Lus10036612 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10036612 pacid=23174500 polypeptide=Lus10036612 locus=Lus10036612.g ID=Lus10036612.BGIv1.0 annot-version=v1.0
ATGGCATTGTTTCGCAAACTTTTCTATAGGAAGCCTCCTGATGGCCTGCTTGAGATCTGCGAGCGCGTCTATGTTTTCGACTGCTGTTTTACTATGGATT
CATGGCAAGAGAAAGATTACAAAGGATATATAGGTGGAATTGTGGGTCAGCTGAAAGAACACTTCCCTGATGCTTCATATCTAGCATTCAATTTCCGTGA
AGGAGAGACACAAAGTCCGTTTGCTGATTTTTTGTCGGATTACGACATGACGATCATGGAGTATCCTCGGCACTATGAAGGTTGCCCTTTGCTCACAATG
GAGGTCGTCCATCATTTTCTAAGATCAGGTGAAAGTTGGCTTTCACTTGGGCAACAAAATTTGTTGCTGATGCATTGTGAAAGGGGCGGTTGGCCAGTTC
TTGCCTTTATGCTGGCTGCATTGCTGATATACAGGAAGCAATACACAGGAGAACAGAAGACATTGGATATGATCTATAGACAGGCGCATTGTGACCTCTT
GCAGTTGTGTCCTCTAAACCCGATTCCTTCTCAGCTGCGGTATCTGCACTATGTTTCAAGAAGAAACGTGGCTACAGAGTGGCCTCCACTTGACAGAGCT
CTCAACTTGGACTGTATCATTATGAGATGCATTCCGAATTTTGATGGTGAGGGTGGGTGCCGTCCTGTATTTCGTATATATGGGCAGGATCCTTTTCTTG
CTACAGATAGAACACCAAAGTTCTTGTACTCCACTCCGAAGAAAAACAAAACCTTCAGGGCTTATAAGCAGACAGAATGTGAATTAGTAAAAATCGATAT
CCGTTGTGCCATCCAAGGTGATGTTGTGCTTGAATGTGTCAGCATACGGGATGAAATGGAATCTGAGGAAATGATGTTTCGGGTAGTGTTCAATACAGCT
TTCATCAGGTCAAACATCTTGATACTCAATCGAGATGAAATTGACATATTATGGGATGCTAAAGTTCTATTCCCAAAGGAATTCAGAGCAGAGATTATTT
TCTCGGAGGTTGATGCCCATGCTACGGTCATTCCGATGGATTTTTCCATCCTGGAGGAGAAAGATGGCCTGCCGATTGAAGCTTTCGCTAAAGTTCATGA
AATCTTCAGCAACGTGGACTGGTCAGCTCCGGATGCAGATGCAGCTTTAAATATGCTTCAGCAAATGCAGGACATGGAGTTGGGACACAGTGCAGAACTA
AGTCCTCAAAGTCCAGAAGCTAGTCTAGGATCCCATAGTGACAGAAACAAGAGAATGGTTTCAGAAAGGTTATTAAGGAACTCATTTCTCTCGTGGGAAA
ATACTGAAAGTCTATACTCACCAGAAATATCTCCACGTTTTAGATTTCTCCGGGATGAGCAGCAAAGTCTTGAAAATAGTCCTAAAGTTGAATCTGATAG
CATCAGCCAAGAGACGCTCCAAACACCTGTGGATCTCACCATTAAAGACTTGTTACGAGACCACACATCACTCCAGAGGATCCTAAAGAACTCACTGGCA
CAAAGTCTTGGGCACCTGGACTCACCAAATTCATTAACGCGTACTGAGTTGGTTGGTTCTGAGTACCAAGACTCTGAAAGAAACATTAAAGTTCCAAGTC
AATCTAATGCTGACACCAAAAATACTTGTGATACATCAACACCTCAACTACAACCTGCTATGGCTTCTGCTGCTGAGAAGCTACTCTTATGTGCACAGTT
TTCAGCACCCAACGGTGCTGCCCCCAAGACTCTTTCAACTCCACAGAACGGGATTTCCTCAAAAATCGATATCCGCTGTGCCATCCAAGGTGATGTTGTG
CTTGAATGTGTCAGCATACGGGATGAAATGGAATCTGAGGAAATGATGTTTCGGGTAGTGTTCAATACAGCTTTCATCAGGTCAAACATCTTGATACTCA
ATCGAGATGAAATTGACATATTATGGGATGCTAAAGATCTATTCCCAAAGGAATTCAGAGCAGAGATTATTTTCTCGGAGGTTGATGCCCATGCTCCGGT
