Lus10036789 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10036789 pacid=23171081 polypeptide=Lus10036789 locus=Lus10036789.g ID=Lus10036789.BGIv1.0 annot-version=v1.0
ATGCAGTCTAGGATCAAGGCAATTGAGGAAGAGCCTGTAGCCTATTCTAACAAATCTACTTTAGCGTGTATGATTAATGTTGAAGTGGGTGCTGTTCTAG
CTGTAATGCGGAGGAACGTGAGGTGGGGCAGTCGGTATATGTCGGGTGATGATCAATTAGAGCACTCTTTGATACAGTCTCTTAAGGGATTGAGGAAGCA
GATATTCTCATGGCAGCAACCGTGGAATTCTGTTAATCCTGCAGATTACCTCCAACCGTTTTTGGAAGTTATTCGATCGGCTGAAACTGGTGCACCAATT
ACTGGGGTTGCTTTATCGTCTGTGTACAAGATACTGACACTGGATGTGATTGATCAAAATACTGTGAATGTGGAAGATGCTATGCACTTGGTGGTTGAAG
CGGTGACTGGTTGTCGATTTGAGCTGACAGATCCTGCATCTGAGGAAGTTGTCCTGATGAAAGTTCTTCAGGTTCTTCTTGCGTGCATGAAAAGTAAAGC
ATCAGTTATGTTGAGCAATCAGGATGTCTGTACAGTTGTAAATACTTGTTTCCGTATAGTCCATCAAGCAGGAAACAAGGGTGAGCTATTGCAGCGGATG
GCTCGTTACACAATGCACGAACTTATTAGATGCATTTTTTCACATCTTCCTGATGTTGATAAGACAGAACAAGCATTGGTGAATGGAGTTAGTACCGTCG
AGCCAGAGATAGGTGGACTAGACAGCAATAATGCATTTGGAAGTAAACAAGTAGAAAATGGAAACAGTAACACTGAATTAGATGGCCAGCCAACATCTTT
GAGCCTAAGTTCCAATTCCACTGCTATGGTGTCAAGTACTATGGAAGATAGCTCAATAGGGGCGGGTAATGGAAGTGATGCCTTTTCTTATGATCAGCAA
CTTATGAAAGAACCATATGGGGTTCCTTGCATGGTGGAAATATTTCATTTCTTGTGTTCATTGTTAAATGCTGTTGAGCATATGGGAATGGATCCCAGAT
CAAATACAATAGCGTTTGATGAAGATGTGCCCCTTTTCGCCTTGGGCTTAATTAATGCAGCTATAGAATTGGCTGGACCTTCAATACGCTATCACCCCAG
GTTACTGACACTGATTCAGGATGAACTGTTCCAGAATCTGATGCAATTTGGTTTGTCAATGAGCCCACTTATTCTCTCAATGGTGTGTAGTATAGTTCTG
AGCCTGTACCATCATTTGTATTCCGAAATTAAGCTACAACTTGAGGCTTTCTTCTCTTGTGTGGTTTTAAGACTGGCACAGAGCAGATATGGTGCTTCAT
ATCAGCAGCAAGAGGTTGCTATGGAGGCTCTTGTAGATTTCTGTAGACAGAAGACTTTCATGGTGGAGATGTATGCTAACTTGGATTGTGACATAACATG
CAGCAATGTGTTTGAAGAACTCGCTAATCTGTTGTCAAAAAGTGCATTTCCTGTTAACTGCCCTTTGTCTGCTATGCACATTCTTGCTTTGGATGGCCTG
ATTGCTGTAATTGAGGGGATGGCTGAGAGGATAGAAAATGAGCCTGGTAATCCTGAGCAGGTTCCAGTGAGCCTTGAGGAGTATACACCATTTTGGGAGG
TGAAATGTGACAATTACAACGATCCTACTCAATGGGTTCCGTTTGTCCGCAGAAGAAAGTACATCAAGCGAAGGTTGATGATTGGAGCTGACCATTTCAA
TCGTGATCCCAAGAAGGGCCTGGAATTTCTCCAAGGAACACATCTATTACCAGAAAAACTTGACCCGCAAAGTGTGGCATGCTTTTTCAGATACACAGCT
GGATTGGATAAAAACCTTGTTGGGGACTTCCTCGGCAATCATGACGAGTTTTGTGTTCAGGTTCTTCATGAGTTTGCTTGTACTTTTGATTTTGAAGATA
