External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G19170 207 / 4e-63
CCD4, NCED4
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
AT4G18350 100 / 3e-24
ATNCED2, NCED2
nine-cis-epoxycarotenoid dioxygenase 2 (.1)
AT1G30100 95 / 3e-22
ATNCED5, NCED5
nine-cis-epoxycarotenoid dioxygenase 5 (.1)
AT3G63520 94 / 8e-22
ATNCED1, ATCCD1, CCD1
carotenoid cleavage dioxygenase 1 (.1)
AT3G14440 92 / 3e-21
SIS7, ATNCED3, STO1, NCED3
SALT TOLERANT 1, SUGAR INSENSITIVE 7, nine-cis-epoxycarotenoid dioxygenase 3 (.1)
AT3G24220 92 / 4e-21
ATNCED6, NCED6
nine-cis-epoxycarotenoid dioxygenase 6 (.1)
AT1G78390 87 / 1e-19
ATNCED9, NCED9
nine-cis-epoxycarotenoid dioxygenase 9 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035696
317 / 2e-105
AT4G19170 813 / 0.0
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Lus10029512
100 / 1e-26
AT3G63520 211 / 9e-67
carotenoid cleavage dioxygenase 1 (.1)
Lus10008443
100 / 1e-24
AT4G19170 315 / 1e-102
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Lus10029513
95 / 3e-22
AT3G63520 901 / 0.0
carotenoid cleavage dioxygenase 1 (.1)
Lus10016410
94 / 6e-22
AT3G63520 842 / 0.0
carotenoid cleavage dioxygenase 1 (.1)
Lus10011750
93 / 2e-21
AT3G24220 657 / 0.0
nine-cis-epoxycarotenoid dioxygenase 6 (.1)
Lus10019710
92 / 3e-21
AT3G63520 842 / 0.0
carotenoid cleavage dioxygenase 1 (.1)
Lus10023673
89 / 3e-20
AT3G24220 706 / 0.0
nine-cis-epoxycarotenoid dioxygenase 6 (.1)
Lus10042482
52 / 1e-07
AT1G78390 374 / 4e-125
nine-cis-epoxycarotenoid dioxygenase 9 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G069100
231 / 3e-72
AT4G19170 812 / 0.0
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Potri.019G093400
229 / 2e-71
AT4G19170 803 / 0.0
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Potri.009G152000
142 / 6e-41
AT4G19170 158 / 5e-44
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Potri.009G151900
146 / 1e-40
AT4G19170 548 / 0.0
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Potri.009G152300
145 / 4e-40
AT4G19170 539 / 0.0
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Potri.009G152200
142 / 5e-39
AT4G19170 546 / 0.0
carotenoid cleavage dioxygenase 4, nine-cis-epoxycarotenoid dioxygenase 4 (.1)
Potri.011G084100
95 / 3e-22
AT1G30100 799 / 0.0
nine-cis-epoxycarotenoid dioxygenase 5 (.1)
Potri.011G112400
94 / 9e-22
AT3G14440 868 / 0.0
SALT TOLERANT 1, SUGAR INSENSITIVE 7, nine-cis-epoxycarotenoid dioxygenase 3 (.1)
Potri.001G393800
93 / 1e-21
AT3G14440 883 / 0.0
SALT TOLERANT 1, SUGAR INSENSITIVE 7, nine-cis-epoxycarotenoid dioxygenase 3 (.1)
Potri.001G265600
87 / 2e-19
AT3G63520 878 / 0.0
carotenoid cleavage dioxygenase 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03055
RPE65
Retinal pigment epithelial membrane protein
Representative CDS sequence
>Lus10035700 pacid=23147592 polypeptide=Lus10035700 locus=Lus10035700.g ID=Lus10035700.BGIv1.0 annot-version=v1.0
ATGGATTCCTTCTCTACATCCTATATCTCGAGATCAGTCTCTCCTCCTTCTACATTGCACCTCCGTCCACTCCAAATACACTCTCTTCCAATTACTCAAG
AAAAAACCCACACCGCCGGCAAACCACCACCTTCTCCTCCTACCCAATCTCCACCGCCTAGTAAGACCACCACCAATCGCCGTTCAATTCGGCCTTTTGT
GCCACAACAGCTGTCCATACCCTCACTAATGTTCAATTCCGTCGACGATTTCATCAACACCTTCGTCGACCCACCAAACAAGCCCGCTACCGACCCCCGC
TTCGTCCTCTCCGGCAACTTCTCCCCTGTCGACGAGCTCCCTCCCACCGAGTGCCACGTCATACACTGCTCCTTACCTCCATGCCTCGATGGGGCCTACA
TACGTAACGGCCCAAACCCACAGTACCCGCCCCGCGTTCCCTACCACCACTTTGATGGCGACGGTATGCTCCACGCGCTCACCATCTCAAGAGGGAAAGC
CACGCTCTCTAGCCGTTATGTCAAGACCTACAAGTACATCACCGAACGCGACGCCGGAGCTCCCATCCTCCCTAATCTCTTTTCCGGCTTCAACGGCCTG
ACCGCCTCCGCGGCCCTTGCTGCCCTTTGTGCCGCGAAAATCGCCACCGTAATGTTTAATCCAGCCAACGGCATCGGTTTGGTTAACACAAGCCTGGACT
TAAATCATTACTAA
AA sequence
>Lus10035700 pacid=23147592 polypeptide=Lus10035700 locus=Lus10035700.g ID=Lus10035700.BGIv1.0 annot-version=v1.0
MDSFSTSYISRSVSPPSTLHLRPLQIHSLPITQEKTHTAGKPPPSPPTQSPPPSKTTTNRRSIRPFVPQQLSIPSLMFNSVDDFINTFVDPPNKPATDPR
FVLSGNFSPVDELPPTECHVIHCSLPPCLDGAYIRNGPNPQYPPRVPYHHFDGDGMLHALTISRGKATLSSRYVKTYKYITERDAGAPILPNLFSGFNGL
TASAALAALCAAKIATVMFNPANGIGLVNTSLDLNHY
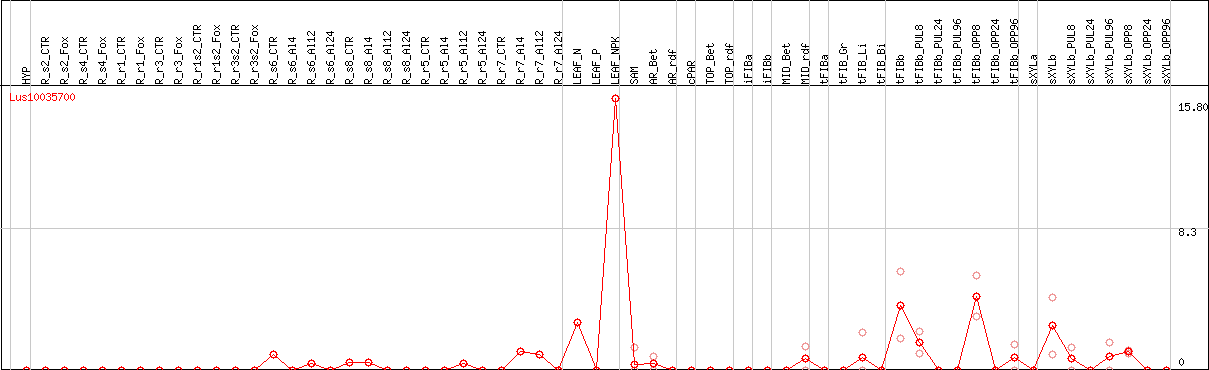
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10035700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.