CATTCTGATGGATTTTGCCAGCCTGGAGGAGAAAGATGGCCTGCCGATTGAAGCTTTCGCTAAAGTTCATGAAATCTTCAGCAACGTGGACTGGTCAGCT
CCAGACGCAGATGCAGCTTTAAATATGCTCCAGCAAATGCAGGACATGGAGTTGGGACACAGTGCAGAACTAAGTCCTCAAAGTCCAGAAGCTAGTCTAG
GATCCCATAGTGACAGAAACAAGAGAATGGTTTCAGAAAGGTTATTAAGGAACTCATTTCTCTCGTGGGAAAATACTGAAAGTCTATACTCACCAGAAAT
ATCTCCACGTTTTAGATTTCTCCGGGATGAACAGAAAAGTCTTGAAAATAGCCCTAAAGATGAATCTAGCATCGGCCACGAGACGCTCCAAACACCTATG
GATCTCACTATTAAAGACTTGTCACGTGACCGCACATCACTCCAGAGGATCCTGAAGAACTCACTGGCACAAACTCTTGGGCACCTGGACTCACCAAATT
CATTAACGCGTACTGAGTTGGTTGGTTCTGAGTACCAAGACTCTGAACGAAACATTGAAGTTCCCAGTCAATCTAATGCTGACACCAAAAATACTCGTGA
TACATCAACACCTCAACTACAGCCTGCTATGGCTTCTGCTGCTGAGAAGCTACTCTTATCTGCACAGTTTTCAGCACCCAACAGTGCTGCCGCCAAGACT
CTTTCAACTCCACAGTTCACTTTTTCTCCTCCCCCACCACCAACACCTCCATTGACTTCGAAAGAACGTGGTGCTGTTAGAGTCAAACCTCCTCCTCCTC
CACCACCTCCACCTCCTGGATTTCTTCCAGGGAGTGCCACAACCTCTCCGCCTCCACCTCCTCCTATTGGTTCATCGCATTCTCATCCTTCAATTCTTGC
ACCTCCTCCTCCTCCGATTCCTTCTCGACACATTCCTACCTCTGTTCCATCTCCCCCGCCACCGCCTCCTGCTCCTCATTCATCTGCACCTTCTGCTCCT
CCTCCACCACCTCCTCCTCTGGTTTCTTCAGCGCCTCCTCCTTCTACCCCTGCTCCACCTCCTCCGCCTCCTTCTGGTAATGGACCTCCTTCCACTGTAC
CACCATCTCCTGCTCCACCAATGAGTTCTGCCCTGGGGAGCAAGGGACGGTTGTCTCGTACTATAAAGAGCAATCAGAAAAAATTGAAACCATTGCATTG
GTTAAAGTTGACAAGAGCTGTACAAGGAAGCTTGTGGGCTGAGACGCAAAAATCAGGAGAAGCCTCCAAGGCTCCAGAGATCGATATGTCAGAGCTTGAG
AGTCTTTTTGCTGCAGCAGCACCAAATCGTGGAGGAAAATCCAATGCATCAGGTCCACGTGCACCTAAGTCCGAAATAGTTCAGCTGATTGACCACAGAA
GAGCGTACAACTGTGAAATCATGCTTTCGAAGGTGAAGATTCCACTGAATGAGTTAACAGGTTATGTCCTTGCTCTGGAAGATTCAACTTTAGATATAGA
TCAAGTAGAAAATCTGATCAAGTTTTGTCCAACAAAAGAAGAGATGGAACTCCTTAAGGGCTATACTGGAGATGAGGAAAAGTTGGGGAAATGTGAACAT
TTCTTCTTAGAGTTGATGAAAGTTCCAAGAATAGAAGCAAAGCTCAGAGTTTTTGCCTTTAAAATGCAGTTTCGTTCGCAGGTCTCTGATCTCAGAAATA
GCCTACAAACGGTCAACTCTGCTGCAGAACAGGCAAGTTACTCTATTGGTCTCTTGGCAATCAGGAATTCTGGAAAGTTGAAGAGAATTATGCAGACTAT
TCTCTCTCTAGGAAATGCTTTGAACCAAGGAACTGCAAGGGGTGCTGCAATTGGGTTTAGGTTGGATAGCCTCTTAAAACTAACCGATACGCGGGCGAAG
AATAATAAAATGACCCTCATGCACTATCTTTGCAAGGTGCTTGCTGACAAACTACCGGAGCTTCTAGATTTTTCGAAAGATTTGACCAGTTTGGAACCTG
CATCCAGGGTGCAATTGAAGTTCTTGGCTGAGGAAATGCAGGCGATAAGCAAAGGATTAGAGAAGGTCATGCAAGAGCTATCCTCGTCGGAGAATGATGG
