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10006547
104 / 2e-26
AT3G04070 337 / 4e-114
NAC domain containing protein 47 (.1.2)
Lus10003269
102 / 9e-26
AT3G04070 324 / 3e-109
NAC domain containing protein 47 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G022800
182 / 7e-59
AT5G64530 231 / 5e-78
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
Potri.001G206900
178 / 4e-57
AT5G64530 221 / 2e-74
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
Potri.007G105000
162 / 1e-50
AT5G64530 183 / 3e-59
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
Potri.005G064100
151 / 1e-46
AT5G64530 199 / 3e-65
Arabidopsis NAC domain containing protein 104, xylem NAC domain 1 (.1)
Potri.004G038000
108 / 1e-28
AT1G61110 332 / 3e-113
NAC domain containing protein 25 (.1)
Potri.001G404400
107 / 6e-28
AT3G15510 345 / 1e-117
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Potri.011G123500
104 / 6e-27
AT3G15510 323 / 4e-109
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Potri.011G046700
102 / 3e-26
AT1G61110 331 / 8e-113
NAC domain containing protein 25 (.1)
Potri.008G089000
97 / 1e-24
AT1G69490 331 / 6e-115
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Potri.011G123300
96 / 7e-24
AT4G27410 354 / 2e-122
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10035647 pacid=23147412 polypeptide=Lus10035647 locus=Lus10035647.g ID=Lus10035647.BGIv1.0 annot-version=v1.0
ATGAACCTGCCTCCTGGTTTCCGATTCTACCCTACTGACGAGGAACTTGTAGTCCACTTCCTCCATCGCAAAGCTACCCTTTTACCATGCCACCCTGACG
TCATCCCTGATCTCGACCTCTACCCTTACGATCCTTGGGACCTCCATGGCAAGGCGTTGGAAGAAGGGAATCAGTGGTACTTCTACAGCAGGAAGACTCA
GAACCGGGTGACCGAGAATGGGTTCTGGAAAACCATAGGGATGGAGGAGCCGATTATTTCATCAAGTGGCAACAACAGGAGGGTTGGAGTTAAGAAGTAT
TTGATGTTCTTCTTGGGTGGAGATGTCAAAACTAACTGGATCATGGAGGAGTTTCGTCTCCCCGATCCCTCCTCTGGTACTGGTACTAGTACCAGTAGGT
CATCATCAAGAAGTACACGATCACGGTCCAGACCGGACTACAACAAGTGGGTAGTTTGCAGAGTGTATGAGCGAAACTCGAGTGACGACGAGGAGGATGG
GTCGACGGGGCTATCGTGCTTGGATGAAGTGTTCCTGTCGTTGGATGATCTCGACGACATTAGTTTGCCTTATAATTAA
AA sequence
>Lus10035647 pacid=23147412 polypeptide=Lus10035647 locus=Lus10035647.g ID=Lus10035647.BGIv1.0 annot-version=v1.0
MNLPPGFRFYPTDEELVVHFLHRKATLLPCHPDVIPDLDLYPYDPWDLHGKALEEGNQWYFYSRKTQNRVTENGFWKTIGMEEPIISSSGNNRRVGVKKY
LMFFLGGDVKTNWIMEEFRLPDPSSGTGTSTSRSSSRSTRSRSRPDYNKWVVCRVYERNSSDDEEDGSTGLSCLDEVFLSLDDLDDISLPYN
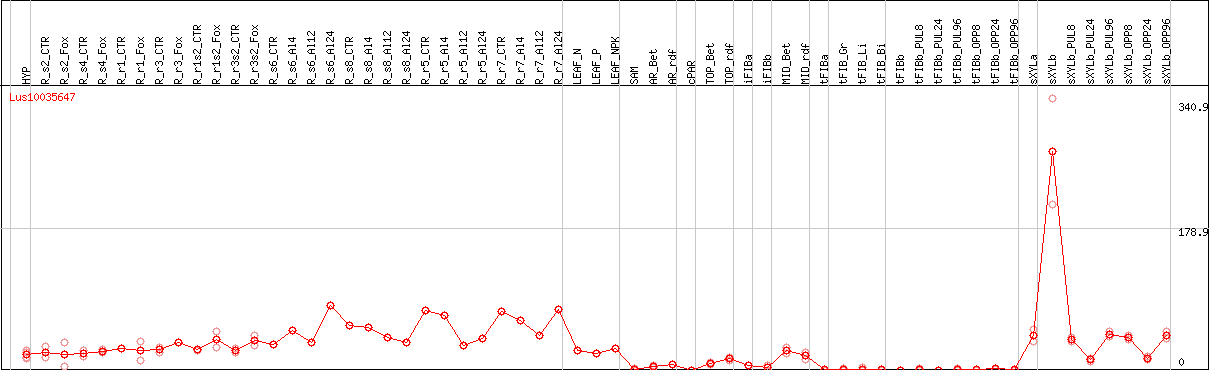
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10035647 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.