TGAATTTGGATACAGCTCTGAGGTTGTTCTTAGAAACTTTTAGGTTGCCAGGAGAATCACAGAAAATTCAAAGAGTGCTTGAGGCATTTTCTGAGAGATA
CTACGAGCAATCACCACAGATTTTAGCTAACAAGGATGCTGCTCTTCTTCTGTCTTACTCTATTATAATGCTCAACACAGATCAGCACAATGTACAGGTG
AAGAAAAAGATGACCGAAGAGGATTTTATCCGGAATAATAGGCATATCAATGGAGGCCAGGATTTGCCACGGGAATTCCTGACTGAACTTTATTACTCAA
TTTGCAAGAATGAGATTCGTACTATTCCTGATCAAGGTGCTGGTTTTGCTGAAATGAACCCCAGCCGCTGGATTGACCTGATGCTCAAGTCCAGGAAAAC
CGCTCCTTATATCGTTTGTGATTCAATAGCCGACCTTGACCACGACATGTTTGCAATAATGTCTGGTCCAACCATAGCTGCCATATCTGTTGTATTTGAT
CATGCAGAACATGAAGAAGTTTACCAAACATGCATCAACGGTTTTCTAGCTGTTGCAAACATTTCAGCTTGCCATCATCTTGAAGACGTACTGGATGATC
TAGTAGTATCTCTGTGTAAGTTCACCACTCTTTTGAATCCATCGTCTATTGAGGAACCTGTGCTAGCATTTGGCGACGATGCAAAGGCTAGGATGGCAAC
TGTGACTGTTTTCACCATTGCAAACAGGTACGGTGACTATATTCGCACTGGGTGGAGAAATATTGTGGACTGCATTCTACGATTGCACAAACTCGGTCTC
CTGCCAGCCCGCGTAGCTAGTGATGCTGCTGATGAATTAGAAACTTTTCCCGAGCTTACACAGGGGAAGCCCGTCACAAATCCTGTGTCTTCTGTTCAAA
TGCAACAGGCTGGAACTCCCAGGAAGTCGTCTGGATTTATGGGTCGGTTCAGTCAACTCTTATCTCTTGATGTTGAGGAGCCTAGATTACAACCTACGGG
GCAAGAGCTTGCTGCTCACCAACGCACTGTCGAAACTATTCAGCAGTGCCACGTAGACAGTATATTCACCGAGAGTAAATTCTTGCAAGCTGAATCCTTA
TCGCAGCTTGCAAAGGCACTAATTTGGGCTGCAGGTCGGCCACAGAAAGGGGCTAGTTCTCCAGAGGATGAAGATACATCCGTATTCTGCCTGGAGTTGC
TGATTGCAATCACGTTAAACAACCGAGATCGGATAATCCTATTATGGCAAGGTGTGTACGAGCACATTGCTAGCATCGTTCAGTCAACTGTGATGCCGTG
TGCTTTAGTAGAGAAGGCTGTATTTGGACTTTTACGAATATGCCAGCGGCTGCTCCCCTACAAAGAAAACTTTGCGGACGAACTTTTAAGGTCACTTAAT
CTTGTTCTGAAGCTTGATGCCCGTGTTGCTGATGCATACTGTGAGCAAATCACGCAGGAAGTAACTCGCCTAGTAAAGGCTAATGCCACTCACATCAGAT
CTCTAATGGGATGGCGAACAATCACATCCTTGCTTTCAATCACAGCTCGGCATCCGGAAGCTTCCGATGCAGGATTCGATGCCCTGCTATTTATCATGAG
CGGTGGTGCTCACCTCTTGCCAGCCAATTACATCCTCTGCGTTGATGCAGCTAGACAATTTGCAGAGTCTCGAGTTGGACAATCTGAAAGGTCTTTACAT
GCTCTAGATCTTATGTCAGGTTCCGTCGATTGCTTAACGCAGTGGTCTAACGAGGCTAAGGCAGCCATGGGAGATGAGGAAGCTGCAAAGCTGTTGCAGG
ATATCGGCGAAATGTGGTTGAGATTGGTGCAGGGGGTAAGAAAAGTTTGTTTGGATCAAAGAGAAGAGGTCAGGAACCACGCCCTTCTATCGTTGCAAAA
ATGTCTCACAAACAATCTTGACCTTCTGCCCGGTTTGTGGTTACAGTGCTTCGACACGGTGATCTTCACAATGCTGGATGACCTGCTCGAAATCGCTCAA