TGGCATTTCAGAACACTTCTGCAAGACACTTAAGGAGTTCCTTCGAGTTGCTGAAGCCGAAGTGAGGTCTCTGGCGTCTTTGTATTCGGGAGTGGGTAGA
AATGTAGATGCATTGATTCTTTACTTTGGCGAAGATCCAGCCCGTTGCCCTTTTGAACAAGTTTGTTCGACGCTGCTAAACTTTGTGAGATTATTCAACA
AAGCTCACGACGAGAACTGCAAGCAGCTCGAAATTGTGCTCAAGAAAGCAGCTGAGAATGAAAAGAACAACAAAGTAGATACGTCCTTGATGCAAACTCC
GGTTCAAACCAGTAGTGTCAAGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10036612 pacid=23174500 polypeptide=Lus10036612 locus=Lus10036612.g ID=Lus10036612.BGIv1.0 annot-version=v1.0
MALFRKLFYRKPPDGLLEICERVYVFDCCFTMDSWQEKDYKGYIGGIVGQLKEHFPDASYLAFNFREGETQSPFADFLSDYDMTIMEYPRHYEGCPLLTM
EVVHHFLRSGESWLSLGQQNLLLMHCERGGWPVLAFMLAALLIYRKQYTGEQKTLDMIYRQAHCDLLQLCPLNPIPSQLRYLHYVSRRNVATEWPPLDRA
LNLDCIIMRCIPNFDGEGGCRPVFRIYGQDPFLATDRTPKFLYSTPKKNKTFRAYKQTECELVKIDIRCAIQGDVVLECVSIRDEMESEEMMFRVVFNTA
FIRSNILILNRDEIDILWDAKVLFPKEFRAEIIFSEVDAHATVIPMDFSILEEKDGLPIEAFAKVHEIFSNVDWSAPDADAALNMLQQMQDMELGHSAEL
SPQSPEASLGSHSDRNKRMVSERLLRNSFLSWENTESLYSPEISPRFRFLRDEQQSLENSPKVESDSISQETLQTPVDLTIKDLLRDHTSLQRILKNSLA
QSLGHLDSPNSLTRTELVGSEYQDSERNIKVPSQSNADTKNTCDTSTPQLQPAMASAAEKLLLCAQFSAPNGAAPKTLSTPQNGISSKIDIRCAIQGDVV
LECVSIRDEMESEEMMFRVVFNTAFIRSNILILNRDEIDILWDAKDLFPKEFRAEIIFSEVDAHAPVILMDFASLEEKDGLPIEAFAKVHEIFSNVDWSA
PDADAALNMLQQMQDMELGHSAELSPQSPEASLGSHSDRNKRMVSERLLRNSFLSWENTESLYSPEISPRFRFLRDEQKSLENSPKDESSIGHETLQTPM
DLTIKDLSRDRTSLQRILKNSLAQTLGHLDSPNSLTRTELVGSEYQDSERNIEVPSQSNADTKNTRDTSTPQLQPAMASAAEKLLLSAQFSAPNSAAAKT
LSTPQFTFSPPPPPTPPLTSKERGAVRVKPPPPPPPPPPGFLPGSATTSPPPPPPIGSSHSHPSILAPPPPPIPSRHIPTSVPSPPPPPPAPHSSAPSAP
PPPPPPLVSSAPPPSTPAPPPPPPSGNGPPSTVPPSPAPPMSSALGSKGRLSRTIKSNQKKLKPLHWLKLTRAVQGSLWAETQKSGEASKAPEIDMSELE
SLFAAAAPNRGGKSNASGPRAPKSEIVQLIDHRRAYNCEIMLSKVKIPLNELTGYVLALEDSTLDIDQVENLIKFCPTKEEMELLKGYTGDEEKLGKCEH
FFLELMKVPRIEAKLRVFAFKMQFRSQVSDLRNSLQTVNSAAEQASYSIGLLAIRNSGKLKRIMQTILSLGNALNQGTARGAAIGFRLDSLLKLTDTRAK
NNKMTLMHYLCKVLADKLPELLDFSKDLTSLEPASRVQLKFLAEEMQAISKGLEKVMQELSSSENDGGISEHFCKTLKEFLRVAEAEVRSLASLYSGVGR
NVDALILYFGEDPARCPFEQVCSTLLNFVRLFNKAHDENCKQLEIVLKKAAENEKNNKVDTSLMQTPVQTSSVK
|
DESeq2's median of ratios [FLAX]
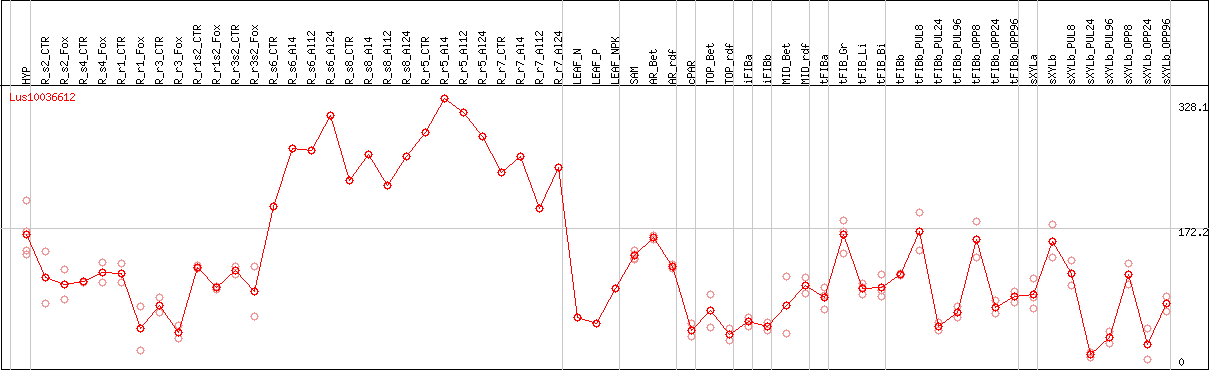
Coexpressed genes
Lus10036612 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
