External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G70520 61 / 6e-11
ASG6, CRK2
ALTERED SEED GERMINATION 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 2 (.1)
AT5G40380 61 / 9e-11
CRK42
cysteine-rich RLK (RECEPTOR-like protein kinase) 42 (.1)
AT5G48540 49 / 7e-07
receptor-like protein kinase-related family protein (.1)
AT1G19090 48 / 2e-06
CRK1, RKF2
CYSTEINE-RICH RLK \(RECEPTOR-LIKE PROTEIN KINASE\) 1, receptor-like serine/threonine kinase 2 (.1)
AT1G70530 46 / 7e-06
CRK3
cysteine-rich RLK (RECEPTOR-like protein kinase) 3 (.1)
AT4G28670 45 / 2e-05
Protein kinase family protein with domain of unknown function (DUF26) (.1)
AT5G37660 44 / 3e-05
PDLP7
plasmodesmata-located protein 7 (.1.2)
AT4G23230 42 / 0.0001
CRK15
cysteine-rich RLK (RECEPTOR-like protein kinase) 15 (.1)
AT3G60720 42 / 0.0002
PDLP8
plasmodesmata-located protein 8 (.1)
AT4G23160 42 / 0.0003
CRK8
cysteine-rich RLK (RECEPTOR-like protein kinase) 8 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009703
353 / 3e-119
AT1G70520 319 / 4e-100
ALTERED SEED GERMINATION 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 2 (.1)
Lus10010960
123 / 2e-32
AT1G70520 373 / 2e-121
ALTERED SEED GERMINATION 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 2 (.1)
Lus10029123
63 / 3e-11
AT1G70520 822 / 0.0
ALTERED SEED GERMINATION 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 2 (.1)
Lus10013039
61 / 1e-10
AT1G70520 699 / 0.0
ALTERED SEED GERMINATION 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 2 (.1)
Lus10030589
61 / 1e-10
AT1G70520 813 / 0.0
ALTERED SEED GERMINATION 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 2 (.1)
Lus10013040
59 / 7e-10
AT1G70530 788 / 0.0
cysteine-rich RLK (RECEPTOR-like protein kinase) 3 (.1)
Lus10029122
57 / 3e-09
AT1G70530 789 / 0.0
cysteine-rich RLK (RECEPTOR-like protein kinase) 3 (.1)
Lus10030901
53 / 5e-08
AT1G70520 803 / 0.0
ALTERED SEED GERMINATION 6, cysteine-rich RLK (RECEPTOR-like protein kinase) 2 (.1)
Lus10020814
52 / 6e-08
AT5G37660 311 / 2e-106
plasmodesmata-located protein 7 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF01657
Stress-antifung
Salt stress response/antifungal
Representative CDS sequence
>Lus10036028 pacid=23141010 polypeptide=Lus10036028 locus=Lus10036028.g ID=Lus10036028.BGIv1.0 annot-version=v1.0
ATGAACGACGAAGTAACAACATGGCCCACAACCAGAAGCATCCTGGTTTTCTACATATTTCTGAATTGGGGATTACTGATCAAGACAGCAGCTTCTGACC
CAGAAATCAATCTGCTGAGTGTCGGATGTAGCCAATATAAAGATGGCGACTTGCATATTTCTAGAGGCAACCTGAAATCAAGCTTCTCTCAGCTCAGAGC
TCAGTTGAATGGTAGTAGCAGCAAGTATTTCGCAACAGCACAGCAAACTTCTGGTCAGGAACCCGTTTACACGATGGGTCAGTGCAGGAATTACATGTCC
AAACCCGATTGCCTCGCTTGCTTTGACGACGCTGTTGCCAGAATCCTCAATTTCTATTCTGCCAATGGTGCCCGTGTGATCTTGGATGGTTGCTTCCTCA
GGTATGAGAGGAACACTTTCTATGACCAGAGCACTCAGGATGGCCACAGTGGGGTTTCTGGGAATCAGACCATATCCACGCAGGATAACAATTATGGCAC
TGCTCGGGAAGATCTTCTGTTGAATCTTGTTATTGCCACACCCAAACTGAATGGCTTTTATGCAGCAAGCAAAAAGGAAGTGGTAGCTGCTGCTGCTGGT
AATCTTTTTTATGCTGTTGAAACTGTCGGCCAGATTGGTTGCAGGATTGCTTAA
AA sequence
>Lus10036028 pacid=23141010 polypeptide=Lus10036028 locus=Lus10036028.g ID=Lus10036028.BGIv1.0 annot-version=v1.0
MNDEVTTWPTTRSILVFYIFLNWGLLIKTAASDPEINLLSVGCSQYKDGDLHISRGNLKSSFSQLRAQLNGSSSKYFATAQQTSGQEPVYTMGQCRNYMS
KPDCLACFDDAVARILNFYSANGARVILDGCFLRYERNTFYDQSTQDGHSGVSGNQTISTQDNNYGTAREDLLLNLVIATPKLNGFYAASKKEVVAAAAG
NLFYAVETVGQIGCRIA
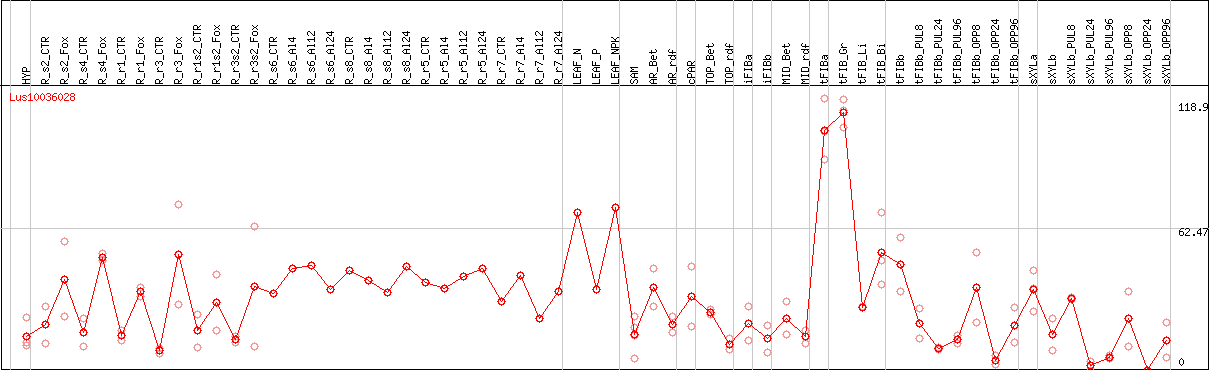
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036028 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.