Lus10035748 [FLAX]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Poplar homologues |
|
|||||||||||||||
| PFAM info | ||||||||||||||||
|
Representative CDS sequence |
>Lus10035748 pacid=23147338 polypeptide=Lus10035748 locus=Lus10035748.g ID=Lus10035748.BGIv1.0 annot-version=v1.0
ATGGTTAAAGACGACGGGAAATTAAGCGCCTCGAAGCGAGCGTACAAGACAGCGCAAGAGGAGGGGAATCGACAAGACGAAGCGCGATGGGCGAACGTGA
TAGGCAACATACTGAAGGACAGGGGAGAGTACGTTGAGGCACTCAGATGGATGCGGAAGGATTATGACGTCTCTGTCAGATACTTACCGGAGAAAGACCT
GTTGGCGACTTGTCAGTCTATCGGGGAGCTCTACCTCCGTCTCGAGAACTTCTCTGATGCATTGGTTTATCAGAAAAGACATCTCGAGCTAGCAAATAGT
GCAAACAACCTTGTCGAGCAGCAAAGAGCAAGCACTCAACTTGGTCGCTCATACCATGCGATGTTCTTGAGGTCCGATGATGATTACTACGCAGTTAAGA
ATGCCAAAAAGTACTTCAAGCTTTCAATGAAGCTAGCTGAGACTCTCAAGGCAAATCCACCGTCTAATAATTCTTCTTTCCTTAAGGAGTACATAGATGC
ACATAACAACATCGGAATGCTTGAATTGGATCTTGATAACTTGGAATTGGCAACTGATATCCTCATGAAAGGACTAAAGATCTGTGACGATGAAGAGGTA
GATGAAAATGATGATGGACGGAGCAGGCTCCATCACAATCTTGGAAATTGTTACATGGAGATACGAAAGTGGGAGAAAGCCCGGGAACATATTGAGAAAG
ATATTCTGATTTGTAAAAAAATTGGTCAGTGCCAAGGGGAGGCAAAGGGCTTCATAAATCTTGGTGAACTACATTATAGGGTCCAAAAGTACGAAGATGC
AGTTCTTTGCTACCAGATGGCTCATAAACTTGCAAAATCCATGGAAGATGAAGATGCATTAGTGAAACAAATTGATCAGAATATCATGACCGTAAGAGAA
GCAGTGAAGGTAATGGATGAGTTGAAAATCGAAGAGCAGAATCTTAAGAAGTTAACCAGAGATATGATTGCTGTAAGAGGCACGGCACGTGAACGGAAGT
CTTTGCTGAAGCTGTATACAAAACTAGATCACCTTATTGAGAAATCCAGCACGATCTCTGCATGGCTGAAGCATCGTGAATTTGCAAAACAGAAAAAGGA
GGTGGCAGCTCAACTATGTGACAAAATCAAGCTGAACGACTCGTATCTGGCTCTAGGAGAGTCATACCGAAAGATTAGAGACTTCAACAAGGCTTACAAA
TGGTTCTCAAAGGCTTGGGAGGGATCTAAGTTGATTGGCTATTTGGAGGGTCAAGCATTAGCTAAAATAAATATCGGTGATGTTCTTGACTGTCGTGGTG
ATTGGTCAGGTGCACTAGCTACTTTTGAAGAAGGCTATAGGATAGCCATGGAAGCTAATGTTCCTTCTGCCCAGATTCAAGCCCTTGAGAATATGCACTA
CAGCCAAATGATGAGATTTCAGAATGCTGAAGAGGCCAGGAGGTTACAACATGAACTTGAAAAATTGAAGCAGTTTGGCGGAAGGGAGGCTGAGAAGAAC
AATTTGGCCGCGGATCACTGCTCTGAGACTGACACAGAAGGTGATTATTTGTCTGATGGTTCATTTTCAAATCCGTGCTCTCCAAACTCTGGCGACTCCT
