External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G53240 331 / 3e-115
mMDH1
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
AT3G15020 327 / 1e-113
mMDH2
mitochondrial malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1.2)
AT2G22780 249 / 1e-82
PMDH1
peroxisomal NAD-malate dehydrogenase 1 (.1)
AT5G09660 224 / 3e-73
PMDH2
peroxisomal NAD-malate dehydrogenase 2 (.1.2.3.4)
AT3G47520 226 / 4e-73
pNAD-MDH, MDH
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
AT3G53910 62 / 2e-12
malate dehydrogenase-related (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038323
364 / 3e-128
AT3G15020 578 / 0.0
mitochondrial malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1.2)
Lus10013680
350 / 4e-123
AT1G53240 520 / 0.0
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Lus10017939
347 / 2e-121
AT1G53240 578 / 0.0
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Lus10039642
243 / 4e-80
AT2G22780 617 / 0.0
peroxisomal NAD-malate dehydrogenase 1 (.1)
Lus10011587
242 / 3e-78
AT2G22780 547 / 0.0
peroxisomal NAD-malate dehydrogenase 1 (.1)
Lus10034458
230 / 1e-74
AT3G47520 572 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
Lus10019096
228 / 9e-74
AT3G47520 575 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
Lus10012459
224 / 1e-72
AT2G22780 578 / 0.0
peroxisomal NAD-malate dehydrogenase 1 (.1)
Lus10021666
223 / 6e-72
AT3G47520 542 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G096300
332 / 1e-115
AT3G15020 541 / 0.0
mitochondrial malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1.2)
Potri.001G376500
332 / 2e-115
AT3G15020 523 / 0.0
mitochondrial malate dehydrogenase 2, Lactate/malate dehydrogenase family protein (.1.2)
Potri.004G054200
327 / 2e-113
AT1G53240 520 / 0.0
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
Potri.009G081600
247 / 6e-82
AT2G22780 602 / 0.0
peroxisomal NAD-malate dehydrogenase 1 (.1)
Potri.001G287400
244 / 6e-81
AT2G22780 588 / 0.0
peroxisomal NAD-malate dehydrogenase 1 (.1)
Potri.007G009100
242 / 4e-80
AT2G22780 612 / 0.0
peroxisomal NAD-malate dehydrogenase 1 (.1)
Potri.017G102000
233 / 6e-76
AT3G47520 591 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
Potri.004G112800
231 / 5e-75
AT3G47520 583 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
Potri.017G101900
230 / 1e-74
AT3G47520 580 / 0.0
plastidic NAD-dependent malate dehydrogenase, malate dehydrogenase (.1)
Potri.017G152000
129 / 5e-37
AT1G53240 277 / 8e-93
mitochondrial malate dehydrogenase 1, Lactate/malate dehydrogenase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0341
LDH_C
PF02866
Ldh_1_C
lactate/malate dehydrogenase, alpha/beta C-terminal domain
Representative CDS sequence
>Lus10036186 pacid=23169439 polypeptide=Lus10036186 locus=Lus10036186.g ID=Lus10036186.BGIv1.0 annot-version=v1.0
ATGATTAGCAATCCAGTGAACTCAACTGTTCCGATAGCGGCTGAGGTCTTCAAGAAAGCTGGAACTTATGATGAGAAAAGGTTATTTGGAGTTACTACTC
TTGATGTGGTCAGGGCTAGGACTTTCTATGCTGGGAAGGCCAATGTCCCTGTCGCAGGGGTCAACATTCCAGTTGTAGGAGGGCATGCCGGTGTAACAAT
TATCCCATTGTTTTCACAGAACTTTTCAGGTGAAGAGATAGTTGCTCTTACAAAGAGAACTCAAGACGGTGGCACGGAAGTCGTGGAAGCAAAGGCTGGA
AAGGGCTCTACAACATCGTCAATGGCCTACGCCGGTGCCCTTTTCGCTGATGCGTGCTTGAAGGGGCTCAATGGTTTTCCAGATTTGGTCGAATGCACAT
TTGTACAATCGAATGTAACCAAACTTCCTTTGTTTGCTTCAAAGGTGAGACTGGGAAAGAATGGCGTCGAGGAAGTCTTGGGATTGGGTCCTCTTTCTGA
CTTTGAGAAACAAGGGTTGGAAAACATGAAGTCTGAGTTGAAATCCTCCATTGAGAAAGGAATTACCTTTGCCAACAAGTGA
AA sequence
>Lus10036186 pacid=23169439 polypeptide=Lus10036186 locus=Lus10036186.g ID=Lus10036186.BGIv1.0 annot-version=v1.0
MISNPVNSTVPIAAEVFKKAGTYDEKRLFGVTTLDVVRARTFYAGKANVPVAGVNIPVVGGHAGVTIIPLFSQNFSGEEIVALTKRTQDGGTEVVEAKAG
KGSTTSSMAYAGALFADACLKGLNGFPDLVECTFVQSNVTKLPLFASKVRLGKNGVEEVLGLGPLSDFEKQGLENMKSELKSSIEKGITFANK
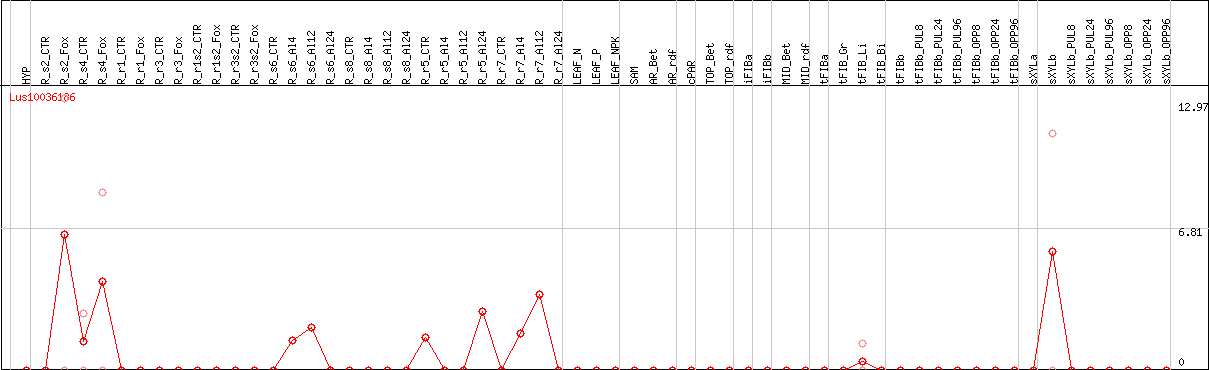
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036186 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.