External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G39620 215 / 2e-67
ATPPR5, EMB2453
EMBRYO DEFECTIVE 2453, A. THALIANA PENTATRICOPEPTIDE REPEAT 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G09820 70 / 7e-14
Pentatricopeptide repeat (PPR-like) superfamily protein (.1)
AT3G18110 70 / 1e-13
EMB1270
embryo defective 1270, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G02420 67 / 5e-13
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G74900 67 / 7e-13
OTP43
organelle transcript processing defect 43, Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT2G35130 63 / 1e-11
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT2G41720 63 / 2e-11
EMB2654
EMBRYO DEFECTIVE 2654, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT1G51965 63 / 2e-11
ABO5
ABA Overly-Sensitive 5 (.1)
AT1G63080 62 / 4e-11
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT5G39980 60 / 2e-10
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035850
410 / 1e-143
AT4G39620 579 / 0.0
EMBRYO DEFECTIVE 2453, A. THALIANA PENTATRICOPEPTIDE REPEAT 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10042362
71 / 5e-14
AT1G02420 613 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10026306
67 / 9e-13
AT1G02420 522 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10025473
66 / 3e-12
AT2G18940 970 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10041429
64 / 8e-12
AT5G39980 976 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10006962
64 / 1e-11
AT2G18940 976 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10013894
62 / 6e-11
AT5G02860 1047 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10002107
61 / 1e-10
AT5G02860 1040 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10018318
61 / 1e-10
AT2G35130 801 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G082400
263 / 3e-86
AT4G39620 625 / 0.0
EMBRYO DEFECTIVE 2453, A. THALIANA PENTATRICOPEPTIDE REPEAT 5, Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.005G046100
69 / 1e-13
AT1G12700 512 / 1e-173
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.012G123600
69 / 2e-13
AT2G35130 852 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.019G049400
67 / 6e-13
AT5G27270 1179 / 0.0
EMBRYO DEFECTIVE 976, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.007G079400
67 / 1e-12
AT1G02420 649 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.013G149800
67 / 1e-12
AT1G12700 482 / 6e-162
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.004G074500
67 / 1e-12
AT1G62930 478 / 1e-162
RNA processing factor 3, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G050180
66 / 1e-12
AT1G12700 488 / 2e-164
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
Potri.015G070200
66 / 2e-12
AT1G74900 590 / 0.0
organelle transcript processing defect 43, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.005G050500
65 / 3e-12
AT1G12700 523 / 2e-178
RNA processing factor 1, ATP binding;nucleic acid binding;helicases (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10036632 pacid=23174415 polypeptide=Lus10036632 locus=Lus10036632.g ID=Lus10036632.BGIv1.0 annot-version=v1.0
ATGGAACAAGTTTTCCAGAGCTTGCTGCAATCCAAGGAGAAGTCAACTCTTCCTACGTTTAATTCTATGATAATCAACTATGGCAAGGCTCGACAGAGAG
ACAGAGCGCAGGATGTGTTCAACAAGATGAATGATTTGAAATATACTCCGAGTTTCATTACCTACGAGAGTCTTATCATGATGTATGGATTTTGTGATTG
TGTCACTAATGCCAGAGGGGTATTTGACTCCATGGTTGAATCTGGGAAGGAGGTCAGAGTTTCGACTTTGAACGCCATGCTTGATGTTTACTGCATGAAT
AAGCTGCACATTGAAGCACACTCCCTTGTCGAAAGTGCACGGGGCCTTGGGTTGTCCCCAGATTCATCGACGTATAAGCTTCTTTATAAGGCCTACACTA
AAGCTAACAACAAAGAGCTTGTACAGAAATTGCTCAAGCGTATGGAGAAAGATGGTATCGTCATCCAGGATAGTAAATTCTTCCTAGATGCCTTGGGTGC
TTCTCAACCTTCTAAACGAGAAGGTTCAGCGGCTTCTAGCAAGGATTCTAGGAGATCAGGTAATTCACAACAGAGTCTGAAGAAAACTGGCCAGAAGCTT
ACAACCAGCCCGGTTAAGACGCATAGCTCTGTGAAGAAGATGAATGGAACTTGA
AA sequence
>Lus10036632 pacid=23174415 polypeptide=Lus10036632 locus=Lus10036632.g ID=Lus10036632.BGIv1.0 annot-version=v1.0
MEQVFQSLLQSKEKSTLPTFNSMIINYGKARQRDRAQDVFNKMNDLKYTPSFITYESLIMMYGFCDCVTNARGVFDSMVESGKEVRVSTLNAMLDVYCMN
KLHIEAHSLVESARGLGLSPDSSTYKLLYKAYTKANNKELVQKLLKRMEKDGIVIQDSKFFLDALGASQPSKREGSAASSKDSRRSGNSQQSLKKTGQKL
TTSPVKTHSSVKKMNGT
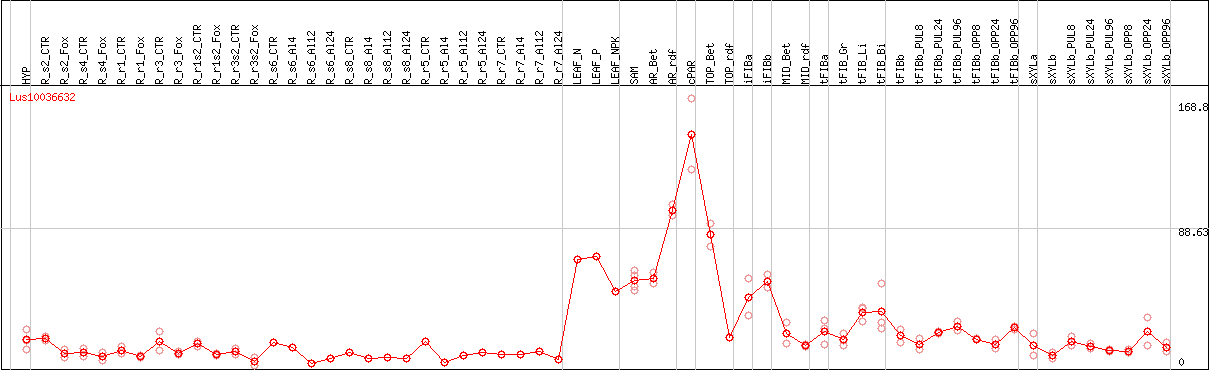
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036632 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.