TGCTTCGGAGATCCACATCTTCCGCATCCGTTGAAGAGTTCGAGGATGACACACCTCTGATTTCACTGCTCAAGTCCACTAAACGTTTGACTAGAAAAAC
TAATACCGATGGCAAAGAGCCTTATGTATCTAATAATCACAATGAAGCTTCTCCAATATGTTCTGCAATCCCAACTACAAGTCAACAATCAGTTGGGCGT
AAGCGTGTTAGGGTCATCCTTTCTGATGATGAAAATGAAACAAATGAGGGTGTGGACTACTTGGAGAGGAAGTTGCCTACCACTCCAGTTGAAGTTGTTG
CTACATCTGACGAGACTAGGAGGAGAAACATTGGAGCATGTTCTTCATCTGAAATTCGGGATTTATCGACGGATGGTTCCAAATATGCCACTGATTCATC
AAATCAAGTTAAAGATGAAGATGTTGCTTTTCTCTACAAGTCTCCAAGTGCTAAAGGGCGAGCGATAACATTCAACATTGCCAACGACCAGATAAGTTTT
GAGGTTGGCTCGAGCTTAGCTTTTAGTGGATTGAGCATTGAATCTTTGAGCGTTGAAGTCGCATGCTTATACTATTTACAGCTTCCTACAGATAAGAGAT
GCCAAGGTTTGTTGCCCATCATTCAGCACTTGAAACATTCTGGACAAATTTTGGAATCATTGGACGACCTTGGAGCTCTCAACGACGCGGGGAACATATT
CATTGAAGTTTCCATAAATGGATGGATCCAGAAGCGGTTGATGAAACTGTACATTGACTGCTGTGGAGAGCTGCATGAAACACCTGATATAAAGTTACTA
AAGAAGCTCTACATTTCGGAGGTGGAGGATGAAGTTATTGCATCCGAATGTGGGCTGCAAGATATATCTATAGCTCCATTATTGAACGCCCTACAAACAC
ATAAAACAGTTCTCATGTTGGATCTTTCTCATAATTTGTTAGGAAATGGAACGGTGGAGAAACTTGAACAAATTTTCACAACTGGCCAAAAACATGGTGA
TCTATCTTTAGACCTGCACTGCAACCGATTTGGTCCAACTGCATTGTTACAGATCTGTGAATGTCCTGTCCTCGTTTCCCGACTTGAGGTGCTAAATATC
TCTGGTAATCGTCTCACTGATGCATGTGGATCATACCTGTCAACTATATTGGACAAATGTAAAGCTCTTTACAGTTTGAATATAGAGCGCTGCTCCATCA
CATCTAGAACGATTCAAAAGGTTGCTGATGCAATACATTCTGGTTCATTCTTGGCACAACTTTCCATAGGATATAATTCTTTATCAGGAACTTCCATCAT
CAATCTATTGATGAAATTATCTACCTTGAATAGCTTTACAGAACTGAATCTAAGTGGTCTGAAGTTGAACAAGCGAGTTATAGATAGCCTCTGTGAACTC
ACAAAAAGCTCTTGTTTGACAAACCTGAGGGTTGGGAGCACTGGCATAGGAAATGAATGGGCTTTGCAGTTGACTGATTCATTGTCCAGCGGATCTCAGG
AATTTCTTAAACTTGACCTATCATACTGCACACTGACATCCACCTTCGTTCAGACACTTACCACGAATTTAAGCCTGACTTGCTGCATTCTTGAGCTTAA
TCTCGAAGGAAATCCTATTCTGCCTGAGGGAGGAAGCGCATTGGTATCACTAGTCAAGCAACCAGGGTGTTCCCTGAGACTTCTCATCCTAAACAAATGT
CAGCTGGGACTTCCCCTCATCCTTCAATTGGTCCAAGCACTCGCAGAGAATGATCAGCTGGAGGAACTCCATGTTGCTGAGAATGGCGATCTTCATAAAT
TAGCCTCAACGACTGATCAACAGACAGCAACAATAGCATCTCATTCCGACCCAGCGGATCACGAGGGAAGATTAGTTGATTGTCAACTAAACATGGAGAG
TGACGGCCTCGAAGTTGCTGATAGTGAGGATTCAACCAGAGTCGAAGCTGCTGCTGCTATTCGATCTGAGCTGAACGACACATGCACAAGCTCCTCCTGG
