Lus10035998 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10035998 pacid=23141109 polypeptide=Lus10035998 locus=Lus10035998.g ID=Lus10035998.BGIv1.0 annot-version=v1.0
ATGGGAATACCTGATACTTCTCTACTGGAGAAGGTTAAGTCTTGGATTGCTTGGGGAGGAAGCGATTTATCACCATCATCTTCAGTGTCCACTGAATTTG
AGATGAACAATAGCGAGTGCAATACAAGTTCTAATGAGCATTCTAACGGATATCATTGTGATAGCTGTGGCCGATGGGTGTGTCTGAAATGCATCAAAGA
ACTGGTGTCTCATCATGTTGTTGAGTCTACTGGCGTTGAACAAAATGTCCATTCCGAGGAACCTATCAAATTATGCATGTTTTGCGGCCTGCCTATAAAG
ATCAGTGAAAAGGTGCATCCCGCAGACTCTCCCCGTGAAAGCCCTGAGCCACCTTCGCCATCTTTTAGCGGCGAATCTATCCAAGGTGATCGTCTTGCCC
GCTATCTTGAATCACGGGACTGTGGATATTCTCCCCTTTCAGTAACCTGCAACAATATGAATGCCTTTGGCCCTCTTCCATGCCTTGCTTCTGTTCGCTG
TTCTCGTAGCAGGGAAGGAGATGAAGAGCAGGCGGAGGATGCCGAGAAGCACTTTCACAGCCCATCTAATGAATATATATCTGATGTAGATTCAAGTAGT
ATTAGCTCTAGGCTTGAGTTTTGCAGTTTTAAGTCGGTGGGTTCAAGTCCTTTTGATAGCCCATCTTCTGGAACTGATTTTACTTCTTATAATAGAATGG
GGCATGCTGTACAGCAGGGGCAAGAAGATAGCCCCTTGTCCCTGAGAGATGGTCCGTTGGATCAAGAAAATATGGCAGTTTTGAGACGGCCTGAAATACG
AGCAGCAGACCCAGATAATGCAGATGATTACTCTGATGATCTCTCAGCTTCAAGAAATCATTATGAAAAGAAACCTTGGGACTTTGAAAGTAACGGACTT
ATCTGGTCCCCCCCTCTTCCCGTGGATGAAAATGATGATGCTGAAAGTAATTTCTTTGCTTATGACGATGACGATGACTATTTTACTGGGGATAGTGTAA
CCTTCTCAACAAGTAGCAGCCTTTCACGTACATTTTCAGCCAAGGAAAGACAGAGTGATGGTAACAAAGAGCCTCTGAAATTCGCAATACAAGAACACTT
TAAGGCTCTGGTATCCCAACTTCTGCAGGGGGAGGGAATCCAAACCAGCGGGGTTGATAGCAGTGAAGATTGGTTGGATATAGTTACTACGTTAGCATGG
CAAGCTGCAAATTTCGTGAAACCTGATACTAGAAGAGGAGGCAGTATGGACCCTGTTGATTATGTAAAGGTGAAGTGTATAGCATCAGGAATTCCAATTG
ATAGTGCCCTAATAAAGGGAGTTGTTTGTTCGAAAAACATAAAGCATAAGCGCATGACAACACAATACAAAAACCCCAGATTACTACTTCTTGGGGGAGC
TCTGGAATATCGAAGTGTTACTAATCAACTGGCATCCTTTAATACATTGGTGCAAGAGGAAAATGATCATCTGAAGATGATTATTTCCAAGATAGAGGCT
CTGCGCCCCAATGTTCTGCTAGTGGAGAAAAATGTATCTCCATATGCCCAAGAATATCTGTTGACAAAGGAAATATCTTTGGTTCCGAATGTGAAGAAAC
CATTGATGGAATGCATAGCACGGTGCACTGGCGCTCTTATTAGTCCATCAATTGATACAATCTCAACTACACGGTTGGGTAACTGTGAACTCTTCCGCGT
