External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G70190 194 / 6e-63
Ribosomal protein L7/L12, oligomerisation;Ribosomal protein L7/L12, C-terminal/adaptor protein ClpS-like (.1.2)
AT4G36420 90 / 2e-22
Ribosomal protein L12 family protein (.1)
AT4G37660 77 / 1e-17
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1)
AT3G06040 75 / 1e-16
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1.2.3)
AT2G03130 71 / 6e-16
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1)
AT3G27850 44 / 1e-05
RPL12-C
ribosomal protein L12-C (.1)
AT3G27830 44 / 2e-05
RPL12-A
ribosomal protein L12-A (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038367
280 / 2e-96
AT1G70190 243 / 3e-82
Ribosomal protein L7/L12, oligomerisation;Ribosomal protein L7/L12, C-terminal/adaptor protein ClpS-like (.1.2)
Lus10028336
94 / 5e-24
AT4G36420 176 / 1e-56
Ribosomal protein L12 family protein (.1)
Lus10041783
91 / 7e-23
AT4G36420 174 / 5e-56
Ribosomal protein L12 family protein (.1)
Lus10031756
88 / 2e-21
AT3G06040 202 / 2e-66
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1.2.3)
Lus10023839
87 / 3e-21
AT4G37660 155 / 1e-48
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1)
Lus10000093
87 / 3e-21
AT4G37660 155 / 1e-48
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1)
Lus10031180
86 / 2e-20
AT3G06040 202 / 2e-66
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1.2.3)
Lus10021011
82 / 3e-19
AT4G37660 157 / 3e-49
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1)
Lus10013078
50 / 2e-07
AT3G27830 175 / 7e-56
ribosomal protein L12-A (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G074800
207 / 7e-68
AT1G70190 204 / 1e-66
Ribosomal protein L7/L12, oligomerisation;Ribosomal protein L7/L12, C-terminal/adaptor protein ClpS-like (.1.2)
Potri.007G019100
93 / 1e-23
AT4G36420 127 / 2e-37
Ribosomal protein L12 family protein (.1)
Potri.015G077200
87 / 4e-21
AT3G06040 132 / 3e-39
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1.2.3)
Potri.004G224300
83 / 1e-19
AT4G37660 154 / 4e-48
Ribosomal protein L12/ ATP-dependent Clp protease adaptor protein ClpS family protein (.1)
Potri.001G346100
46 / 4e-06
AT3G27830 146 / 1e-44
ribosomal protein L12-A (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00542
Ribosomal_L12
Ribosomal protein L7/L12 C-terminal domain
Representative CDS sequence
>Lus10036228 pacid=23169395 polypeptide=Lus10036228 locus=Lus10036228.g ID=Lus10036228.BGIv1.0 annot-version=v1.0
ATGAGTTTGGTTCTAAGATTGCGACATCGCTTGCAAAATAGCTTGTATGCAAAATCAGCCCTTCCACTCTTGAATCAGACAAGCATCTTCAATGGTGTTC
CATCTCGTAGCCTTGCACAAGCTGCTATGAAAGAAGAAGAGGAAGAAGAAGTAGAAATTGATCAGAGGCGGCTCCCAGCAGACTACGACCCCGCAACATT
CGATCCCACCGAGCACCGTAGCCCTCCCACGGAGCGAGTTTTCAGGCTAGTTGACGAGGTTGCCAATCTGACATTAATGGAGGTGGCAGAGCTGGGTTCG
ATTATAATGAAAAGGAAAGGAATGACTGAACCGCCAGTTGTGGGAGTTATCAAGGGCGGAGTAGCTGCTGGGATGGCAATGCAGAAGGGAGGAGGATCGG
GTTCGGCCAAGGAAGAGAAGAAGCCAGAGAAGACGGTTTTCGAGCTGAAGTTGGAGTCTTATGAGGCGGCTTCGAAGATTAAGATCATTAAGGAAGTGAG
GAGCTTTACTGACTTGGGACTTAAGGAAGCGAAGGATCTGGTGGAGAAGACTCCGTCGGTGTTGAAATCGGGCGTATCGAAGGAGGAAGGCGAGAAGATT
ATCGAGAAGTTGAAAGCTTTAGGTGCTAAAGTTGTGTTGGAATGA
AA sequence
>Lus10036228 pacid=23169395 polypeptide=Lus10036228 locus=Lus10036228.g ID=Lus10036228.BGIv1.0 annot-version=v1.0
MSLVLRLRHRLQNSLYAKSALPLLNQTSIFNGVPSRSLAQAAMKEEEEEEVEIDQRRLPADYDPATFDPTEHRSPPTERVFRLVDEVANLTLMEVAELGS
IIMKRKGMTEPPVVGVIKGGVAAGMAMQKGGGSGSAKEEKKPEKTVFELKLESYEAASKIKIIKEVRSFTDLGLKEAKDLVEKTPSVLKSGVSKEEGEKI
IEKLKALGAKVVLE
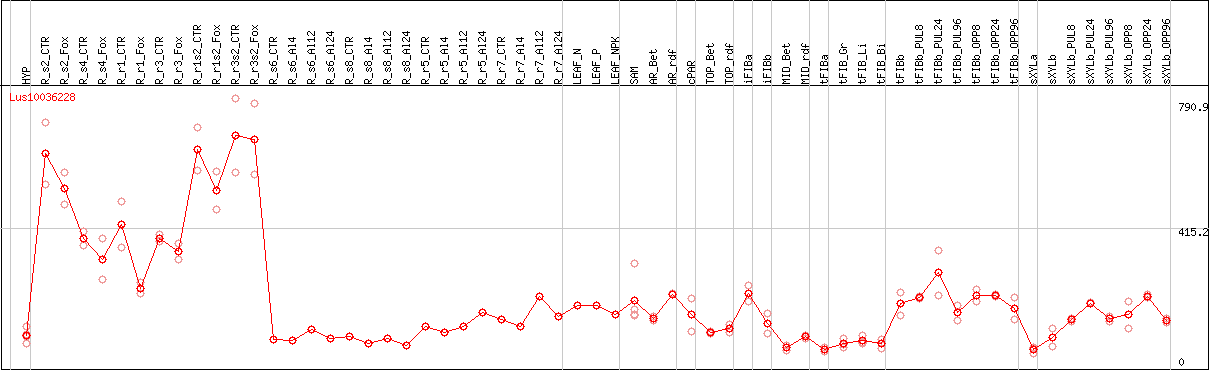
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036228 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.