GGACACTCCCAAAAGGAGTACAGAAACATGGAAGGGACACTTATCATCGCAATCAAACTCATGTCGAAAGTGTTCCTCCAGCTCCTGCAAGACCTCGCGC
AGCAACTGACGACATTCTGCAAGTTGTGGCTGGGAGTGCTGGGGAGGACGGAGAAGTACATGAAGGTGAAAATAAGAGGGAAGAAGAGCGAAAAGCTTCA
GGAAACTATACCCGAGCTCCTCAAGAACATCTTACTCATCATGAAGACCACTGGAGTTCTGGTTCGCCCCCTGGAGTTCTGGTCCAGAGCAGGCCCAGCC
TGTGGGAGCTGA
|
||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10036789 pacid=23171081 polypeptide=Lus10036789 locus=Lus10036789.g ID=Lus10036789.BGIv1.0 annot-version=v1.0
MQSRIKAIEEEPVAYSNKSTLACMINVEVGAVLAVMRRNVRWGSRYMSGDDQLEHSLIQSLKGLRKQIFSWQQPWNSVNPADYLQPFLEVIRSAETGAPI
TGVALSSVYKILTLDVIDQNTVNVEDAMHLVVEAVTGCRFELTDPASEEVVLMKVLQVLLACMKSKASVMLSNQDVCTVVNTCFRIVHQAGNKGELLQRM
ARYTMHELIRCIFSHLPDVDKTEQALVNGVSTVEPEIGGLDSNNAFGSKQVENGNSNTELDGQPTSLSLSSNSTAMVSSTMEDSSIGAGNGSDAFSYDQQ
LMKEPYGVPCMVEIFHFLCSLLNAVEHMGMDPRSNTIAFDEDVPLFALGLINAAIELAGPSIRYHPRLLTLIQDELFQNLMQFGLSMSPLILSMVCSIVL
SLYHHLYSEIKLQLEAFFSCVVLRLAQSRYGASYQQQEVAMEALVDFCRQKTFMVEMYANLDCDITCSNVFEELANLLSKSAFPVNCPLSAMHILALDGL
IAVIEGMAERIENEPGNPEQVPVSLEEYTPFWEVKCDNYNDPTQWVPFVRRRKYIKRRLMIGADHFNRDPKKGLEFLQGTHLLPEKLDPQSVACFFRYTA
GLDKNLVGDFLGNHDEFCVQVLHEFACTFDFEDMNLDTALRLFLETFRLPGESQKIQRVLEAFSERYYEQSPQILANKDAALLLSYSIIMLNTDQHNVQV
KKKMTEEDFIRNNRHINGGQDLPREFLTELYYSICKNEIRTIPDQGAGFAEMNPSRWIDLMLKSRKTAPYIVCDSIADLDHDMFAIMSGPTIAAISVVFD
HAEHEEVYQTCINGFLAVANISACHHLEDVLDDLVVSLCKFTTLLNPSSIEEPVLAFGDDAKARMATVTVFTIANRYGDYIRTGWRNIVDCILRLHKLGL
LPARVASDAADELETFPELTQGKPVTNPVSSVQMQQAGTPRKSSGFMGRFSQLLSLDVEEPRLQPTGQELAAHQRTVETIQQCHVDSIFTESKFLQAESL
SQLAKALIWAAGRPQKGASSPEDEDTSVFCLELLIAITLNNRDRIILLWQGVYEHIASIVQSTVMPCALVEKAVFGLLRICQRLLPYKENFADELLRSLN
LVLKLDARVADAYCEQITQEVTRLVKANATHIRSLMGWRTITSLLSITARHPEASDAGFDALLFIMSGGAHLLPANYILCVDAARQFAESRVGQSERSLH
ALDLMSGSVDCLTQWSNEAKAAMGDEEAAKLLQDIGEMWLRLVQGVRKVCLDQREEVRNHALLSLQKCLTNNLDLLPGLWLQCFDTVIFTMLDDLLEIAQ
GHSQKEYRNMEGTLIIAIKLMSKVFLQLLQDLAQQLTTFCKLWLGVLGRTEKYMKVKIRGKKSEKLQETIPELLKNILLIMKTTGVLVRPLEFWSRAGPA
CGS
|
DESeq2's median of ratios [FLAX]
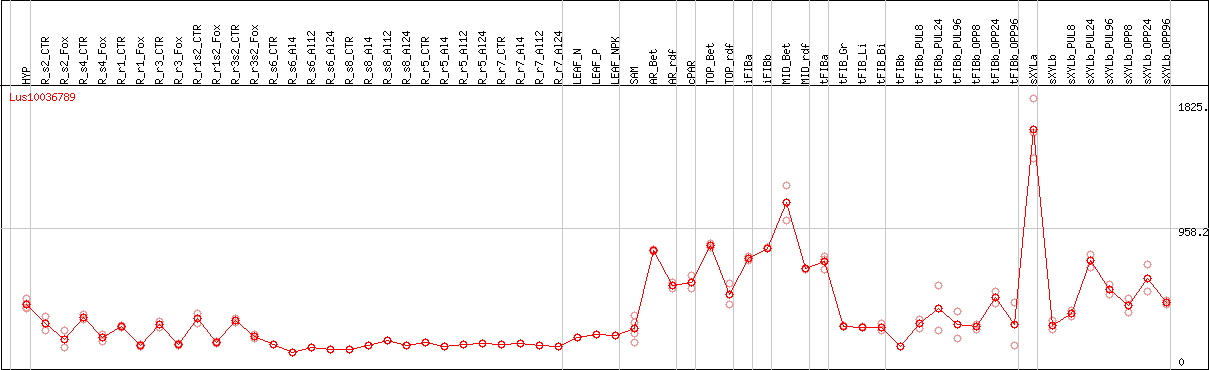
Coexpressed genes
Lus10036789 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