GGAGAGATTTTTGGAGGAACATGAGCCTGCTAATCAGTTCAACAAGAAGCCAACCAAGACTTTGATGTTCTTCGAAGGATGTCCAAGACGTTTGGGCTGC
ACGGTCCTTCTGAGGGGCAAGTGCTGCGATGAACTCAAGAAAATTAAGCATGTTGTTCAATATGCTGTTTTTGCCGCATATCACTTATCCCTTGAGACTT
CGTTCCTTGCTGATGAAGGTGCAAGTCTTCCTAAGATGAAATCTGAACAAACGGTTACCGTGCCAGCAGACGAAGGGATGTCAGTCGTTCCTTCAAACAA
GTTTGATGCAATTGCTGACTATTGTTCCGATGATAATGATCATACAGGATTGGAGTTGGTGTCTGAAAATATTGATGCTTATGGCACTACTCAGTCTCTT
GTTCCTGTCAATAATAGCATTGGAGATACACTCCCTGATTTATGCCATATTCCTCTGTCGAACTCAGTCTTACAACACTTTGCTACTTGTGGTTGTAATA
TGCCGATGACGTCTAGAGTTCCTGTCCTGGATGTTAATAGTCTTTCTCATCCTGAGTTGTTAGATATCATTGCCGATCAACCGAGCCAAGTAGATGAGAC
CTTAGAGTTGGAGAAATCTGACCGGACTGATGGACATGGTGTATCTAATGATTATTTATTGGCATCTGATACTTATCAGAGCATATTGGTCTCATTTTCA
AGCCGCTGTGTACCAAAAGGAACTGTATGTGAACGTTCTCGACTCCTGCGTATAAAATTTTATGGATCATTTGATAAACCACTTGGAAGATATCTTCGTG
ATGACCTCTTTGATCAGACATCTTTTTGTAGGAGTTGTAAGGAGCCAGCTGAATCGCATGTATTATGTTATACTCACCAGCAAGGGAATCTTACAATTAA
CGTGAGGTCCCTTCCCTCTATTAAGCTTCCTGGCGAACAAGATGGTAAAATATGGATGTGGCACAGATGCTTGAGGTGTGCTCATGTAGATGGAGTCCCA
CCGTCAACTCGTAGGGTAATTATGTCAGATGCTGCTTGGGGACTTTCATTTGGGAAGTTCTTGGAATTGAGCTTTTCAAACCATGCAACTGCAGATCGTG
TTGCTCCATGTGGTCATTCATTACAGAAAGATTGTCTTCGCTTCTATGGGTTTGGGAGCATGGTTGGATTCTTTCGTTATTCTCCTATTGATATTCTTAA
TGTACATCTGCCACCTTCGGTGCTGGAATTTACGGGCATTTGTCAGCAAGAGTGGATCAAAAAAGAGGCAGCCGAGCTTCTTAGCAAGGTTGAAATCTTT
TACTCGGAGATTTCTAATGTGCTTGATAGCTTGGAAGGGAGAAGTAAGTCATATGGTTGTCAATTGTCCGAGGCAACTGAGCTGCAGAATCATATTCTTG
ATTTGAAAGTTCAACTTGTAAAGGAGAGGAGCAACTATTCTGAAACTCTGAAACTTGCTGTCATGGAGAGTCTACCACCAGGCCAGGAAGGTTTAGACAT
TCTGGAGCTCAATCGTACAAGGCGTTCCCTCATAATTGACTCGCGTGTCTGGGACCGTCAAATATATTCGCTAGATTCTCTCCTTAAGACAATTGCGACT
ATTCGTCCTAAACAAAAGGTTGTGTGTCAAAATATCACGATGGCAGGAAGTGGCGATTCTTTGGGATGTCCAGAACCACTTTGTTCTGGTAATGATCACC
CGTTTGAGCAGGGAGTGATCAGTCACTTAGCTGAGCAATCTCTTTCTGAAGATTTAGTATTATCTTCTCGTCATGTTCATAGACAGGAGGATGTGCATTC
