External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G13900 309 / 1e-100
Purple acid phosphatases superfamily protein (.1)
AT2G03450 300 / 3e-97
PAP9, ATPAP9
purple acid phosphatase 9 (.1)
AT1G13750 103 / 1e-24
Purple acid phosphatases superfamily protein (.1)
AT4G24890 99 / 9e-23
ATPAP24, PAP24
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 24, purple acid phosphatase 24 (.1)
AT5G50400 97 / 3e-22
ATPAP27, PAP27
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
AT3G52780 48 / 5e-06
ATPAP20, PAP20
Purple acid phosphatases superfamily protein (.1.2)
AT3G52820 45 / 6e-05
ATPAP22, PAP22
purple acid phosphatase 22 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037091
448 / 7e-155
AT1G13900 899 / 0.0
Purple acid phosphatases superfamily protein (.1)
Lus10004661
339 / 3e-112
AT1G13900 860 / 0.0
Purple acid phosphatases superfamily protein (.1)
Lus10026652
332 / 1e-109
AT1G13900 875 / 0.0
Purple acid phosphatases superfamily protein (.1)
Lus10039565
101 / 1e-23
AT5G50400 362 / 6e-117
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
Lus10016356
92 / 1e-20
AT4G24890 332 / 7e-106
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 24, purple acid phosphatase 24 (.1)
Lus10012144
91 / 3e-20
AT5G50400 885 / 0.0
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
Lus10037079
90 / 1e-19
AT1G13750 928 / 0.0
Purple acid phosphatases superfamily protein (.1)
Lus10036904
89 / 1e-19
AT1G13750 928 / 0.0
Purple acid phosphatases superfamily protein (.1)
Lus10019771
89 / 1e-19
AT5G50400 317 / 4e-100
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G160500
354 / 2e-118
AT1G13900 903 / 0.0
Purple acid phosphatases superfamily protein (.1)
Potri.003G202200
100 / 3e-23
AT5G50400 359 / 8e-116
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
Potri.010G158250
100 / 3e-23
AT1G13750 967 / 0.0
Purple acid phosphatases superfamily protein (.1)
Potri.001G023400
98 / 2e-22
AT5G50400 345 / 1e-110
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
Potri.010G158400
95 / 2e-21
AT1G13750 962 / 0.0
Purple acid phosphatases superfamily protein (.1)
Potri.015G095900
95 / 2e-21
AT5G50400 838 / 0.0
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
Potri.012G097400
92 / 2e-20
AT5G50400 947 / 0.0
ARABIDOPSIS THALIANA PURPLE ACID PHOSPHATASE 27, purple acid phosphatase 27 (.1)
Potri.008G096000
91 / 3e-20
AT1G13750 927 / 0.0
Purple acid phosphatases superfamily protein (.1)
PFAM info
Representative CDS sequence
>Lus10036895 pacid=23171286 polypeptide=Lus10036895 locus=Lus10036895.g ID=Lus10036895.BGIv1.0 annot-version=v1.0
ATGTCGACTCCCCGTCGAAGCTTGACTGGCTCGGAATCTACTCCCCTCCTTCTTCCCCTCACGACCACTTCATCGGGTACAAGTTCCTATCCTCCTCCCC
CGGGTGGGAATCCGGGTCGGGTTCCCTGTCCATACCAATCACCAACCTCCGATCCAACTACTCGTTCCGGATCTTCCGCTGGACGGAAGACCGCCAAGCT
CCGTGACCAGGACGAGAATCCGCTTCCGGGGACGAAGCACCTGCTAGCGGAGTCTGATGGAGTCGTCCTGTTCGGGTCGGGGAATGGGCCGGAGCAGATC
CATCTCGCGTTTACGGATCGAGCTGACGAGATGAGGGTGATGTTCTTGACTGGAGATGGCGACGAGAGGAAGGTCCGGTGGGGGGAGACCGACGGCGCGT
GGAGTAAGGAAGCGGCGGCGCGTGTGGGGAGGTATGAAAAGGAGGATATGTGTGATTCGCCGGCGAATAGCTCTGTTGGGTGGAGAGATCCGGGTTGGAT
TCATGATGGGGTAATGAAGGATTTGAAGCAAGGTTTCAGATACTATTACCAGGTTGGAAGTGACTCAAAGGGATGGAGTAAAGCTCAAACCTTTGTGTCA
AGGAACGAAGATTCAGACGAGACAACAGCATTCCTATTTGGCGACATGGGAACATCAGCACCATATGCAACATTCATCCGTACGCAATCCGAAAGCGTAT
CAACAGTGAACTTGATCCTCCGTGATATGGAGAGACTCGGGGAGAAGCCAGCATTTGTTTCCCACATTGGTGACATAAGCTACGCAAGGGGGTATTCATG
GATATGGGACCATTTCTTCACACAAATTGAGCCGATCTCTTCGAGATTGGCATACCCGGAACCTCTACTGCTCGTTCGATTTCGGGGCGGTGCATTTCGT
TTACATCTCAACTGA
AA sequence
>Lus10036895 pacid=23171286 polypeptide=Lus10036895 locus=Lus10036895.g ID=Lus10036895.BGIv1.0 annot-version=v1.0
MSTPRRSLTGSESTPLLLPLTTTSSGTSSYPPPPGGNPGRVPCPYQSPTSDPTTRSGSSAGRKTAKLRDQDENPLPGTKHLLAESDGVVLFGSGNGPEQI
HLAFTDRADEMRVMFLTGDGDERKVRWGETDGAWSKEAAARVGRYEKEDMCDSPANSSVGWRDPGWIHDGVMKDLKQGFRYYYQVGSDSKGWSKAQTFVS
RNEDSDETTAFLFGDMGTSAPYATFIRTQSESVSTVNLILRDMERLGEKPAFVSHIGDISYARGYSWIWDHFFTQIEPISSRLAYPEPLLLVRFRGGAFR
LHLN
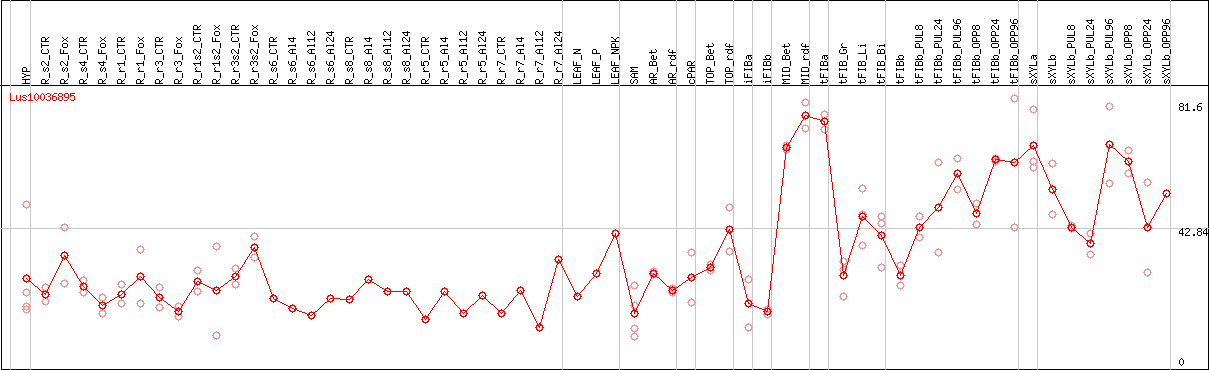
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10036895 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.