External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G46264 217 / 2e-68
HSF
SCZ, AT-HSFB4
SCHIZORIZA, heat shock transcription factor B4 (.1)
AT4G36990 153 / 2e-44
HSF
AT-HSFB1, ATHSF4, HSF4, HsfB1
ARABIDOPSIS THALIANA HEAT SHOCK FACTOR 4, ARABIDOPSIS THALIANA CLASS B HEAT SHOCK FACTOR B1, heat shock factor 4 (.1)
AT5G62020 147 / 5e-42
HSF
AT-HSFB2A
ARABIDOPSIS THALIANA HEAT SHOCK TRANSCRIPTION FACTOR B2A, heat shock transcription factor B2A (.1)
AT4G11660 147 / 3e-41
HSF
AT-HSFB2B, HSFB2B
HEAT SHOCK TRANSCRIPTION FACTOR B2B, winged-helix DNA-binding transcription factor family protein (.1)
AT2G41690 136 / 3e-38
HSF
AT-HSFB3, HSFB3
HEAT SHOCK TRANSCRIPTION FACTOR B3, heat shock transcription factor B3 (.1)
AT4G17750 139 / 2e-37
HSF
ATHSFA1A, ATHSF1, HSF1
ARABIDOPSIS THALIANA CLASS A HEAT SHOCK FACTOR 1A, ARABIDOPSIS THALIANA HEAT SHOCK FACTOR 1, heat shock factor 1 (.1)
AT1G32330 138 / 3e-37
HSF
ATHSFA1D
heat shock transcription factor A1D (.1)
AT3G02990 138 / 4e-37
HSF
ATHSFA1E
heat shock transcription factor A1E (.1)
AT5G16820 137 / 8e-37
HSF
ATHSF3, ATHSFA1B, HSF3
ARABIDOPSIS THALIANA CLASS A HEAT SHOCK FACTOR 1B, ARABIDOPSIS HEAT SHOCK FACTOR 3, heat shock factor 3 (.1.2)
AT5G03720 132 / 1e-35
HSF
AT-HSFA3
ARABIDOPSIS THALIANA HEAT SHOCK TRANSCRIPTION FACTOR A3, heat shock transcription factor A3 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026819
221 / 2e-72
AT1G46264 167 / 2e-51
SCHIZORIZA, heat shock transcription factor B4 (.1)
Lus10001133
201 / 3e-61
AT1G46264 319 / 4e-107
SCHIZORIZA, heat shock transcription factor B4 (.1)
Lus10042646
195 / 3e-59
AT1G46264 308 / 8e-103
SCHIZORIZA, heat shock transcription factor B4 (.1)
Lus10008007
161 / 1e-46
AT4G11660 306 / 3e-102
HEAT SHOCK TRANSCRIPTION FACTOR B2B, winged-helix DNA-binding transcription factor family protein (.1)
Lus10024508
161 / 4e-46
AT4G11660 308 / 3e-102
HEAT SHOCK TRANSCRIPTION FACTOR B2B, winged-helix DNA-binding transcription factor family protein (.1)
Lus10019348
149 / 5e-43
AT4G36990 254 / 3e-84
ARABIDOPSIS THALIANA HEAT SHOCK FACTOR 4, ARABIDOPSIS THALIANA CLASS B HEAT SHOCK FACTOR B1, heat shock factor 4 (.1)
Lus10014994
145 / 4e-41
AT5G62020 259 / 2e-85
ARABIDOPSIS THALIANA HEAT SHOCK TRANSCRIPTION FACTOR B2A, heat shock transcription factor B2A (.1)
Lus10009351
142 / 5e-40
AT4G36990 248 / 1e-81
ARABIDOPSIS THALIANA HEAT SHOCK FACTOR 4, ARABIDOPSIS THALIANA CLASS B HEAT SHOCK FACTOR B1, heat shock factor 4 (.1)
Lus10011065
145 / 7e-40
AT1G32330 441 / 5e-152
heat shock transcription factor A1D (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G273700
223 / 7e-72
AT1G46264 251 / 2e-82
SCHIZORIZA, heat shock transcription factor B4 (.1)
Potri.009G068000
216 / 5e-69
AT1G46264 258 / 4e-85
SCHIZORIZA, heat shock transcription factor B4 (.1)
Potri.002G124800
202 / 2e-62
AT1G46264 353 / 7e-121