GGATGGTGAGCCTAGCATAACCAAGACTATGACTGACCGCATTGCTTGTGATGCATCCAATTTGTCTGACAGAATCGATTCTGCATGGAGTGGTACTAAT
CAATGTGCTATGAAAGTTCAGGAACTGGACTCATCTGACAATAGTGGGCTGAGGGTTGCTACTAATTCTCCTCTGAAAAGAATGATGGGACCAGTAAGAG
TTAATTCTTTTGACTCTGCATTGAGAGTCCAAGAACGAATACAGAAAGGAACGCCGTTCTCGAATCTCCGATCATTCCATGCTTCTGGAGATTACAGGAC
TATGTTGAGAGATCCCGTATCAAACACTTCGAGGACTTACTCTCAATCTCAAGCATCACCTTTGGAGACACAGAGGTCAAATGTATTATCTGCGACACCT
TCGTTTATATCTTCTGCATCTGACATGACTGGGGGAGCTCGATTTCTGCTTCCTCACAGGACTCACAGTGACGTAGTGATTGGTGTTTATGATGATGATC
CTGCCAGCATAGTATCTTATGCTCTCAATTTGAAAGAACACGAGGATTGGGTAGCAGCTAACAAGTCAAATGAGATTGGAGGCAAATGCAATATAGGTGT
TCTGGGGAGGGAAGATTCTGCTGCTTCCATGATATCAGGATGGCAATCTTTTGGTTCCTTGGACTTCGACTATATGAATTACGGATCTGAAGGAATATCA
TCTTCCATCAGTAACTTGTTCACAGATTCCAAGAAATCCCCACATTTTGGAGTATCTTATAATGATGCTTCTTCTGGTGTTGGAGGTAAAGTGAAGTACT
CAGTGACCTGCTATTTTGCCAACCAGTTTCACTTGTTGAGGAAGAAATGCTGCCCGAGTGATGTTGATTTTGTTCGGTCCTTAAGCCGCTGCCAGAAATG
GAGTGCACAGGGTGGGAAAAGCAACGTCTACTTTGCTAAATCATTGGATGAGCGTTTCATTTTAAAACAAGTTAAGAAGACAGAGTTGGAATCCTTTGAG
GAATTTGCACCTGAATATTTCAAATATGTAACAGAGTCTTTGAACTCTGGAAGTCCAACTTGCCTTGCAAAAATTCTTGGAATATATCAGGTTGCTGTAA
AGCACTTAAAAAGTGGCAGAGAAACAAAGATGGATTTAATGGTAATGGAGAACCTGTTTTTTAGGAGAAAAATATCTAGGGTATATGACCTTAAGGGCTC
TACACGGTCTCGTTACAATCCTGATAGTACAGGAGTGTTGCTGGATATGAATTTAGTGGAGGAATTGCGAACAGAACCCATATTTCTGGGGAGCAAAGCA
AAAAGAAGCCTAGAGAGAGCAATATGGAATGACACATCTTTTCTAGCGTCAGTAGATGTCATGGACTACTCGTTGCTGGTAGGAGTGGACGAGGAGAAGA
AGGAATTGGTTTTAGGGATAATAGACTTCATGAGACAATATACTTGGGACAAGCACTTGGAGACATGGGTAAAAGCATCCGGAATACTTGGGGGTCCGAA
AAATGCTTCTCCGACGATCATTTCTCCGGTTCAGTACAAAAAGAGATTTCGAAAGGCAATGACTTCCTACTTTCTCACGGTTCCTGATCAGTGGTCTTCA
TGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10035998 pacid=23141109 polypeptide=Lus10035998 locus=Lus10035998.g ID=Lus10035998.BGIv1.0 annot-version=v1.0
