External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G70630 449 / 8e-157
Nucleotide-diphospho-sugar transferase family protein (.1)
AT2G35610 76 / 5e-15
XEG113
xyloglucanase 113 (.1)
AT1G56550 66 / 6e-12
RXGT1
RhamnoGalacturonan specific Xylosyltransferase 1 (.1)
AT1G19360 62 / 1e-10
RRA3
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
AT1G75120 59 / 2e-09
RRA1
REDUCED RESIDUAL ARABINOSE 1, Nucleotide-diphospho-sugar transferase family protein (.1)
AT1G75110 59 / 2e-09
RRA2
REDUCED RESIDUAL ARABINOSE 2, Nucleotide-diphospho-sugar transferase family protein (.1)
AT1G28695 56 / 9e-09
Nucleotide-diphospho-sugar transferase family protein (.1)
AT4G01750 55 / 2e-08
RGXT2
rhamnogalacturonan xylosyltransferase 2 (.1)
AT4G01220 55 / 2e-08
MGP4
male gametophyte defective 4, Nucleotide-diphospho-sugar transferase family protein (.1.2)
AT4G01770 54 / 7e-08
RGXT1
rhamnogalacturonan xylosyltransferase 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006212
569 / 0
AT1G70630 610 / 0.0
Nucleotide-diphospho-sugar transferase family protein (.1)
Lus10005034
76 / 5e-15
AT2G35610 886 / 0.0
xyloglucanase 113 (.1)
Lus10027804
74 / 2e-14
AT2G35610 927 / 0.0
xyloglucanase 113 (.1)
Lus10026327
69 / 6e-13
AT1G19360 610 / 0.0
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
Lus10032710
65 / 1e-11
AT1G19360 629 / 0.0
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
Lus10018812
65 / 1e-11
AT1G19360 636 / 0.0
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
Lus10042340
63 / 5e-11
AT1G19360 603 / 0.0
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
Lus10017981
59 / 9e-10
AT1G14590 303 / 9e-101
Nucleotide-diphospho-sugar transferase family protein (.1)
Lus10041973
59 / 9e-10
AT1G14590 302 / 3e-100
Nucleotide-diphospho-sugar transferase family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G186800
455 / 1e-157
AT1G70630 668 / 0.0
Nucleotide-diphospho-sugar transferase family protein (.1)
Potri.010G046100
347 / 4e-116
AT1G70630 497 / 5e-171
Nucleotide-diphospho-sugar transferase family protein (.1)
Potri.004G235800
71 / 2e-13
AT2G35610 976 / 0.0
xyloglucanase 113 (.1)
Potri.009G113300
69 / 4e-13
AT1G19360 655 / 0.0
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
Potri.014G042700
69 / 7e-13
AT1G19360 635 / 0.0
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
Potri.014G051600
68 / 1e-12
AT1G28710 344 / 5e-117