SCHIZORIZA, heat shock transcription factor B4 (.1)
Potri.014G027100
198 / 9e-61
AT1G46264 335 / 9e-114
SCHIZORIZA, heat shock transcription factor B4 (.1)
Potri.001G108100
162 / 2e-47
AT4G11660 285 / 2e-94
HEAT SHOCK TRANSCRIPTION FACTOR B2B, winged-helix DNA-binding transcription factor family protein (.1)
Potri.007G043800
154 / 5e-45
AT4G36990 241 / 3e-79
ARABIDOPSIS THALIANA HEAT SHOCK FACTOR 4, ARABIDOPSIS THALIANA CLASS B HEAT SHOCK FACTOR B1, heat shock factor 4 (.1)
Potri.012G138900
147 / 5e-42
AT5G62020 230 / 2e-74
ARABIDOPSIS THALIANA HEAT SHOCK TRANSCRIPTION FACTOR B2A, heat shock transcription factor B2A (.1)
Potri.016G056500
144 / 9e-42
AT2G41690 218 / 1e-71
HEAT SHOCK TRANSCRIPTION FACTOR B3, heat shock transcription factor B3 (.1)
Potri.017G059600
145 / 1e-39
AT4G13980 499 / 1e-174
HEAT SHOCK TRANSCRIPTION FACTOR A5, winged-helix DNA-binding transcription factor family protein (.1)
Potri.015G141100
140 / 2e-39
AT5G62020 233 / 8e-76
ARABIDOPSIS THALIANA HEAT SHOCK TRANSCRIPTION FACTOR B2A, heat shock transcription factor B2A (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0123
HTH
PF00447
HSF_DNA-bind
HSF-type DNA-binding
Representative CDS sequence
>Lus10036062 pacid=23141084 polypeptide=Lus10036062 locus=Lus10036062.g ID=Lus10036062.BGIv1.0 annot-version=v1.0
ATGCAGAGAAGTTGTAATGAAGAGGAAGAAGAAGAAGAAGAAGAAGAGGAGGAGAGTATAATAATGCAGGGATTGAACAAGGGCGGTGGTGGAGTGGTGG
CGCCATTCTTAACCAAGACATACGAGCTGGTGGATGATCCAATGACAGACCACATTGTGTCTTGGGGAAGTAATATGAATGATCCAATGAGCACATTCGT
GGTATGGAGACCTCCGGAGTTCGCAAGAGATCTCCTCCCTAACTACTTCAAACACAACAACTTCTCCAGCTTCGTCCGCCAACTCAACACCTACGGATTC
AAGAAGGTAGTAGCAGAAAGATGGGAGTTTGGTAACGACTACTTCAGGAAAGGAGCAAAGCAGTTGCTCTCCCATATCCACCGTAGAAAATCACCCCATC
ATCACCATCACCATCATCATAACGATCATCATCATCATTATCATAATCATGACTTCATATTTCAGCTTCCACCACCTAATTACATTAACCCTCATCACCA
ACCCCATCACATATTCCTCCCACCAGCAGCAGCAGCAGTACATTCGGATGATGATGATGAGATGATGATGTTGAGTACACTTTCGGAGGACAACCAGAGG
CTGAAGAAGAGGAACTTGGCTCTGCTCTCTGAGCTCTCCCACATGAAGTCCCTCTACAACGACATCATCTACTTCATCCAAAACCATCTCAAGCCCAACC
ACCCATTTCCTCATCATCATCATCATCCCCCTACTACTACTACTGCTACTAATAATAATAATAATAATAACAAGCTCGTGGAACTGCAGGATCTCTCTGA
TGATACTAAGGAGGATGACAGGGACGGTGGGGGCTGCTCTGGTGCGGTTAAGCTGTTTGGAGTGTCGCTCTGCGGGAAGAAGAGGCAGCTCCATTCTGAT
CATCATTTCACTGCTCTTCCTCCTCCTCTTTCATCCTCCTCTTCCTTATGA
AA sequence
>Lus10036062 pacid=23141084 polypeptide=Lus10036062 locus=Lus10036062.g ID=Lus10036062.BGIv1.0 annot-version=v1.0
MQRSCNEEEEEEEEEEEESIIMQGLNKGGGGVVAPFLTKTYELVDDPMTDHIVSWGSNMNDPMSTFVVWRPPEFARDLLPNYFKHNNFSSFVRQLNTYGF
KKVVAERWEFGNDYFRKGAKQLLSHIHRRKSPHHHHHHHHNDHHHHYHNHDFIFQLPPPNYINPHHQPHHIFLPPAAAAVHSDDDDEMMMLSTLSEDNQR
LKKRNLALLSELSHMKSLYNDIIYFIQNHLKPNHPFPHHHHHPPTTTTATNNNNNNNKLVELQDLSDDTKEDDRDGGGCSGAVKLFGVSLCGKKRQLHSD
HHFTALPPPLSSSSSL
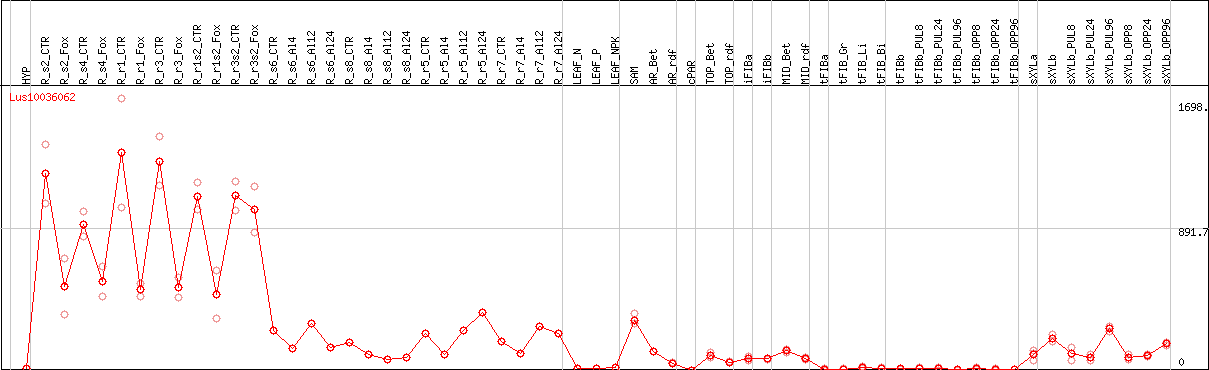
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036062 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.