MGIPDTSLLEKVKSWIAWGGSDLSPSSSVSTEFEMNNSECNTSSNEHSNGYHCDSCGRWVCLKCIKELVSHHVVESTGVEQNVHSEEPIKLCMFCGLPIK
ISEKVHPADSPRESPEPPSPSFSGESIQGDRLARYLESRDCGYSPLSVTCNNMNAFGPLPCLASVRCSRSREGDEEQAEDAEKHFHSPSNEYISDVDSSS
ISSRLEFCSFKSVGSSPFDSPSSGTDFTSYNRMGHAVQQGQEDSPLSLRDGPLDQENMAVLRRPEIRAADPDNADDYSDDLSASRNHYEKKPWDFESNGL
IWSPPLPVDENDDAESNFFAYDDDDDYFTGDSVTFSTSSSLSRTFSAKERQSDGNKEPLKFAIQEHFKALVSQLLQGEGIQTSGVDSSEDWLDIVTTLAW
QAANFVKPDTRRGGSMDPVDYVKVKCIASGIPIDSALIKGVVCSKNIKHKRMTTQYKNPRLLLLGGALEYRSVTNQLASFNTLVQEENDHLKMIISKIEA
LRPNVLLVEKNVSPYAQEYLLTKEISLVPNVKKPLMECIARCTGALISPSIDTISTTRLGNCELFRVERFLEEHEPANQFNKKPTKTLMFFEGCPRRLGC
TVLLRGKCCDELKKIKHVVQYAVFAAYHLSLETSFLADEGASLPKMKSEQTVTVPADEGMSVVPSNKFDAIADYCSDDNDHTGLELVSENIDAYGTTQSL
VPVNNSIGDTLPDLCHIPLSNSVLQHFATCGCNMPMTSRVPVLDVNSLSHPELLDIIADQPSQVDETLELEKSDRTDGHGVSNDYLLASDTYQSILVSFS
SRCVPKGTVCERSRLLRIKFYGSFDKPLGRYLRDDLFDQTSFCRSCKEPAESHVLCYTHQQGNLTINVRSLPSIKLPGEQDGKIWMWHRCLRCAHVDGVP
PSTRRVIMSDAAWGLSFGKFLELSFSNHATADRVAPCGHSLQKDCLRFYGFGSMVGFFRYSPIDILNVHLPPSVLEFTGICQQEWIKKEAAELLSKVEIF
YSEISNVLDSLEGRSKSYGCQLSEATELQNHILDLKVQLVKERSNYSETLKLAVMESLPPGQEGLDILELNRTRRSLIIDSRVWDRQIYSLDSLLKTIAT
IRPKQKVVCQNITMAGSGDSLGCPEPLCSGNDHPFEQGVISHLAEQSLSEDLVLSSRHVHRQEDVHSDGEPSITKTMTDRIACDASNLSDRIDSAWSGTN
QCAMKVQELDSSDNSGLRVATNSPLKRMMGPVRVNSFDSALRVQERIQKGTPFSNLRSFHASGDYRTMLRDPVSNTSRTYSQSQASPLETQRSNVLSATP
SFISSASDMTGGARFLLPHRTHSDVVIGVYDDDPASIVSYALNLKEHEDWVAANKSNEIGGKCNIGVLGREDSAASMISGWQSFGSLDFDYMNYGSEGIS
SSISNLFTDSKKSPHFGVSYNDASSGVGGKVKYSVTCYFANQFHLLRKKCCPSDVDFVRSLSRCQKWSAQGGKSNVYFAKSLDERFILKQVKKTELESFE
EFAPEYFKYVTESLNSGSPTCLAKILGIYQVAVKHLKSGRETKMDLMVMENLFFRRKISRVYDLKGSTRSRYNPDSTGVLLDMNLVEELRTEPIFLGSKA
KRSLERAIWNDTSFLASVDVMDYSLLVGVDEEKKELVLGIIDFMRQYTWDKHLETWVKASGILGGPKNASPTIISPVQYKKRFRKAMTSYFLTVPDQWSS
|
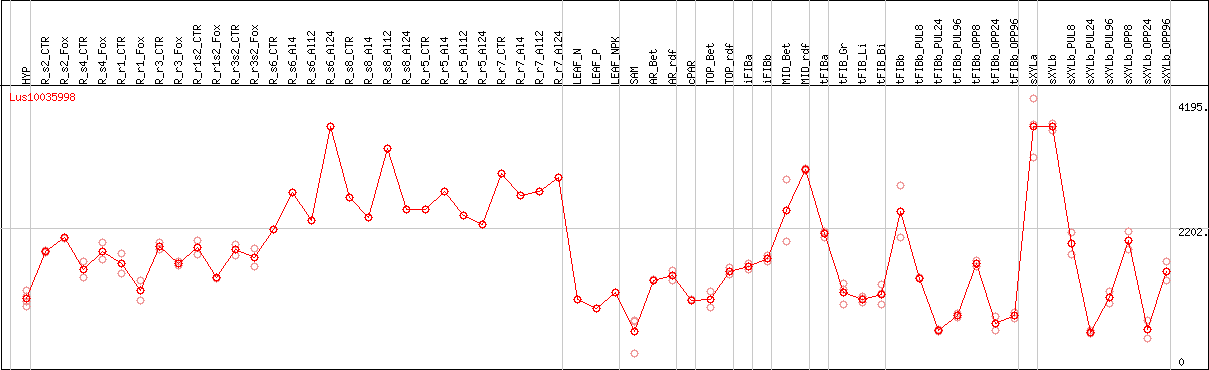
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10035998 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
