External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G24050 229 / 1e-71
eIFiso4G2
eukaryotic translation Initiation Factor isoform 4G2, MIF4G domain-containing protein / MA3 domain-containing protein (.1)
AT5G57870 224 / 2e-69
eIFiso4G1
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
AT4G30680 210 / 4e-69
Initiation factor eIF-4 gamma, MA3 (.1)
AT4G24800 66 / 8e-13
ECIP1
EIN2 C-terminus Interacting Protein 1, MA3 domain-containing protein (.1.2.3)
AT5G63190 64 / 3e-12
MA3 domain-containing protein (.1.2)
AT3G48390 61 / 2e-11
MA3 domain-containing protein (.1)
AT1G22730 53 / 2e-08
MA3 domain-containing protein (.1)
AT3G60240 48 / 1e-06
CUM2, EIF4G
CUCUMOVIRUS MULTIPLICATION 2, eukaryotic translation initiation factor 4G (.2.3.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036594
340 / 2e-114
AT5G57870 625 / 0.0
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
Lus10010782
289 / 3e-94
AT5G57870 952 / 0.0
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
Lus10022168
286 / 1e-92
AT5G57870 976 / 0.0
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
Lus10028156
59 / 2e-10
AT3G60240 1275 / 0.0
CUCUMOVIRUS MULTIPLICATION 2, eukaryotic translation initiation factor 4G (.2.3.4)
Lus10042856
57 / 8e-10
AT3G60240 970 / 0.0
CUCUMOVIRUS MULTIPLICATION 2, eukaryotic translation initiation factor 4G (.2.3.4)
Lus10012205
54 / 1e-08
AT1G22730 560 / 0.0
MA3 domain-containing protein (.1)
Lus10011324
53 / 2e-08
AT1G22730 816 / 0.0
MA3 domain-containing protein (.1)
Lus10018085
52 / 4e-08
AT4G24800 995 / 0.0
EIN2 C-terminus Interacting Protein 1, MA3 domain-containing protein (.1.2.3)
Lus10042080
49 / 4e-07
AT4G24800 974 / 0.0
EIN2 C-terminus Interacting Protein 1, MA3 domain-containing protein (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G104100
261 / 3e-83
AT5G57870 882 / 0.0
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
Potri.006G182100
255 / 5e-81
AT5G57870 884 / 0.0
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
Potri.006G265300
224 / 8e-70
AT5G57870 790 / 0.0
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
Potri.018G017600
221 / 2e-68
AT5G57870 809 / 0.0
eukaryotic translation Initiation Factor isoform 4G1, MIF4G domain-containing protein / MA3 domain-containing protein (.1.2)
Potri.012G091900
59 / 2e-10
AT5G63190 1022 / 0.0
MA3 domain-containing protein (.1.2)
Potri.015G088300
57 / 5e-10
AT4G24800 1008 / 0.0
EIN2 C-terminus Interacting Protein 1, MA3 domain-containing protein (.1.2.3)
Potri.014G052600
57 / 7e-10
AT3G60240 998 / 0.0
CUCUMOVIRUS MULTIPLICATION 2, eukaryotic translation initiation factor 4G (.2.3.4)
Potri.005G199200
54 / 8e-09
AT1G22730 828 / 0.0
MA3 domain-containing protein (.1)
Potri.002G140200
52 / 4e-08
AT3G60240 942 / 0.0
CUCUMOVIRUS MULTIPLICATION 2, eukaryotic translation initiation factor 4G (.2.3.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF02847
MA3
MA3 domain
Representative CDS sequence
>Lus10035812 pacid=23147417 polypeptide=Lus10035812 locus=Lus10035812.g ID=Lus10035812.BGIv1.0 annot-version=v1.0
ATGAATGCCGATGAACTCAAGAGGAAAACGGTCTCAATCTTGAAGGAGTATTTCAGCATTCGACTTGTGGACGAAGCATTCCAGTGCATTGAAGAACTCA
AGTCTCCAGCATACCACCCGGAAGTCATCAAGGAAGGCATTGCCCTTGGCCTGGAGGAAAACCCTCCTTGCGTGGAACCTGTCTTCAAGCTCCTCGAATA
CTTGTCAACCAAGAAAACTCTCACAGCTCGAGACATAACGACCGGGTGTATGCTCTACGCAACGATGCTGGATGACATTGGAATCGACTTGCCCAGAGCG
CCCAACAACTTTGGCGAGATCATCGGGAAACTAATCCTCTCTGGTCTACTGGAGTTCAAATCGGTGACGGAGATCCTAAAGAAGATGGAGGACGACATGT
ACCAGAAATCCGTGCTGGATGCAGCAGTCAGGATTGTGAAATCCAGCTCCGTAGGCCAAGGTATACTTGAATCCCAAGCTGCTGATATCGAGGCTTGCCA
GAGCTTGTTCCAGTAA
AA sequence
>Lus10035812 pacid=23147417 polypeptide=Lus10035812 locus=Lus10035812.g ID=Lus10035812.BGIv1.0 annot-version=v1.0
MNADELKRKTVSILKEYFSIRLVDEAFQCIEELKSPAYHPEVIKEGIALGLEENPPCVEPVFKLLEYLSTKKTLTARDITTGCMLYATMLDDIGIDLPRA
PNNFGEIIGKLILSGLLEFKSVTEILKKMEDDMYQKSVLDAAVRIVKSSSVGQGILESQAADIEACQSLFQ
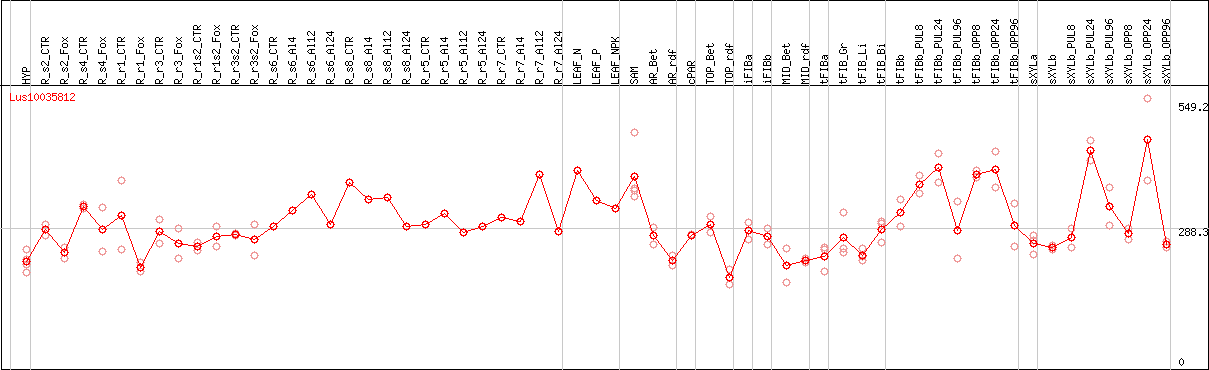
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10035812 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.