Nucleotide-diphospho-sugar transferase family protein (.1.2.3)
Potri.002G139800
67 / 2e-12
AT1G28710 337 / 1e-114
Nucleotide-diphospho-sugar transferase family protein (.1.2.3)
Potri.002G134400
64 / 2e-11
AT1G19360 622 / 0.0
reduced residual arabinose 3, Nucleotide-diphospho-sugar transferase family protein (.1)
Potri.002G166000
60 / 6e-10
AT4G01220 524 / 0.0
male gametophyte defective 4, Nucleotide-diphospho-sugar transferase family protein (.1.2)
Potri.014G092400
54 / 3e-08
AT4G01220 530 / 0.0
male gametophyte defective 4, Nucleotide-diphospho-sugar transferase family protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF03407
Nucleotid_trans
Nucleotide-diphospho-sugar transferase
Representative CDS sequence
>Lus10036860 pacid=23171327 polypeptide=Lus10036860 locus=Lus10036860.g ID=Lus10036860.BGIv1.0 annot-version=v1.0
ATGTCGGGTTGCTCAGTTAGTAATATTGAGTTGGGGAACTTGCAGCCATTGCTTGAGTTCCAAGCTTCCCTGGAATCGTTGCTTGAGATAACTGCGGACA
AGACTCGGACAGTTGTGCTTGCTGTTGCTGGATATAGTTACAAGGACATGTTGATGAGTTGGGTTTGCAGATTGCGCCGTTTGAAAGTTACCAACTTTAT
CATTTGTGCTCTGGACCATGAAACCTTTGAATTTTCTTTTCTTCAGGGTTTGCCAGTTTTCTATGATCCATCAGCAGCACCAAGGAACATCAGCTTCAAC
GATTGCCATTTCGGAACGAAATGCTTCCAGAGAGTAACAAAGGTTAAATCCAGGTTAGTTCTCAAGATATTGAAGCTAGGATACAATGTACTCCTCAGTG
ACGTAGATGTCAACTGGTTTCGAAATCCACTGCAGTTCCTCACCTCATTCGGTCCCGGGTTTCTCGTTGCTCAGGCCGATGAATACAACAAAACAGGTCC
AATAAACTTACCAAGGCGCCTGAACTCTGGCTTTTACTTTGCTCGTTCAGATGCTCCAACTATCACCGTCATCGATAAGGTAGTGATCCATGCTGCTAAT
TCAGGTCTTTCAGAGCAGCCAAGCTTTTACGACGTAATGTGTGGTGAAGGTGGATCGAATCGAATCAGTGATAACCGATGCCTAGATCCGGAAACGAATC
TGACTGTTCATTTCTTGGATAGAGACCTTTTCCCCAACGGTGCATACTTCGACCTGTGGGATGAGCCGAACGTAACAGCATCTTGTATGAAGAGAAGTTG
CTTTGTTCTTCACAATAACTGTTTGAATGGAAGATTGAAGAAGCTCGAACGCCAGGTTTCGTCCGGTCTGTGGGAGTATGACGGTAGCAAAGAATGTGTC
TTTCCTGCATAA
AA sequence
>Lus10036860 pacid=23171327 polypeptide=Lus10036860 locus=Lus10036860.g ID=Lus10036860.BGIv1.0 annot-version=v1.0
MSGCSVSNIELGNLQPLLEFQASLESLLEITADKTRTVVLAVAGYSYKDMLMSWVCRLRRLKVTNFIICALDHETFEFSFLQGLPVFYDPSAAPRNISFN
DCHFGTKCFQRVTKVKSRLVLKILKLGYNVLLSDVDVNWFRNPLQFLTSFGPGFLVAQADEYNKTGPINLPRRLNSGFYFARSDAPTITVIDKVVIHAAN
SGLSEQPSFYDVMCGEGGSNRISDNRCLDPETNLTVHFLDRDLFPNGAYFDLWDEPNVTASCMKRSCFVLHNNCLNGRLKKLERQVSSGLWEYDGSKECV
FPA
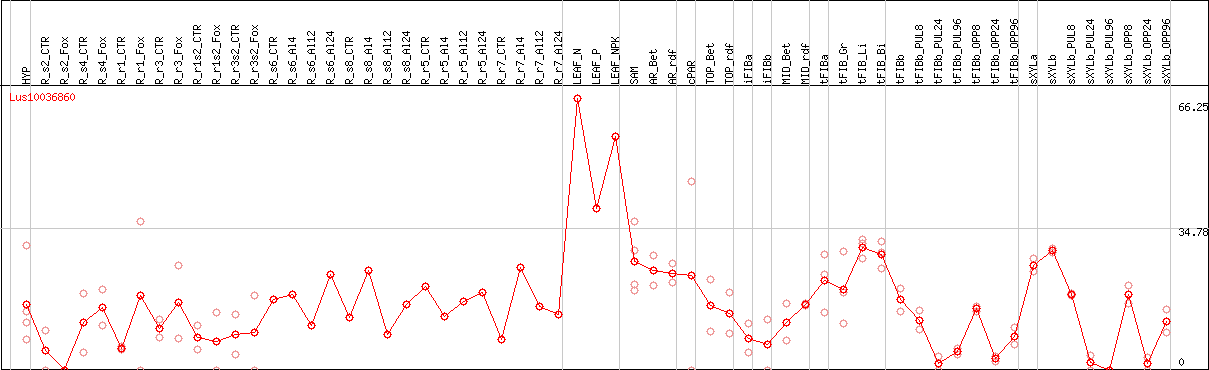
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036860 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.