GGGAAGAAGAAGAAGAAGAAGAATGAAGACGAGCCAATAATATCGTCTCAAATCGTTAAGGAGCTGTCGGATGGCATAAGTGCAGCCAAGGCATTGCAGG
TGTTGGATTTGAGCAGGAATGGACTTTCGGTGAAAGAAGCGGAAGCATTGTATGGTTCGTGGAGAGTGGGGAAGAAGGGTTCCTCAGCTCAGAAGCACGT
TAAAGACCAAATAGTCCATTTCTCAGTTGCAGGAAAAAAGTGTTGCAGCAAGCCGTGCTGCAGAAGGGAATGA
|
|||||||||||||||
|
AA sequence
|
>Lus10035748 pacid=23147338 polypeptide=Lus10035748 locus=Lus10035748.g ID=Lus10035748.BGIv1.0 annot-version=v1.0
MVKDDGKLSASKRAYKTAQEEGNRQDEARWANVIGNILKDRGEYVEALRWMRKDYDVSVRYLPEKDLLATCQSIGELYLRLENFSDALVYQKRHLELANS
ANNLVEQQRASTQLGRSYHAMFLRSDDDYYAVKNAKKYFKLSMKLAETLKANPPSNNSSFLKEYIDAHNNIGMLELDLDNLELATDILMKGLKICDDEEV
DENDDGRSRLHHNLGNCYMEIRKWEKAREHIEKDILICKKIGQCQGEAKGFINLGELHYRVQKYEDAVLCYQMAHKLAKSMEDEDALVKQIDQNIMTVRE
AVKVMDELKIEEQNLKKLTRDMIAVRGTARERKSLLKLYTKLDHLIEKSSTISAWLKHREFAKQKKEVAAQLCDKIKLNDSYLALGESYRKIRDFNKAYK
WFSKAWEGSKLIGYLEGQALAKINIGDVLDCRGDWSGALATFEEGYRIAMEANVPSAQIQALENMHYSQMMRFQNAEEARRLQHELEKLKQFGGREAEKN
NLAADHCSETDTEGDYLSDGSFSNPCSPNSGDSLLRRSTSSASVEEFEDDTPLISLLKSTKRLTRKTNTDGKEPYVSNNHNEASPICSAIPTTSQQSVGR
KRVRVILSDDENETNEGVDYLERKLPTTPVEVVATSDETRRRNIGACSSSEIRDLSTDGSKYATDSSNQVKDEDVAFLYKSPSAKGRAITFNIANDQISF
EVGSSLAFSGLSIESLSVEVACLYYLQLPTDKRCQGLLPIIQHLKHSGQILESLDDLGALNDAGNIFIEVSINGWIQKRLMKLYIDCCGELHETPDIKLL
KKLYISEVEDEVIASECGLQDISIAPLLNALQTHKTVLMLDLSHNLLGNGTVEKLEQIFTTGQKHGDLSLDLHCNRFGPTALLQICECPVLVSRLEVLNI
SGNRLTDACGSYLSTILDKCKALYSLNIERCSITSRTIQKVADAIHSGSFLAQLSIGYNSLSGTSIINLLMKLSTLNSFTELNLSGLKLNKRVIDSLCEL
TKSSCLTNLRVGSTGIGNEWALQLTDSLSSGSQEFLKLDLSYCTLTSTFVQTLTTNLSLTCCILELNLEGNPILPEGGSALVSLVKQPGCSLRLLILNKC
QLGLPLILQLVQALAENDQLEELHVAENGDLHKLASTTDQQTATIASHSDPADHEGRLVDCQLNMESDGLEVADSEDSTRVEAAAAIRSELNDTCTSSSW
GKKKKKKNEDEPIISSQIVKELSDGISAAKALQVLDLSRNGLSVKEAEALYGSWRVGKKGSSAQKHVKDQIVHFSVAGKKCCSKPCCRRE
|
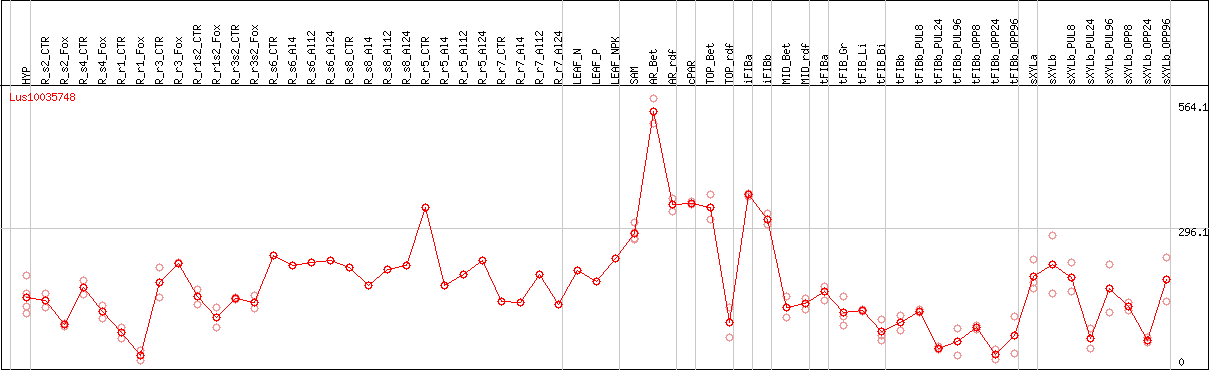
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10035748 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
